CRISPR/Cas-based Diagnostics and Gene Therapy
1Graduate School of Medical Science and Engineering, Korea Advanced Institute of Science and Technology (KAIST), Daejeon 34141, Republic of Korea
2Department of Materials Science and Engineering, Korea Advanced Institute of Science and Technology (KAIST), Daejeon 34141, Republic of Korea
*Correspondence to: Pei Li. E-mail: lipei@kaist.ac.kr
Received: January 7 2021; Revised: February 21 2021; Accepted: March 11 2021; Published Online: April 23 2021
Cite this paper:
Meiyu Qiu and Pei Li. CRISPR/Cas-based Diagnostics and Gene Therapy. BIO Integration 2021; 2(3): 121–129.
DOI: 10.15212/bioi-2020-0048. Available at: https://bio-integration.org/
Download citation
© 2021 The Authors. This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See https://bio-integration.org/copyright-and-permissions/
Abstract
Clustered regularly interspaced short palindromic repeats (CRISPR) technology, an easy, rapid, cost-effective, and precise gene-editing technique, has revolutionized diagnostics and gene therapy. Fast and accurate diagnosis of diseases is essential for point-of-care-testing (POCT) and specialized medical institutes. The CRISPR-associated (Cas) proteins system shed light on the new diagnostics methods at point-of-care (POC) owning to its advantages. In addition, CRISPR/Cas-based gene-editing technology has led to various breakthroughs in gene therapy. It has been employed in clinical trials for a variety of untreatable diseases, including cancer, blood disorders, and other syndromes. Currently, the clinical application of CRISPR/Cas has been mainly focused on ex vivo therapies. Recently, tremendous efforts have been made in the development of ex vivo gene therapy based on CRISPR-Cas9. Despite these efforts, in vivo CRISPR/Cas gene therapy is only in its initial stage. Here, we review the milestones of CRISPR/Cas technologies that advanced the field of diagnostics and gene therapy. We also highlight the recent advances of diagnostics and gene therapy based on CRISPR/Cas technology. In the last section, we discuss the strength and significant challenges of the CRISPR/Cas technology for its future clinical usage in diagnosis and gene therapy.
Keywords
Clinical trails, CRISPR/Cas, diagnostics, gene therapy, point-of-care-testing.
Introduction of the CRISPR/Cas system
Over the past decades, site-specific DNA-binding proteins have significantly changed the field of biotechnology and medical research. Nevertheless, protein engineers are often required for gene editing due to the complexity of developing DNA-binding proteins for specific target sites [1]. Advances in CRISPR/Cas technology have dramatically expanded the molecular toolbox for bioresearchers [2]. The CRISPR/Cas system is an adaptive immune system developed in bacteria and archaea, which can be used for defending against exogenous genetic elements. In bacteria and archaea, evolved Cas proteins cleave the nucleic acids of invading viruses and plasmids [1]. Moreover, the bacterial cells can use CRISPR-Cas systems to protect themselves from renewed infection. CRISPR/Cas grants a kind of immune memory after the first infection defense by inserting a short fragment of exogenic DNA [3]. Therefore, the CRISPR-Cas system can provide protection for the host through the cleavage of foreign DNAs by Cas proteins under the guidance of CRISPR RNA (crRNA). Notably, CRISPR/Cas target specificity depends on the specific base pairing of nucleic acids instead of the protein–DNA recognition [4]. Since its discovery, CRISPR/Cas technology has been regarded as a universal gene editing tool to edit the genetic materials with high precision for biological applications, such as molecular diagnostics, genome engineering, and bacterial metabolic engineering [5–13].
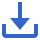
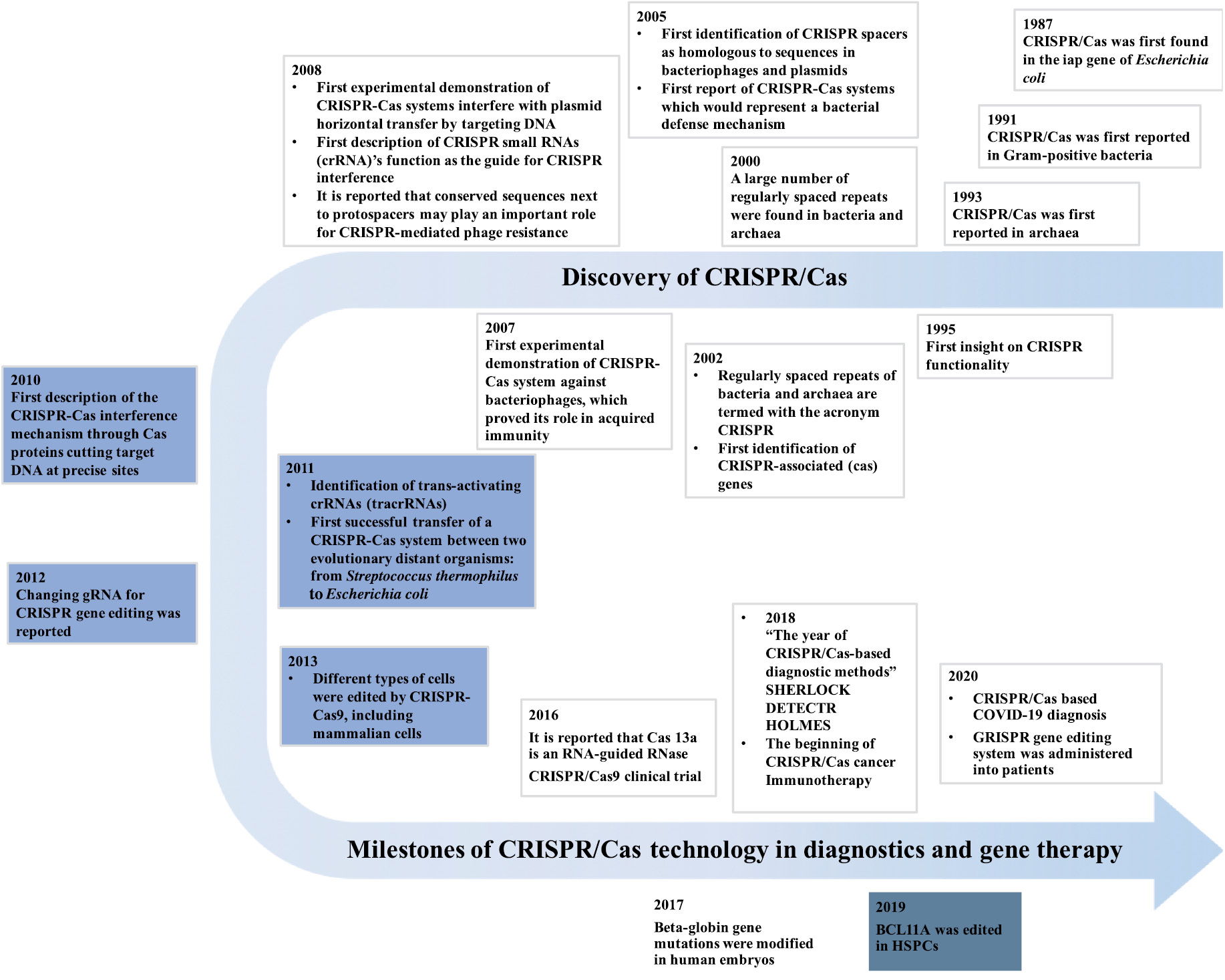
As stated above, CRISPR/Cas technology originated from bacteria and archaea [1] (Figure 1) The regularly spaced direct repeats (DR) were discovered in the Escherichia coli’s iap gene in 1987, which was the first hint of the CRISPR/Cas system’s existence [14, 15]. Since then, six types and 22 subtypes of CRISPR have been reported [16]. CRISPR/Cas was first found in Gram-positive bacteria in 1991. Interspaced DR were identified in strains of Mycobacterium tuberculosis complex (MTBC) [17]. Following the discoveries of CRISPR/Cas in Gram-negative and Gram-positive bacteria, a research group in Spain discovered repeated-spacer clusters in archaea in 1993 [18]. From 2000 to 2011, more evidence showed that the CRISPR/Cas system is involved in bacterial immunity [19, 20].
