IL2RB and TUBA1B are Potential Targets of Music Therapy for Alzheimer’s Disease and Cognitive Impairment
1Kangwon National University, Gangwon-do, Korea
2Ningde Municipal Hospital, Fujian Key Laboratory of Toxicant and Drug Toxicology, Medical College, Ningde Normal University, Ningde, China
3The Third Hospital, Hebei Medical University, Hebei, China
4Huashan Hospital, Fudan University, Shanghai, China
5The Second Affiliated Hospital of Nanchang University, Nanchang, China
6Zherong Vocational and Technical School, Fujian Province, China
7Faculty of Chinese Medicine and State Key Laboratory of Quality Research in Chinese Medicines, Macau University of Science and Technology, Macau, Macau SAR, China
8Department of Neurosurgery, Department of Physiology and Pharmacology, Department of Neurosurgery and Anesthesiology, School of Medicine, Loma Linda University, Risley Hall, Room 219, 11041 Campus Street, Loma Linda, CA 92354, USA
9The Affiliated Hospital of Youjiang Medical University for Nationalities, Baise City, China
aThese authors contributed equally to this work.
*Correspondence to: Jianling Liu, Ningde Municipal Hospital, Fujian Key Laboratory of Toxicant and Drug Toxicology, Medical College, Ningde Normal University, Ningde, China; and Huashan Hospital, Fudan University, Shanghai, China. E-mail: 2571333932@qq.com; Chengwan Shen, Fujian Key Laboratory of Toxicant and Drug Toxicology, Medical College, Ningde Normal University, Ningde, China. E-mail: shchwan@sin.com; Haojun Shi, Faculty of Chinese Medicine and State Key Laboratory of Quality Research in Chinese Medicines, Macau University of Science and Technology, Macau, Macau SAR, China. E-mail: haojunhjs@163.com; John H. Zhang, Department of Neurosurgery, Department of Physiology and Pharmacology, Department of Neurosurgery and Anesthesiology, School of Medicine, Loma Linda University, Loma Linda, CA, 92354, USA. E-mail: johnzhang3910@yahoo.com. Bin Huang, Ph.D., The Affiliated Hospital of Youjiang Medical University for Nationalities, Baise City, China. E-mail: 905849580@qq.com; Maowei Chen, Ph.D., Kangwon National University, Gangwon-do, Korea. E-mail: muwi@kangwon.ac.kr; Yisheng Chen, Ph.D., Fujian Key Laboratory of Toxicant and Drug Toxicology, Medical College, Ningde Normal University, Ningde, China; and Huashan Hospital, Fudan University, Shanghai, China. E-mail: yschen21@m.fudan.edu.cn
Received: December 31 2024; Revised: April 30 2025; Accepted: June 25 2025; Published Online: September 2 2025.
Cite this paper:
Wu K, Liu J, Su Y et al. IL2RB and TUBA1B are Potential Targets of Music Therapy for Alzheimer’s Disease and Cognitive Impairment. BIO Integration 2025; 1–17.
DOI: 10.15212/bioi-2025-1002. Available at: https://bio-integration.org/
Download citation
© 2025 The Authors. This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See https://bio-integration.org/copyright-and-permissions/
Abstract
Objective: This study examined the complex interactions among differential gene expression, immune responses, therapy-associated genes in Alzheimer’s disease (AD), and cognitive impairment, with a distinct focus on the integration of music therapy. Our objective was to delineate the landscape of differentially expressed genes (DEGs), particularly the interconnections between gene expression changes and therapeutic interventions, including exercise and music therapy, to discover immune-associated therapeutic targets.
Methods: Through evaluation of the gene expression datasets GSE140831 and GSE48624, we identified DEGs and investigated their interactions with genes whose expression is induced by exercise and music therapy. Through protein-protein interaction analysis, gene set variation analysis, immune infiltration studies, and correlation analyses, we revealed the regulatory interactions and pathway enrichments. An mRNA-miRNA interaction network was constructed to elucidate regulatory mechanisms, and a drug-target interaction analysis was performed to discover potential therapeutic avenues.