Figure 1 Discovery of CRISPR/Cas and the milestones of CRISPR/Cas development in diagnostics and gene therapy.
Despite the numerous discoveries in the CRISPR/Cas system, its applications in diagnostics and gene therapies have only been developed in recent decades. As recently as 2012, it was reported that a CRISPR nuclease can be programmed by merely varying its guide RNA (gRNA) sequence, which makes it extremely useful for genome editing [21]. In 2013, scientists successfully edited different types of cells by using a specific Type II Cas protein termed Cas9 [22]. The gRNA is the key for CRISPR-Cas9 systems. It can recognize the target sequence with high specificity [2, 5].
Subsequently, Cas9 protein cuts the targeted sequence near a protospacer adjacent motif (PAM) [2]. In 2015, two Type V CRISPR/Cas proteins, Cas-12a and Cas-13a, were discovered for genome editing, and they were initially named Cpf1 and C2c2, respectively [23–25]. Notably, Science named the powerful genome-editing technique–CRISPR as the breakthrough of the year [26]. In 2016, Abudayyeh et al. discovered that Cas-13a is an RNA-guided RNase, which provides protection against RNA phages [27, 28].
Besides diagnostics, CRISPR/Cas systems revolutionized the area of gene therapy-based healthcare. In 2013, Cong et al. engineered a CRISPR/Cas system and proved that Cas9 could precisely induce cleavage in mammalian cells [29]. This study raised the possibility of the implementation of CRISPR-Cas9 as a therapeutic tool for gene-editing. Since then, CRISPR/Cas technology has been tested in many clinical trials for a large variety of diseases, such as metastatic gastrointestinal epithelial cancer, T-cell malignancies, and refractory B-cell malignancies [30]. In 2017, the first CRISPR/Cas-based clinical trial to treat the human immunodeficiency virus (HIV) was carried out by several Chinese scientists [31]. In 2018, a T-cell receptor (TCR) gene-modified T cell was tested in clinical trials to treat patients with different cancers [32]. This study proved the possibility of using CRISPR-Cas9 in cancer treatment [33]. In 2019, the treatment of Leber’s congenital amaurosis (LCA) based on in vivo CRISPR-Cas9 gene therapy was tested in clinical trials [34].
This article reviews the recent progress of the CRISPR/Cas system in clinical diagnostics and gene therapy-based treatments in clinical trials. We also discuss the CRISPR/Cas systems’ strengths as well as its limitations, such as off-target effects (OTEs), tissue accessibility, and ethical concerns for their future applications.
CRISPR-Cas12a and 13a-based diagnostics
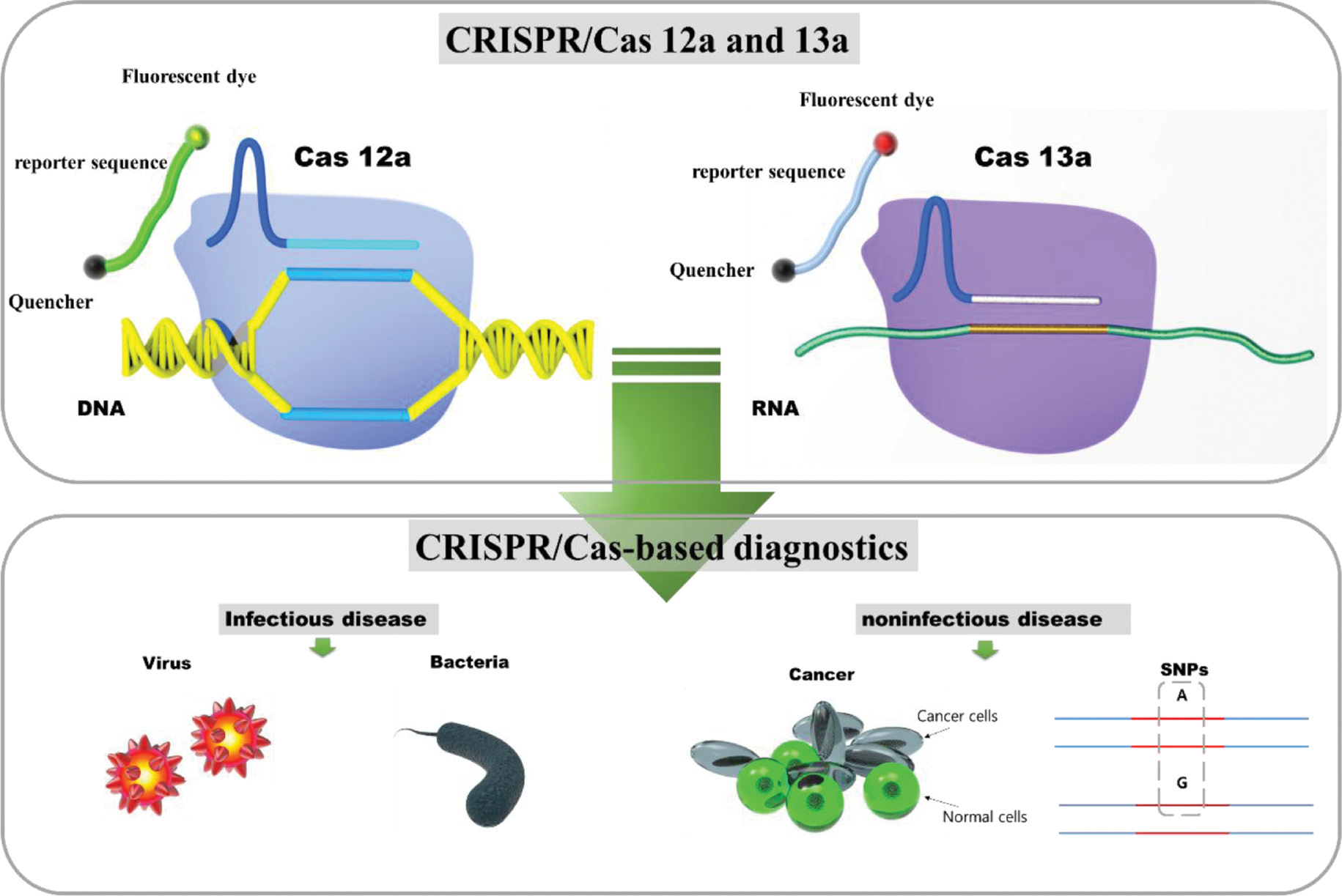
Accurate and rapid detection of infectious disease is crucial for managing clinical care and controlling the spread of disease in the community [35]. Cas proteins brought about radical changes in gene editing for inexpensive, accurate, rapid testing for POC [35]. Type II, V, and VI Cas proteins have often been utilized for the CRISPR systems diagnostic purpose [16]. Although various robust biosensing strategies had utilized CRISPR-Cas9, the two recently discovered Cas proteins, CRISPR-Cas13a and CRISPR-Cas12a, significantly advanced the technique to detect nucleic acid [16, 36]. Cas-13a utilizes the Cas13-crRNA to guide and bind the target RNA. Once it binds to the target, the nonspecific RNase activity of Cas 13 is activated, which results in collateral cleavage of any nearby single-stranded RNAs without a complementary crRNA guide sequence [27, 28]. In contrast, Cas-12a is an RNA-guided, DNA-targeting enzyme that targets DNA and collaterally cleaves ssDNA [37]. Hence, CRISPR/Cas systems allow the detection of viral and bacterial-induced infectious diseases and noninfectious diseases [16]. Owning to the high specificity and accuracy of CRISPR/Cas, it has also been utilized in the discrimination of human single nucleotide polymorphism (SNP) sites [37] (Figure 2) Diagnostics based on CRISPR/Cas systems combined with isothermal nucleic acid amplification strategies can be detected using a lateral flow strip or a hand-held fluorometer with colorimetric or fluorescent reporters, respectively [38–40]. Very recently, the CRISPR/Cas-powered electrochemical biosensors have provided free pre-amplification approaches for nucleic acid diagnostics [41, 42]. Therefore, CRISPR/Cas systems shed light on rapid, robust, and cost-effective molecular diagnosis at POC.
Figure 2 Illustration of CRISPR-Cas-12a and 13a sensing mechanisms and their diagnostic applications.
The well-known CRISPR/Case-based diagnostic platform termed specific high-sensitivity enzymatic reporter unlocking (HERLOCK) established by Gootenberg et al. exploits Cas13a to detect RNA molecules [43]. After a year, a diagnostic platform based on SHERLOCK combined with isothermal pre-amplification was developed, which allows single molecules to detect RNA or DNA at an attomolar level [44]. It has been applied to detect dengue or Zika virus single-stranded RNA as well as mutations in patient liquid biopsy samples via a lateral flow, which provides a quantitative platform to detect the target RNA or DNA at POC [44]. Moreover, the SHERLOCK technology has been harnessed for the detection of HIV, which continuously arouses significant concern worldwide [45]. Notably, in vitro transcription of DNA to RNA is required before the test when using SHERLOCK [37].
Cas12a is another powerful enzyme that is widely used in CRISPR/Cas-based biosensing strategies. Doudna et al. reported that Cas-12a has single-stranded DNA threading activity. By combining Cas12a single-stranded deoxyribonuclease (ssDNase) activation with isothermal amplification, They created a technology termed DNA endonuclease-targeted CRISPR trans reporter (DETECTR). It achieved attomolar DNA detection sensitivity [46]. DETECTR enabled selective detection of human papillomavirus (HPV) in patient samples in a short time [46]. Similarly, Li et al. developed one-Hour Low-cost Multipurpose highly Efficient System (HOLMES) based on Cas-12a for fast human SNP genotype detection [37]. Recently, Yin et al. reported a dynamic aqueous multiphase reaction (DAMR) system for quantitative HPV detection with sensitivity based on CRISPR-Cas-12a and recombinase polymerase amplification (RPA) [38].
Notably, CRISPR/Cas has emerged as a sensing tool for rapidly developed severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) RNA for quick, on-site diagnosis in the current pandemic of coronavirus disease 2019 (COVID-19) [16, 47–49] (Figure 1) The current “gold-standard” COVID-19 detection method is reverse transcription polymerase chain reaction (RT-PCR) [50]. This method is highly sensitive and specific, and it can quantify viral RNA. However, it requires proper sampling procedures, and it suffers from PCR inhibition, and a long detection time [51, 52].
CRISPR-Cas12a and 13a combined with isothermal nucleic acid amplification strategies, such as RPA and loop-mediated isothermal amplification (LAMP), can provide sensitive detection results within an hour [48, 49, 53]. To improve the sample-to-result time, Ramachandran et al. developed an electric field gradient for automatically purifying target RNA in raw nasopharyngeal swab samples based on a selective ionic focusing technique, termed isotachophoresis (ITP) implemented on a microfluidic chip [53]. They combined the ITP technology with a LAMP CRISPR assay to detect SARS-CoV-2 RNA. It can be achieved in about 35 min from raw sample-to-result-time [53]. Additionally, CRISPR/Cas has been utilized for the multiplex detection of SARS-CoV-2 and HIV. Ding et al. developed an all-in-one dual CRISPR-Cas-12a (AIOD-CRISPR) assay for detecting SARS-CoV-2 and HIV in one-spot. They achieved sensitive detection of virus RNAs down to a few copies. It has achieved comparable results with real-time RT-PCR in human plasma samples [54]. Recently, Fozouni et al. reported a CRISPR-Cas13a-based direct SARS-CoV-2 RNA detection strategy, eliminating the viral RNA’s pre-amplification step. The authors measured the fluorescence by utilizing a smart phone camera in a compact device, which yields quantitative results in real-time. They achieved sensitivity of ∼100 copies/μL within 30 min for SARS-CoV-2 RNA detection [55]. This sensing platform has the potential to allow fast, cost-effective and accurate COVID diagnostics at POC [55].
Moreover, the CRISPR/Cas system has been exploited in the field of bacterial detection, such as for identifying antimicrobial drug-resistant bacteria [16]. One strategy termed finding low abundance sequences by hybridization (FLASH) employs Cas9 and different gRNAs to identify pathogens using gene sequencing [56], which achieved sub-attomolar sensitivity. Although the authors only applied this strategy to examine the antimicrobial resistance genes in respiratory fluid and dried blood spots, in theory, FLASH next-generation genes sequencing can be applied to other samples that rely on multiplex PCR [56].
Also, the CRISPT/Cas system has been utilized in the diagnosis of noninfectious diseases, such as identifying oncogenes and other mediators in almost all areas of cancers [57]. Additionally, CRISPR-based genome editing strategies show high-accuracy in the identification of single-base variation in target sequences. Therefore it provides a sufficient distinction of specific alleles, which is crucial for early disease diagnosis [58, 59].
CRISPR-Cas9-based gene therapy
Gene therapies, which deliver nucleic acids to patients’ cells, have been utilized to treat diseases that are not curable with conventional medicines [60]. Current gene therapies use nucleases to modify genomes by generating DNA lesions and initiating endogenous DNA repair systems [61].Conventional nucleases for gene therapy include zinc finger nucleases (ZFNs), and transcription activator-like effector nucleases (TALENs) [62]. ZFNs and TALENs are engineered enzymes that are made by fusing binding and cleavage domains. Substitution of the natural cleavage domain with engineered enzymes enables ZFNs and TALENs to target specific DNA sequences [63]. However, it is challenging to predict the target’s final arrangement by using ZENs [63]. DNA target cloning is a complicated design process and the low genome screening efficiency has limited the application of TALENs in gene therapy [64]. Recently, gene therapy based on CRISPR-Cas9 has been applied to treat various diseases in clinical trials due to its simplicity, various genomic target sites, multiplex genomic modification, and the possibility of easy off-target site prediction [65, 66]. Different from ZFNs and TALENs, the process of CRISPR-Cas9 is easier and faster because it only requires one Cas9 endonuclease and a short single gRNA [67]. In addition to this, unlike traditional methods that introduce functional genetic materials via direct viral vector-mediated over-expression, CRISPR-Cas9 can introduce functional genes in patients that can be expressed in their natural context [68]. Therefore, CRISPR/Cas therapy is promising for treating diseases related to gene disorders [22].