Results: Our study revealed the complex organization of DEGs. In addition to identifying IL2RB and TUBA1B, we propose these genes as critical modulators in the context of exercise, music therapy, and AD. Our results indicated a substantial upregulation of pathways, such as glycolysis and TGF-beta signaling, in response to exercise, music therapy, and AD. These pathways revealed significant changes in gene expression compared to baseline conditions, highlighting their involvement in immune response modulation. The network of mRNA-miRNA interactions revealed key regulators of immune response and cognition. Our examination of drug and target interactions provided insights into potential mechanisms of action for disease modulation.
Conclusions: This research provided a comprehensive overview of DEGs in AD and cognitive impairment, and uniquely incorporated evaluation of music therapy. Our findings underscore the importance of IL2RB and TUBA1B as potential therapeutic targets; provide initial insights into the mechanisms of immune response; and suggest new pathways for targeted treatment development.
Keywords
Alzheimer’s disease, cognitive impairment, differential gene expression, drug-target interaction, immune response, mRNA-miRNA network, music therapy, therapeutic targets.
Background
Alzheimer’s disease (AD) remains a formidable public health challenge, whose incidence is rising in tandem with global population aging [1]. This disorder affects the brain and is caused by loss of neurons. Progressive impairment occurs in multiple cognitive domains and consequently poses risks of altered social behavior. This neurodegenerative disorder is associated with progressive memory loss, impaired executive function, and cognitive impairment, and affects the daily lives of both patients and their caregivers [2].
AD has been a major clinical research focus, and the growing body of literature has highlighted the importance of genetic factors in the etiology of this disease [3]. The unprecedented development of high-throughput sequencing technology and bioinformatics tools has ushered in an era of genetic studies. Analysis of differentially expressed genes (DEGs) has led to important advances in knowledge regarding AD [4]. Despite considerable evidence of CIRS’s pathophysiology, substantial evidence indicates potential roles of immunity in CIRS; immunity might therefore serve as a treatment target. A first step in the fight against AD is directing therapies against neuroimmune inflammation, primarily because immune mechanisms are essential for AD progression [5–8].
Exercise and music therapy are non-pharmacological interventions that might benefit patients diagnosed with AD. Given that telerehabilitation with transcranial direct current stimulation has been found to be effective in stroke rehabilitation, these treatments might also improve cognition and quality of life in patients with AD [9]. Among the non-pharmacological therapies for the treatment of neurological diseases, music therapy is frequently used. Many studies have shown that music therapy improves cognition and emotion in patients with AD [10, 11]. Music’s therapeutic value might arise from its ability to engage various cognitive faculties, including memory, attention, and emotion, all of which are often impaired in AD [12]. By enhancing musical skills in individuals, it becomes possible to focus on the impact of music on brain function and condition, which may help improve cognitive and emotional aspects in AD patients. [13, 14]. Researchers are beginning to understand how music therapy can help people with AD. Music activates a broad range of areas in the brain, several of which are affected in AD [15–17]. The activation of pathways related to glycolysis and TGF-beta signaling might enhance synaptic plasticity and promote the development of new neural connections, compensating for the neurodegeneration associated with AD [18]. Moreover, music therapy stimulates the release of growth factors by engaging emotional and cognitive processes, which helps improve neuroplasticity and supports brain health. [15, 19]. Including music therapy in a larger therapy system would provide patients with AD with more comprehensive, holistic AD care [20].