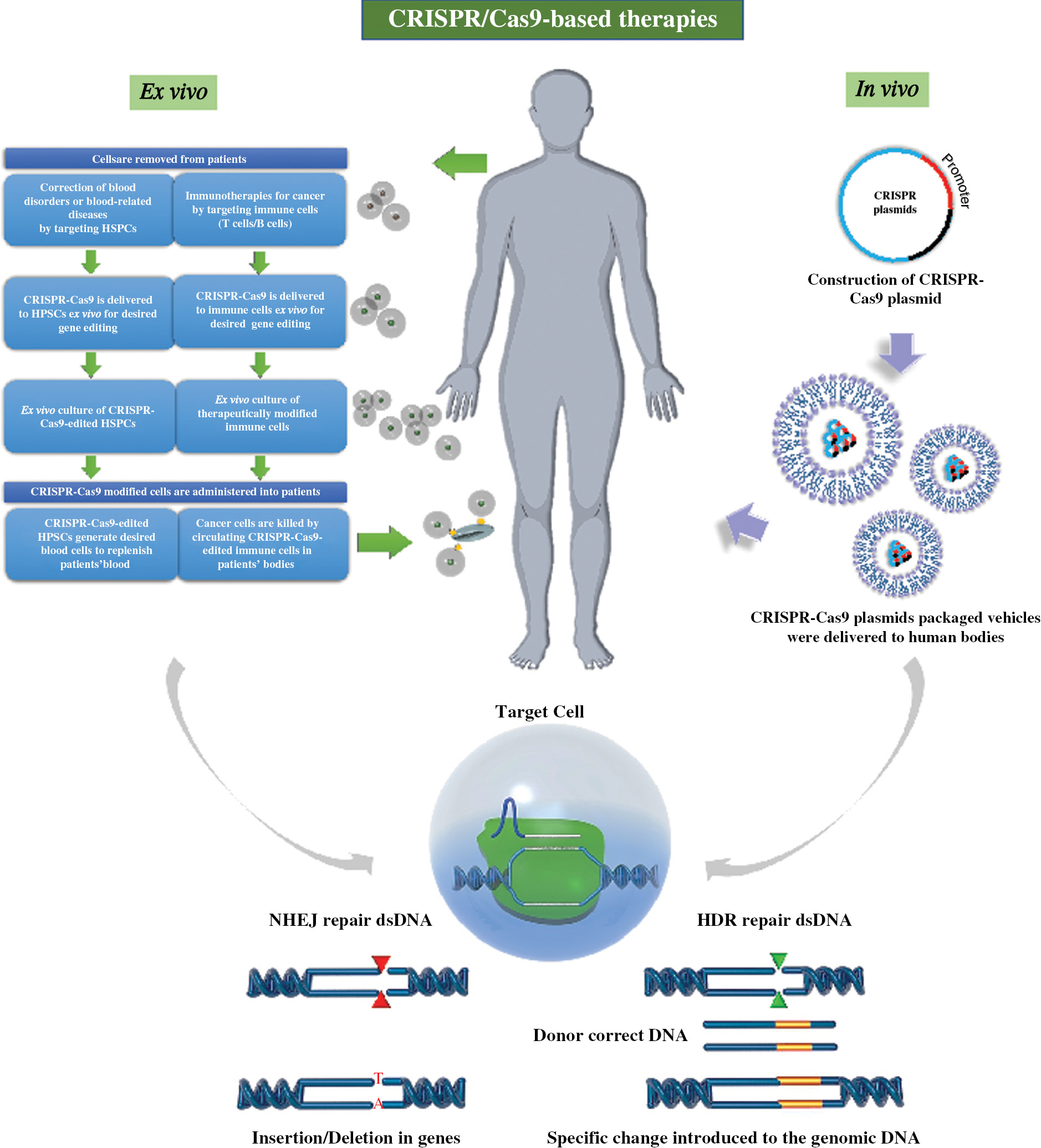
Since its discovery, CRISPR/Cas-based therapy has been investigated in a wide spectrum of cells types and in various animals. In brief, CRISPR-Cas9-based gene-editing can be accomplished by inserting, deleting, or replacing sections of the DNA sequence in the DNA double-strand breaks DSBs’ locus. (Figure 3). In ex vivo CRISPR-Cas9 gene therapies, cells are isolated from the patient, applied to the gene-editing process, and then injected back into the patient. It is possible to choose the cells with high viability to increase the survival rate of cells after transplantation to achieve its therapeutic effect [69]. In terms of in vivo CRISPR-Cas9 gene therapies, CRISPR components are directly delivered to specific tissues (such as the eye, brain, or muscle) through systematic administration or local injection [69]. Subsequently, CRISPR-Cas9-based gene-editing systems recognize particular target DNA sequences by gRNA, generating DSBs using the cas9 protein, which then initiates endogenous DNA repair pathways. Nonhomologous end-joining (NHEJ) is the major DNA repair mechanism that induces indels to deactivate gene functions. Homology-directed repair (HDR) is essential for precise editing, such as replacing a gene sequence with a desired sequence with the low possibility of introducing random mutations [68, 70]. Ex vivo CRISPR-Cas9 gene therapies have been widely utilized to treat several blood diseases, acquired immunodeficiency syndrome (AIDS), and cancer. In vivo CRISPR-Cas9 gene therapy technology is only at its initial stage. To the best of our knowledge, it has only been tested to treat LCA10 [34]. Below, we present the up-to-date progress of CRISPR-Cas9-based gene therapy and its clinical applications according to specific diseases.
Figure 3 Illustration of CRISPR-Cas9-based gene therapy.
Ex vivo CRISPR-Cas9-based gene therapy for blood disorders
Initially CRISPR-Cas9 was applied for the treatment inherited blood disorders [71]. Sickle-cell disease (SCD) and β-thalassemia genetic disorders are caused by mutations in a single gene, which affect the production of β-globin, a component of fetal hemoglobin (HbF) [72, 73]. These two diseases can be alleviated by re-expressing the γ-globin gene, the paralogous gene of HbF. Wu et al. applied CRISPR-Cas9 to human hematopoietic stem cells to increase the expression of γ-globin by modifying the enhancer of a gene called B-cell lymphoma 11 A (BCL11A), which is a repressor gene of γ-globin [71].
Ex vivo CRISPR-Cas9-based gene therapy for AIDS
Another ex vivo CRISPR-Cas9 therapy approach has been applied to patients infected with HIV [74]. Continuous deduction of CD4+ T lymphocytes is the main symptom in HIV patients. This reduction leads to the inability of the immune system to defend against pathogens [75]. HIV can bind and enter macrophages or CD4+ T lymphocytes. Transmembrane C–C chemokine receptor type 5 (CCR5) plays an important role in mediating the entry of HIV into lymphocytes [76]. Previous studies showed that a specific mutation (the deletion of particular 32-base pair of the gene) of the CCR5 gene could destroy its function, thus suppressing the expression of CCR5 on the cell surface [77, 78]. Xu et al. applied CRISPR/Cas9 to ablate CCR5 with same strategy which has been utilized in hematopoietic stem and progenitor cells (HSPCs). The results of this study showed that these edited HSPCs in mice could resist HIV for up to 47 weeks [31].
Ex vivo CRISPR-Cas9-based gene therapy for cancer
Furthermore, CRISPR-Cas9-based ex vivo gene therapies combined with immunotherapy advanced the clinical trials for cancer treatment. Chimeric antigen receptor (CAR)-engineered T-cell therapy is the most commonly used immunotherapy that modifies T cells’ genes from patients to kill cancer cells [79]. Firstly, T lymphocytes were isolated from patients’ blood. Next, target genes (TCR and PD-1) were deleted to attenuate the immune response. Subsequently, a specific TCR encoding gene for the NY-ESO-1 antigen, which is highly up-regulated in the relapsed tumor, was transduced into the T lymphocytes using a lentivirus [33]. Therefore, immune cells can be transformed into trained “soldier cells” to locate and kill cancer cells through gene modification. This clinical trial paved the way for a broader implication of CRISPR/Cas-based gene therapies. A recent study published in Nature Medicine showed favorable results in the treatment of late-stage lung cancer patients with these modified T cells. Donor cells exist in patients for more than 19 months. This trial confirmed the feasibility and safety of CAR T-cell therapy in treating late-stage lung cancer [80]. The authors also used PD-1 deleted-T cells for the treatment of different kinds of cancers in clinical trials [80].
Recent advances in in vivo CRISPR/Cas-based gene therapy
Ex vivo CRISPR-Cas9 gene therapy has been extensively reviewed, however in vivo gene therapy based on CRISPR-Cas9 has been less extensively employed [80]. Risk-free and efficient delivery of the CRISPR-Cas9 element to the target tissue is the foundation of successful treatment [81]. Currently, adeno-associated viral (AAV) vectors are the frequently used delivery vehicles for in vivo CRISPR/Cas-based gene therapy [82]. CRISPR-Cas9-based in vivo gene therapy that employs AAV vectors has been applied in treating Duchenne muscular dystrophy (DMD) in mouse models through intramuscular, intraperitoneal, or intravenous injection to rescue the disease phenotype [83–86]. These studies showed the potential of using AAV vectors for in vivo CRISPR-Cas9 delivery in human clinical trials. Recently, for the very first time, THE CRISPR-Cas9 technique was employed in a clinical trial through direct delivery of the CRISPR/Cas9 component to the target tissue of human body by using AAV vectors. This trial is aiming to treat LCA10 by correcting a defective gene named centrosomal protein 290 (CEP290) [34].The outcome of this clinical trial is of great importance for the further applications of in vivo CRISPR/Cas gene therapies.
Future perspective and challenges of CRISPR/Cas-based diagnostics and therapeutic applications
CRISPR/Cas technology has been regarded as a versatile genome-editing tool with significant potential for diagnostic and therapeutic applications. CRISPR/Cas-based molecular assays provide fast, sensitive, and specific diagnostics in patient samples suitable for POCT. Moreover, CRISPR/Cas-based diagnostics can be easily transformed into a universal diagnostic platform by simply changing the design of the specific sgRNA for different pathogens and cancer cell lines. CRISPR/Cas-based SARS-CoV-2 detection strategies for POCT have rapidly emerged since the outbreak of the COVID-19 pandemic due to the simplicity of its design. Despite the high sensitivity and specificity, CRISPR/Cas-based fluorescent nucleic acid detection has several limitations, including the necessity of pre-amplification methods, such as LAMP and RPA, expensive fluorescent signal read-out devices, and expertise in biotechnology. On the other hand, although CRISPR/Cas-based lateral flow colorimetric detection provides merely qualitative detection results, it has been regarded as a rapid, robust, and cost-effective approach for on-site diagnosis by the naked-eye. Moreover, the CRISPR/Cas-based electrochemical biosensors also provide a highly desirable free pre-amplification method for rapid nucleic acid diagnostics.
In terms of therapeutic applications, despite the remarkable advances of CRISPR/Cas-based gene therapies and their clinical applications, several challenges should be addressed. A significant concern of utilizing CRISPR-Cas9 gene therapy in the clinic is the OTEs. The binding of gRNA to a similar region with target sequence of genome can lead to the deletions or insertions in an unwanted locus. Notably, these random deletions and insertions of the genome can cause detrimental mutations [87]. Since undetectable OTEs are not understood completely, the method for the detection and evaluation of the consequences of OTEs for various organs needs to be explored further [88].
In addition to OTEs, different delivery methods of CRISPR/Cas gene therapy have their own merits and challenges. Compared with in vivo delivery, ex vivo delivery exhibits several advantages, including technical feasibility, clinical safety, and the possibility of a tight quality check on transplanted cells. But this approach was challenged by the viability and time limitation of retaining functional cells in vitro after genetic editing and a lengthy culture process [80]. Moreover, ex vivo therapy is limited to specific cell types, such as HSPCs and T lymphocytes, because of their ability to survive and expand in culture. In contrast, many other tissues are not suited for this method [89, 90]. In vivo gene therapy is a promising technology that can expand CRISPR/Cas technology application to treat genetic disorders with a widespread range. Methods of in vivo CRISPR components delivery include intravenously injecting to specific tissues. Compared with local injections, systematic administration is more capable of reaching different organs through the circulatory system. Besides, it is also possible to control the expression of Cas9 precisely in desired tissue [91]. However, CRISPR components delivered in vivo might be degraded by various enzymes and thus lose their function [80].
Besides the technical challenges, several ethical (moral) and safety concerns limited the CRISPR applications, such as germline editing [91, 93]. Firstly, OTEs over the lifetime caused by germline gene editing remains critical for utilizing this technology broadly in the human body. Also, the possibility of artificial selection of humans has raised concern for ethicists and the public. Therefore, a good documentation guideline will be of great help to support the further clinical applications of CRISPR-Cas9 technology.