Despite notable advancements, an important knowledge gap persists regarding the interactions among DEGs, immune responses, and the genetics associated with therapeutic interventions (e.g., exercise and music therapy). To study the functions of complex tissues and microenvironments, single-cell sequencing and multi-omics analyses are increasingly being applied to gain insight into immune microenvironments and metabolic regulation [21]. Bioinformatics analysis of large-scale datasets, such as biomarker identification, signaling pathway exploration, and immune characteristic studies, has afforded novel insights into disease diagnosis [22, 23]. The advent of these technologies has broadened the utility of transcriptomics, metabolomics, and proteomics in disease diagnosis and treatment [24, 25]. This study was aimed at bridging the existing knowledge gap through multimodal examination of gene expression, immunology, and therapeutic studies. We combined biomedical evidence from diverse sources to investigate the roles of music and exercise in modifying the trajectory of AD. This study uniquely performed extensive analysis of the connections among music, exercise, and AD, to gain novel perspectives in exploiting molecular targets for therapeutic purposes. With the aim of improving targeted interventions that might markedly enhance the quality of life of patients with AD, this study provides insights into the molecular and immune status of AD. The findings expand clinical knowledge and practice regarding AD, and may help ease the burden of this condition worldwide.
Materials and methods
Data acquisition and identification of key genes
We extracted two datasets related to AD and musical intervention, GSE140831 and GSE48624, from the Gene Expression Omnibus (GEO) database at NCBI. The limma package [26] was used for differential gene expression analysis. Genes with a p-value less than 0.05 were classified as DEGs. Genes from the GeneCards database were associated with physical activity. We intersected DEGs from both the AD and musical intervention datasets with the exercise-associated gene sets to identify overlapping genes. In preprocessing, we used quality control measures to ensure that all samples were of high quality. This process involved determining the distributions of expression values, the amounts of background noise present, and the presence of outliers. Quantile normalization was used to normalize the raw expression values. Moreover, the ComBat method was used to remove batch effects. The cut-off criteria for DEG selection were chosen as p-value < 0.05 and false discovery rate < 0.05, to control for multiple testing.
Protein-protein interaction and gene set variation analysis enrichment studies
To describe the interaction networks of IL2RB and TUBA1B, important genes identified in our study, we used STRING to map the protein-protein interaction (PPI) network. The resulting networks were carefully visualized with the Cytoscape application to understand the interactions among essential proteins. For gene set enrichment analysis, we divided the dataset GSE140831 into high-expression and low-expression groups based on the expression levels of IL2RB and TUBA1B. The GSVA package from BMC Bioinformatics [27] was used to obtain enrichment scores for each sample, which were then evaluated for inter-group differences. We used a permutation test with 1000 permutations to determine whether the pathway enrichments were significant. The false discovery rate significance threshold was set at 0.05.
Assessment of immune infiltration
Research on immune cellular infiltration in the GSE140831 dataset was conducted with the TIMER internet tool, which facilitated comprehensive assessment of immune cell infiltration across samples. The correlation between the expression of the identified essential genes (IL2RB and TUBA1B) and the abundance of various immune cells was quantitatively assessed with Spearman’s rank correlation analysis. Furthermore, the immune landscape of the samples was quantified with the ESTIMATE algorithm, which provided immune and stromal scores along with an average ESTIMATE score for each sample. This process was instrumental in gauging the composition of the tumor microenvironment. The correlation between those scores and the expression levels of key genes was tested with Pearson’s correlation coefficient, which enabled a nuanced exploration of how genes might influence or be influenced by the tumor’s immune and stromal components.
Development of an mRNA-miRNA interaction network
To elucidate the regulatory frameworks governing the observed gene expression patterns, specifically in the context of music therapy, we obtained statistics from the GSE115071 dataset in the GEO database from NCBI. Differential expression evaluation was performed with the limma package, as previously described [26], with a preset of p < 0.05 to distinguish DEGs. Concurrently, we used the miRTarBase database to source microRNAs (miRNAs) associated with the genes in the PPI network under consideration. The integration of DEGs and the comparison of miRNAs were facilitated by Venn diagrams identifying overlaps between miRNAs in the database and those associated with gene expression patterns observed in the music therapy intervention dataset.