References
- Wright AV, Nuñez JK, Doudna JAJC. Biology and applications of CRISPR systems: harnessing nature’s toolbox for genome engineering. Cell 2016;164:29-44. [PMID: 26771484 DOI: 10.1016/j.cell.2015.12.035]
- Pickar-Oliver A, Gersbach CA. The next generation of CRISPR-Cas technologies and applications. Nat Rev Mol Cell Biol 2019;20:490-507. [PMID: 31147612 DOI: 10.1038/s41580-019-0131-5]
- Loureiro A, da Silva GJ. Crispr-cas: converting a bacterial defence mechanism into a state-of-the-art genetic manipulation tool. Antibiotics 2019;8:18. [PMID: 30823430 DOI: 10.3390/antibiotics8010018]
- Sarewitz D. CRISPR: science cant solve it. Nature 2015;522:413-14. [PMID: 26108836 DOI: 10.1038/522413a]
- Liu X, Homma A, Sayadi J, Yang S, Ohashi J, et al. Sequence features associated with the cleavage efficiency of CRISPR/Cas9 system. Sci Rep 2016;6:1-9. [PMID: 26813419 DOI: 10.1038/srep19675]
- Adli MJ. The CRISPR tool kit for genome editing and beyond. Nat Commun 2018;9:1-13. [PMID: 29765029 DOI: 10.1038/s41467-018-04252-2]
- Wu SS, Li QC, Yin CQ, Xue W, Song CQ. Advances in CRISPR/Cas-based gene therapy in human genetic diseases. Theranostics 2020;10:4374-82. [PMID: 32292501 DOI: 10.7150/thno.43360]
- Lau A, Ren CL, Lee LP. Critical review on where CRISPR meets molecular diagnostics. Prog Biomed Eng 2020;3. [DOI: 10.1088/2516-1091/abbf5e]
- Srivastava S, Upadhyay DJ, Srivastava A. Next-generation molecular diagnostics development by CRISPR/Cas tool: rapid detection and surveillance of viral disease outbreaks. Front Mol Biosci 2020;7:582499. [PMID: 33425987 DOI: 10.3389/fmolb.2020.582499]
- Afzal S, Sirohi P, Singh NK. A review of CRISPR associated genome engineering: application, advances and future prospects of genome targeting tool for crop improvement. Biotechnol Lett 2020;42:1611-32. [PMID: 32642978 DOI: 10.1007/s10529-020-02950-w]
- Moon SB, Ko J-H, Kim Y-S. Recent advances in the CRISPR genome editing tool set. Exp Mol Med 2019;51:1-11. [PMID: 31685795 DOI: 10.1038/s12276-019-0339-7]
- Ding W, Zhang Y, Shi S. Development and application of CRISPR/Cas in microbial biotechnology. Front Bioeng Biotechnol 2020;8:711. [PMID: 32695770 DOI: 10.3389/fbioe.2020.00711]
- Wu Y, Liu Y, Lv X, Li J, Du G, et al. Applications of CRISPR in a microbial cell factory: from genome reconstruction to metabolic network reprogramming. ACS Synthetic Biol 2020;9:2228-38. [PMID: 32794766 DOI: 10.1021/acssynbio.0c00349]
- Ishino Y, Krupovic M, Forterre P. History of CRISPR-Cas from encounter with a mysterious repeated sequence to genome editing technology. J Bacteriol Res 2018;200:e00580-17. [PMID: 29358495 DOI: 10.1128/JB.00580-17]
- Jia F, Li X, Zhang C, Tang X. The expanded development and application of CRISPR system for sensitive nucleotide detection. Protein Cell 2020;11:1-6. [PMID: 32246439 DOI: 10.1007/s13238-020-00708-8]
- Katalani C, Booneh HA, Hajizade A, Sijercic A, Ahmadian G. CRISPR-based diagnosis of infectious and noninfectious diseases. Biol Proced Online 2020;22:1-14. [PMID: 32939188 DOI: 10.1186/s12575-020-00135-3]
- Hermans P, Van Soolingen D, Bik E, De Haas P, et al. Insertion element IS987 from Mycobacterium bovis BCG is located in a hot-spot integration region for insertion elements in Mycobacterium tuberculosis complex strains. Infect Immun 1991;59:2695-705. [PMID: 1649798 DOI: 10.1128/IAI.59.8.2695-2705.1991]
- Mojica FJ, Juez G, Rodriguez-Valera F. Transcription at different salinities of Haloferax mediterranei sequences adjacent to partially modified PstI sites. Mol Microbiol 1993;9:613-21. [PMID: 8412707 DOI: 10.1111/j.1365-2958.1993.tb01721.x]
- Mojica FJ, Montoliu L. On the origin of CRISPR-Cas technology: from prokaryotes to mammals. Trends Microbiol 2016;24:811-20. [PMID: 27401123 DOI: 10.1016/j.tim.2016.06.005]
- Xu Y, Li Z. CRISPR-Cas systems: overview, innovations and applications in human disease research and gene therapy. Comput Struct Biotechnol J 2020;18:2401-15. [PMID: 33005303 DOI: 10.1016/j.csbj.2020.08.031]
- Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, et al. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 2012;337:816-21. [PMID: 22745249 DOI: 10.1126/science.1225829]
- Foss DV, Hochstrasser ML, Wilson RC. Clinical applications of CRISPR-based genome editing and diagnostics. Transfusion 2019;59:1389-99. [PMID: 30600536 DOI: 10.1111/trf.15126]
- Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 2015;163:759-71. [PMID: 26422227 DOI: 10.1016/j.cell.2015.09.038]
- Shmakov S, Abudayyeh OO, Makarova KS, Wolf YI, Gootenberg JS, et al. Discovery and functional characterization of diverse class 2 CRISPR-Cas systems. Mol Cell 2015;60:385-97. [PMID: 26593719 DOI: 10.1016/j.molcel.2015.10.008]
- Tang Y, Fu Y. Class 2 CRISPR/Cas: an expanding biotechnology toolbox for and beyond genome editing. Cell Biosci 2018;8:1-13. [PMID: 30459943 DOI: 10.1186/s13578-018-0255-x]
- Travis J. CRISPR genome-editing technology shows its power. Science 2015;350:1456-7. [DOI: 10.1126/science.350.6267.1456].
- Abudayyeh OO, Gootenberg JS, Konermann S, Joung J, Slaymaker IM, et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 2016;353:aaf5573. [PMID: 27256883 DOI: 10.1126/science.aaf5573]
- East-Seletsky A, O’Connell MR, Knight SC, Burstein D, Cate JH, et al. Two distinct RNase activities of CRISPR-C2c2 enable guide-RNA processing and RNA detection. Nature 2016;538:270-3. [PMID: 27669025 DOI: 10.1038/nature19802]
- Cong L, Ran FA, Cox D, Lin S, Barretto R, et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013;339:819-23. [PMID: 23287718 DOI: 10.1126/science.1231143]
- Hirakawa MP, Krishnakumar R, Timlin JA, Carney JP, Butler KS. Gene editing and CRISPR in the clinic: current and future perspectives. Biosci Rep 2020;40. [PMID: 32207531 DOI: 10.1042/BSR20200127]
- Xu L, Yang H, Gao Y, Chen Z, Xie L, et al. CRISPR/Cas9-mediated CCR5 ablation in human hematopoietic stem/progenitor cells confers HIV-1 resistance in vivo. Mol Ther 2017;25:1782-9. [PMID: 28527722 DOI: 10.1016/j.ymthe.2017.04.027]