Differential expression and improvement investigation methodology
The evaluation of gene expression changes between disease-affected and control groups in the context of PPI systems was conducted with t-tests executed in R. This measurable approach facilitated the identification of DEGs across comparative cohorts. To further investigate the regulatory mechanisms suggested by the mRNA-miRNA network, we performed enrichment analysis with the clusterProfiler package, as previously described [28]. Correlation analyses were performed with Pearson’s correlation method to determine the strength and direction of the associations between genes of interest. A p-value threshold of < 0.05 was used as the criterion for statistical significance, to ensure that the observed associations were both significant and robust.
Bayesian network inference
For Bayesian network (BN) inference, we used the CBNplot R package on the EA results with curated natural pathway data [29]. This package allowed us to determine the structure of the BN and provide a visualization enabling assessment of interactions between genes and the design of a probabilistic model of the regulatory network. The expression values of eigengenes for pathways identified in the EA served as the basis for BN inference.
Probabilistic reasoning and classification
Probabilistic reasoning was applied to the constructed BN to evaluate the conditional probabilities of gene expression, such as that of the matrix metalloproteinase-2 (MMP-2) gene, given certain clinical factors (e.g., tumor stage). This aim was achieved with the cpdist function in bnlearn, which enables the examination of conditional distributions and visualization of the results with the ggdist package [29].
Network comparison and stability assessment
The constructed BNs were compared against reference systems, and the stability of the systems was evaluated through bootstrap strategies. This process consisted of resampling the gene expression data, substituting the gene expression data, and recomputing the network structure to assess consistency. We additionally visualized the BN with the graphite package, which converts pathway topology to a molecular network [29]. The analyzed visualizations helped us reveal the relationships among genes and pathways, identify key genes for future work, and formulate new hypotheses for future validation.
Analysis of the shortest pathways between drug targets and disease-associated genes
Using previously developed methods, we sought to identify the shortest common pathways between drug targets and disease-associated genes. We used the Human Gene Interaction Network, in which the connections formed among genes are known biological interactions [30]. The rationale for using this method was that shorter network distances between a drug’s target genes and genes associated with a disease might suggest more favorable therapeutic potential for that drug against that disease. We refer to the network distance between drug target genes and disease-associated genes as their “proximity.” To examine the statistical significance of the proximity for each drug-disease gene pair, we conducted 1,000 random perturbation experiments within the international Human Gene Interaction Network. In those experiments, the positions of drug targets and disease-associated genes were randomly reassigned within the network to establish a baseline distribution of proximity measures.
Results
IL2RB and TUBA1B are key genes linking music therapy, exercise, and Alzheimer’s disease, according to integrated bioinformatics analysis
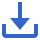
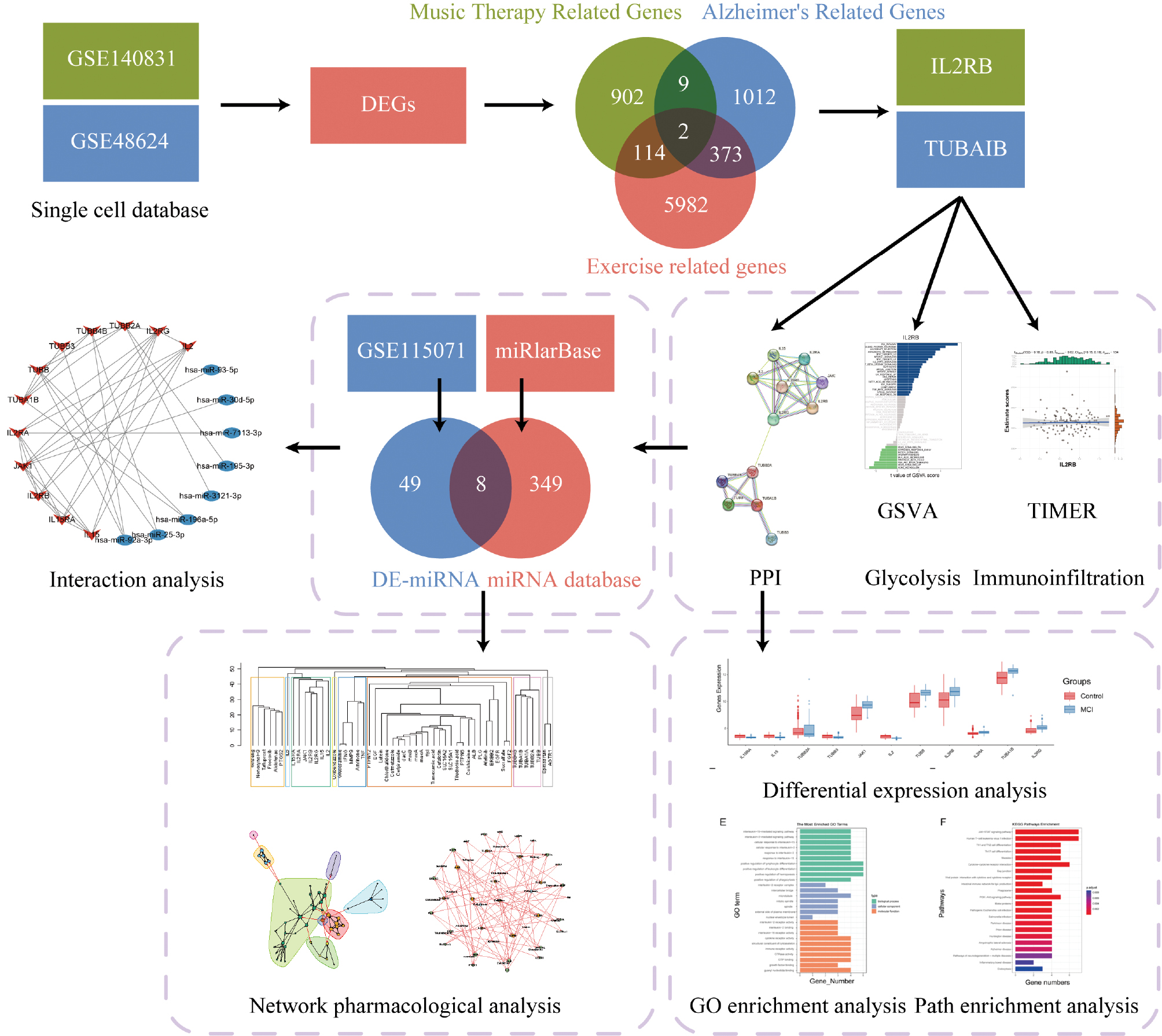
Through integrated bioinformatics analysis, we explored the relationships among music therapy, AD, and exercise-associated genes. Differential expression analysis of two single-cell RNA-seq datasets (GSE140831 and GSE48624) identified two core overlapping genes, IL2RB and TUBA1B (Figure 1), associated with both AD and exercise-responsive pathways. Further miRNA analysis with GSE115071 and miRTarBase revealed eight key DE-miRNAs potentially regulating IL2RB and TUBA1B. A gene-miRNA network confirmed interactions relevant to neurodegeneration and neuroprotection. PPI and functional enrichment analyses indicated that these genes play a role in the immune response and glycolysis pathways.Analyses with GSVA and TIMER demonstrated differential expression of these genes in AD samples and their potential roles in immune cell infiltration. GO and KEGG pathway analyses highlighted enrichment in metabolic regulation and immune signaling functions. Moreover, network pharmacology suggested IL2RB and TUBA1B as potential therapeutic targets.