- Azangou-Khyavy M, Ghasemi M, Khanali J, Boroomand-Saboor M, Jamalkhah M, et al. CRISPR/Cas: from tumor gene editing to T cell-based immunotherapy of cancer. Front Immunol 2020;11. [PMID: 33117331 DOI: 10.3389/fimmu.2020.02062]
- Stadtmauer EA, Fraietta JA, Davis MM, Cohen AD, Weber KL, et al. CRISPR-engineered T cells in patients with refractory cancer. Science 2020;367. [PMID: 32029687 DOI: 10.1126/science.aba7365]
- Ledford H. CRISPR treatment inserted directly into the body for first time. Nature 2020;579:185. [PMID: 32157225 DOI: 10.1038/d41586-020-00655-8]
- Chertow DS. Next-generation diagnostics with CRISPR. Science 2018;360:381-2. [PMID: 29700254 DOI: 10.1126/science.aat4982]
- Myhrvold C, Freije CA, Gootenberg JS, Abudayyeh OO, Metsky HC, et al. Field-deployable viral diagnostics using CRISPR-Cas13. Science 2018;360:444-8. [PMID: 29700266 DOI: 10.1126/science.aas8836]
- Li S-Y, Cheng Q-X, Wang J-M, Li X-Y, Zhang Z-L, et al. CRISPR-Cas12a-assisted nucleic acid detection. Cell Discov 2018;4:1-4. [PMID: 29707234 DOI: 10.1038/s41421-018-0028-z]
- Yin K, Ding X, Li Z, Zhao H, Cooper K, et al. Dynamic aqueous multiphase reaction system for one-pot CRISPR-Cas12a based ultrasensitive and quantitative molecular diagnosis. Anal Chem 2020;92:8561-8. [PMID: 32390420 DOI: 10.1021/acs.analchem.0c01459]
- Huang Z, Tian D, Liu Y, Lin Z, Lyon C, et al. Ultra-sensitive and high-throughput CRISPR-powered COVID-19 diagnosis. Biosens Bioelectron 2020;164:112316. [PMID: 32553350 DOI: 10.1016/j.bios.2020.112316]
- Mukama O, Wu J, Li Z, Liang Q, Yi Z, et al. An ultrasensitive and specific point-of-care CRISPR/Cas12 based lateral flow biosensor for the rapid detection of nucleic acids. Biosens Bioelectron 2020;159:112143. [PMID: 32364943 DOI: 10.1016/j.bios.2020.112143]
- Bruch R, Baaske J, Chatelle C, Meirich M, Madlener S, et al. CRISPR/Cas13a-powered electrochemical microfluidic biosensor for nucleic acid amplification-free miRNA diagnostics. Adv Mater 2019;31:1905311. [PMID: 31663165 DOI: 10.1002/adma.201905311]
- Xu W, Jin T, Dai Y, Liu CC. Surpassing the detection limit and accuracy of the electrochemical DNA sensor through the application of CRISPR Cas systems. Biosens Bioelectron 2020;155:112100. [PMID: 32090878 DOI: 10.1016/j.bios.2020.112100]
- Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 2017;356:438-42. [PMID: 28408723 DOI: 10.1126/science.aam9321]
- Gootenberg JS, Abudayyeh OO, Kellner MJ, Joung J Collins JJ, et al. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 2018;360:439-44. [PMID: 29449508 DOI: 10.1126/science.aaq0179]
- Bhattacharyya RP, Thakku SG, Hung DT. Harnessing CRISPR effectors for infectious disease diagnostics. ACS Infectious Dis 2018;4:1278-82. [PMID: 30113801 DOI: 10.1021/acsinfecdis.8b00170]
- Chen JS, Ma E, Harrington LB, Da Costa M, Tian X, et al. CRISPR-Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 2018;360:436-9. [PMID: 29449511 DOI: 10.1126/science.aar6245]
- Hou T, Zeng W, Yang M, Chen W, Ren L, et al. Development and evaluation of a CRISPR-based diagnostic for 2019-novel coronavirus. PLoS Pathog 2020;16:e1008705. [PMID: 32853291 DOI: 10.1371/journal.ppat.1008705]
- Metsky HC, Freije CA, Kosoko-Thoroddsen T-SF, Sabeti PC, Myhrvold CJB, et al. CRISPR-based surveillance for COVID-19 using genomically-comprehensive machine learning design. bioRxiv 2020. [DOI: 10.1101/2020.02.26.967026]
- Esbin MN, Whitney ON, Chong S, Maurer A, Darzacq X, et al. Overcoming the bottleneck to widespread testing: a rapid review of nucleic acid testing approaches for COVID-19 detection. RNA 2020;26:771-83. [PMID: 32358057 DOI: 10.1261/rna.076232.120]
- Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill 2020;25. [PMID: 31992387 DOI: 10.2807/1560-7917.ES.2020.25.3.2000045]
- Tahamtan A, Ardebili A. Real-time RT-PCR in COVID-19 detection: issues affecting the results. Expert Rev Mol Diagn 2020;20:453-4. [PMID: 32297805 DOI: 10.1080/14737159.2020.1757437]
- Wang X, Yao H, Xu X, Zhang P, Zhang M, et al. Limits of detection of 6 approved RT–PCR kits for the novel SARS-Coronavirus-2 (SARS-CoV-2). Clin Chem 2020;66:977-9. [PMID: 32282874 DOI: 10.1093/clinchem/hvaa099]
- Ramachandran A, Huyke DA, Sharma E, Sahoo MK, Huang C, et al. Electric field-driven microfluidics for rapid CRISPR-based diagnostics and its application to detection of SARS-CoV-2. Proc Nat Acad Sci U S A 2020;117:29518-25. [PMID: 33148808 DOI: 10.1073/pnas.2010254117]
- Ding X, Yin K, Li Z, Liu CJ. All-in-one dual CRISPR-Cas12a (AIOD-CRISPR) assay: a case for rapid, ultrasensitive and visual detection of novel coronavirus SARS-CoV-2 and HIV virus. bioRxiv 2020. [PMID: 32511323 DOI: 10.1101/2020.03.19.998724]
- Fozouni P, Son S, de León Derby MD, Knott GJ, Gray CN, et al. Direct detection of SARS-CoV-2 using CRISPR-Cas13a and a mobile phone. Cell 2020;184:323-33.e9. [PMID: 33306959 DOI: 10.1016/j.cell.2020.12.001]
- Quan J, Langelier C, Kuchta A, Batson J, Teyssier N, et al. FLASH: a next-generation CRISPR diagnostic for multiplexed detection of antimicrobial resistance sequences. Nucleic Acids Res 2019;47:e83. [PMID: 31114866 DOI: 10.1093/nar/gkz418]
- Tian X, Gu T, Patel S, Bode AM, Lee M-H, et al. CRISPR/Cas9–An evolving biological tool kit for cancer biology and oncology. NPJ Precision Oncol 2019;3:1-8. [PMID: 30911676 DOI: 10.1038/s41698-019-0080-7]
- Lee SH, Park Y-H, Jin YB, Kim S-U, Hur JK. CRISPR diagnosis and therapeutics with single base pair precision. Trends Molec Med 2020;26:337-50. [PMID: 31791730 DOI: 10.1016/j.molmed.2019.09.008]
- Courtney D, Moore J, Atkinson S, Maurizi E, Allen E, et al. CRISPR/Cas9 DNA cleavage at SNP-derived PAM enables both in vitro and in vivo KRT12 mutation-specific targeting. Gene Ther 2016;23:108-12. [PMID: 26289666 DOI: 10.1038/gt.2015.82]
- Kumar SR, Markusic DM, Biswas M, High KA, Herzog RW. Clinical development of gene therapy: results and lessons from recent successes. Mol Ther Methods Clin Dev 2016;3:16034. [PMID: 27257611 DOI: 10.1038/mtm.2016.34]