Figure 1 This study presents an integrative analysis of music therapy, Alzheimer’s disease-associated genes, and exercise-associated genes. Differentially expressed genes (DEGs) were extracted from single-cell databases (GSE140831 and GSE48624), and an intersection analysis was performed with genes associated with music therapy, Alzheimer’s disease, and exercise, thus resulting in the identification of two core genes, IL2RB and TUBA1B. Additionally, differential expression analysis of miRNAs (DE-miRNAs) conducted with the GSE115071 and miRTarBase databases led to the identification of eight key miRNAs. A gene-miRNA interaction network was constructed, and protein-protein interaction (PPI) analysis was performed. Functional exploration with the GSVA and TIMER tools was conducted to investigate the roles of these genes in glycolysis and immune infiltration. Differential expression analysis revealed the expression differences between the control and Alzheimer’s disease groups, and GO functional enrichment and pathway analysis further elucidated their biological functions and pathways. Furthermore, network pharmacology analysis highlighted the potential applications of these genes as drug targets. Overall, this study systematically analyzed the relationships and potential mechanisms among music therapy, Alzheimer’s disease, and exercise-associated genes by integrating various bioinformatics approaches.
Differential gene expression and intersection with therapy-associated genes
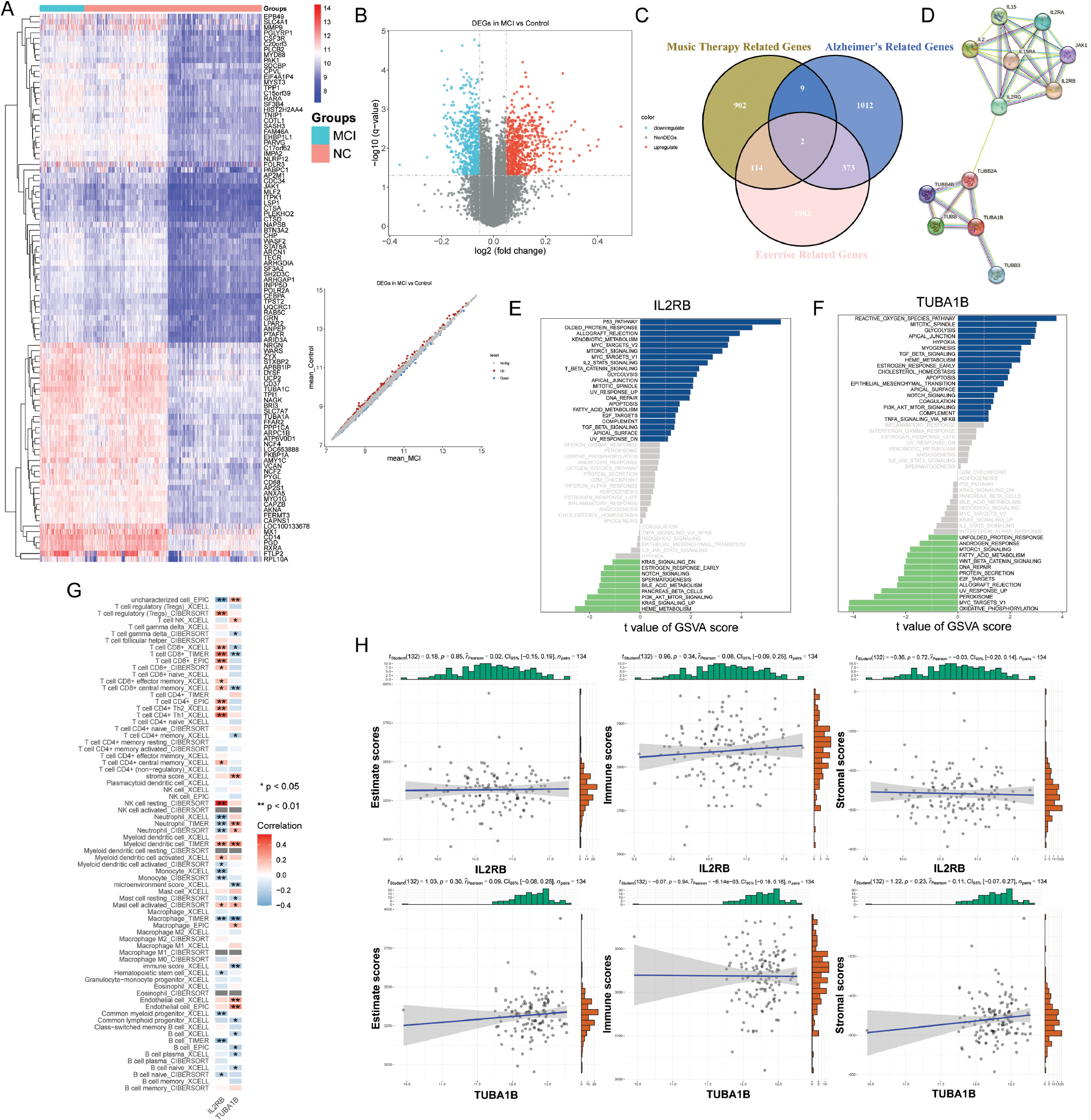
Our comprehensive analysis revealed distinct DEG profiles between two datasets, GSE140831 and GSE48624. In GSE140831, we identified 1,027 DEGs, comprising 600 downregulated and 427 upregulated genes (Figure 2A). In contrast, GSE48624 exhibited a homogeneous expression profile comprising 1,096 DEGs, all of which were upregulated. An exhaustive search for exercise-associated genes identified 6,471 genes (Figure 2B). Notably, a Venn diagram analysis highlighted two key genes, IL2RB and TUBA1B, implicated in exercise, music therapy, and AD (Figure 2C). These findings provided a basis for further investigation.