- Bulaklak K, Gersbach CA. The once and future gene therapy. Nat Commun 2020;11:1-4. [PMID: 33199717 DOI: 10.1038/s41467-020-19505-2]
- Maeder ML, Gersbach CA. Genome-editing technologies for gene and cell therapy. Mol Ther 2016;24:430-46. [PMID: 26755333 DOI: 10.1038/mt.2016.10]
- Cathomen T, Joung JK. Zinc-finger nucleases: the next generation emerges. Mol Ther 2008;16:1200-7. [PMID: 18545224 DOI: 10.1038/mt.2008.114]
- Juillerat A, Dubois G, Valton J, Thomas S, Stella S, et al. Comprehensive analysis of the specificity of transcription activator-like effector nucleases. Nucleic Acids Res 2014;42:5390-402. [PMID: 24569350 DOI: 10.1093/nar/gku155]
- Gupta RM, Musunuru K. Expanding the genetic editing tool kit: ZFNs, TALENs, and CRISPR-Cas9. J Clin Invest 2014;124:4154-61. [PMID: 25271723 DOI: 10.1172/JCI72992]
- Dai W-J, Zhu L-Y, Yan Z-Y, Xu Y, Wang Q-L, et al. CRISPR-Cas9 for in vivo gene therapy: promise and hurdles. Mol Therapy-Nucleic Acids 2016;5:e349. [PMID: 28131272 DOI: 10.1038/mtna.2016.58]
- Doudna JA, Charpentier E. The new frontier of genome engineering with CRISPR-Cas9. Science 2014;346. [PMID: 25430774 DOI: 10.1126/science.1258096]
- Hsu PD, Lander ES, Zhang F. Development and applications of CRISPR-Cas9 for genome engineering. Cell 2014;157:1262-78. [PMID: 24906146 DOI: 10.1016/j.cell.2014.05.010]
- Li H, Yang Y, Hong W, Huang M, Wu M, et al. Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther 2020;5:1-23. [PMID: 32296011 DOI: 10.1038/s41392-019-0089-y]
- Jasin M, Haber JE. The democratization of gene editing: insights from site-specific cleavage and double-strand break repair. DNA Repair 2016;44:6-16. [PMID: 27261202 DOI: 10.1016/j.dnarep.2016.05.001]
- Wu Y, Zeng J, Roscoe BP, Liu P, Yao Q, et al. Highly efficient therapeutic gene editing of human hematopoietic stem cells. Nat Med 2019;25:776-83. [PMID: 30911135 DOI: 10.1038/s41591-019-0401-y]
- Heywood JD, Karon M, Weissman S. Amino acids: incorporation into α-and β-chains of hemoglobin by normal and thalassemic reticulocytes. Science 1964;146:530-1. [PMID: 14190243 DOI: 10.1126/science.146.3643.530]
- Telen MJ, Malik P, Vercellotti GM. Therapeutic strategies for sickle cell disease: towards a multi-agent approach. Nat Rev Drug Discov 2019;18:139-58. [PMID: 30514970 DOI: 10.1038/s41573-018-0003-2]
- Johnson D. Ryan White dies of AIDS at 18; his struggle helped pierce myths. New York Times. 1990;D10.
- Ekere EF, Useh MF, Okoroiwu HU, Mirabeau TY. Cysteine-cysteine chemokine receptor 5 (CCR5) profile of HIV-infected subjects attending University of Calabar Teaching Hospital, Calabar, Southern Nigeria. BMC Infect Dis 2020;20:1-9. [PMID: 31900106 DOI: 10.1186/s12879-019-4737-1]
- Connor RI, Ho DD. Human immunodeficiency virus type 1 variants with increased replicative capacity develop during the asymptomatic stage before disease progression. J Virol 1994;68:4400-8. [PMID: 7515972 DOI: 10.1128/JVI.68.7.4400-4408.1994]
- Hütter G, Nowak D, Mossner M, Ganepola S, Müßig A, et al. Long-term control of HIV by CCR5 Delta32/Delta32 stem-cell transplantation. N Engl J Med 2009;360:692-8. [PMID: 19213682 DOI: 10.1056/NEJMoa0802905]
- Barmania F, Pepper MS. CC chemokine receptor type five (CCR5): an emerging target for the control of HIV infection. Appl Transl Genom 2013;2:3-16. [PMID: 27942440 DOI: 10.1016/j.atg.2013.05.004]
- Miliotou AN, Papadopoulou LC. CAR T-cell therapy: a new era in cancer immunotherapy. Curr Pharm Biotechnol 2018;19:5-18. [PMID: 29667553 DOI: 10.2174/1389201019666180418095526]
- Uddin F, Rudin CM, Sen T. CRISPR gene therapy: applications, limitations, and implications for the future. Front Oncol 2020;10. [PMID: 32850447 DOI: 10.3389/fonc.2020.01387]
- Wilbie D, Walther J, Mastrobattista E. Delivery aspects of CRISPR/Cas for in vivo genome editing. Acc Chem Res 2019;52:1555-64. [PMID: 31099553 DOI: 10.1021/acs.accounts.9b00106]
- Xu X, Wan T, Xin H, Li D, Pan H, et al. Delivery of CRISPR/Cas9 for therapeutic genome editing. J Gene Med 2019;21:e3107. [PMID: 31237055 DOI: 10.1002/jgm.3107]
- Nelson CE, Hakim CH, Ousterout DG, Thakore PI, Moreb EA, et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 2016;351:403-7. [PMID: 26721684 DOI: 10.1126/science.aad5143]
- Yin H, Song C-Q, Dorkin JR, Zhu LJ, Li Y, et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat Biotechnol 2016;34:328-33. [PMID: 26829318 DOI: 10.1038/nbt.3471]
- Yang Y, Wang L, Bell P, McMenamin D, He Z, et al. A dual AAV system enables the Cas9-mediated correction of a metabolic liver disease in newborn mice. Nat Biotechnol 2016;34:334-8. [PMID: 26829317 DOI: 10.1038/nbt.3469]
- Tabebordbar M, Zhu K, Cheng JK, Chew WL, Widrick JJ, et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 2016;351:407-11. [PMID: 26721686 DOI: 10.1126/science.aad5177]
- Schacker M, Seimetz D. From fiction to science: clinical potentials and regulatory considerations of gene editing. Clin Transl Med 2019;8:27. [PMID: 31637541 DOI: 10.1186/s40169-019-0244-7]
- Wang H-X, Li M, Lee CM, Chakraborty S, Kim H-W, et al. CRISPR/Cas9-based genome editing for disease modeling and therapy: challenges and opportunities for nonviral delivery. Chemical Rev 2017;117:9874-906. [PMID: 28640612 DOI: 10.1021/acs.chemrev.6b00799]
- Yu K-R, Natanson H, Dunbar CE. Gene editing of human hematopoietic stem and progenitor cells: promise and potential hurdles. Hum Gene Ther 2016;27:729-40. [PMID: 27483988 DOI: 10.1089/hum.2016.107]
- Baylis F, McLeod M. First-in-human phase 1 CRISPR gene editing cancer trials: are we ready? Curr Gene Ther 2017;17:309-19. [PMID: 29173170 DOI: 10.2174/1566523217666171121165935]
- Ablain J, Zon L. Tissue-specific gene targeting using CRISPR/Cas9. Methods Cell Biol 2016;135:189-202. [PMID: 27443926 DOI: 10.1016/bs.mcb.2016.03.004]
- Scheufele DA, Xenos MA, Howell EL, Rose KM, Brossard D, et al. U.S. attitudes on human genome editing. Science 2017;357:553-4. [PMID: 28798120 DOI: 10.1126/science.aan3708]
- Brokowski C. Do CRISPR germline ethics statements cut it? CRISPR J 2018;1:115-25. [PMID: 31021208 DOI: 10.1089/crispr.2017.0024]