Figure 2 Identification of key genes and their functions. (A) Heatmaps of the top 100 differentially expressed genes (DEGs) from the GSE140831 dataset, displaying expression patterns throughout several samples. (B) Volcano plot illustrating DEGs within the GSE48624 dataset, highlighting markedly upregulated (red) or downregulated (blue) genes. (C) Venn diagram depicting the overlap among DEGs from GSE140831, music therapy-associated DEGs, and exercise-associated genes. (D) Protein-protein interaction (PPI) network indicating interactions between the IL2RB and TUBA1B proteins. (E, F) Gene set variation analysis (GSVA) enrichment plots for the key genes IL2RB and TUBA1B, demonstrating the enrichment rankings throughout distinct samples or conditions. (G) Correlation evaluation among immune cells and key genes, with statistical importance indicated with color coding. (H) Correlation evaluation of immune rating, stromal rating, and ESTIMATE rating with the key genes IL2RB and TUBA1B, indicating scatter plots and trend lines. Experimental techniques included gene expression profiling and bioinformatics analyses, and statistical relevance is denoted by p-values where applicable.
Protein-protein interaction and gene set variation analysis
PPI network analysis suggested potential regulatory interactions between TUBB2A and IL2RG in relation to the key genes, IL2RB and TUBA1B (Figure 2D). Additionally, gene set variation analysis (GSVA) revealed significant enrichment of these genes in pathways such as glycolysis, apical junctions, and TGF-beta signaling, thereby suggesting synergistic modulation of gene expression (Figure 2E-F).
Immune infiltration and correlation analysis
Investigation of immune cell infiltration revealed contrasting roles of the key genes. IL2RG positively correlated with CD8+ T cells and CD8+ central memory T cells, but negatively correlated with neutrophil populations. In contrast, TUBB2A exhibited an inverse relationship, correlating negatively with T cell subsets and positively with neutrophils (Figure 2G). These findings suggested opposing regulatory roles in the immune response. However, the absence of significant correlations between the immune score, stromal score, or ESTIMATE score and the key genes IL2RB and TUBA1B (Figure 2H) suggested a modulatory role rather than a primary role in immune regulation.
Construction and analysis of the mRNA-miRNA network
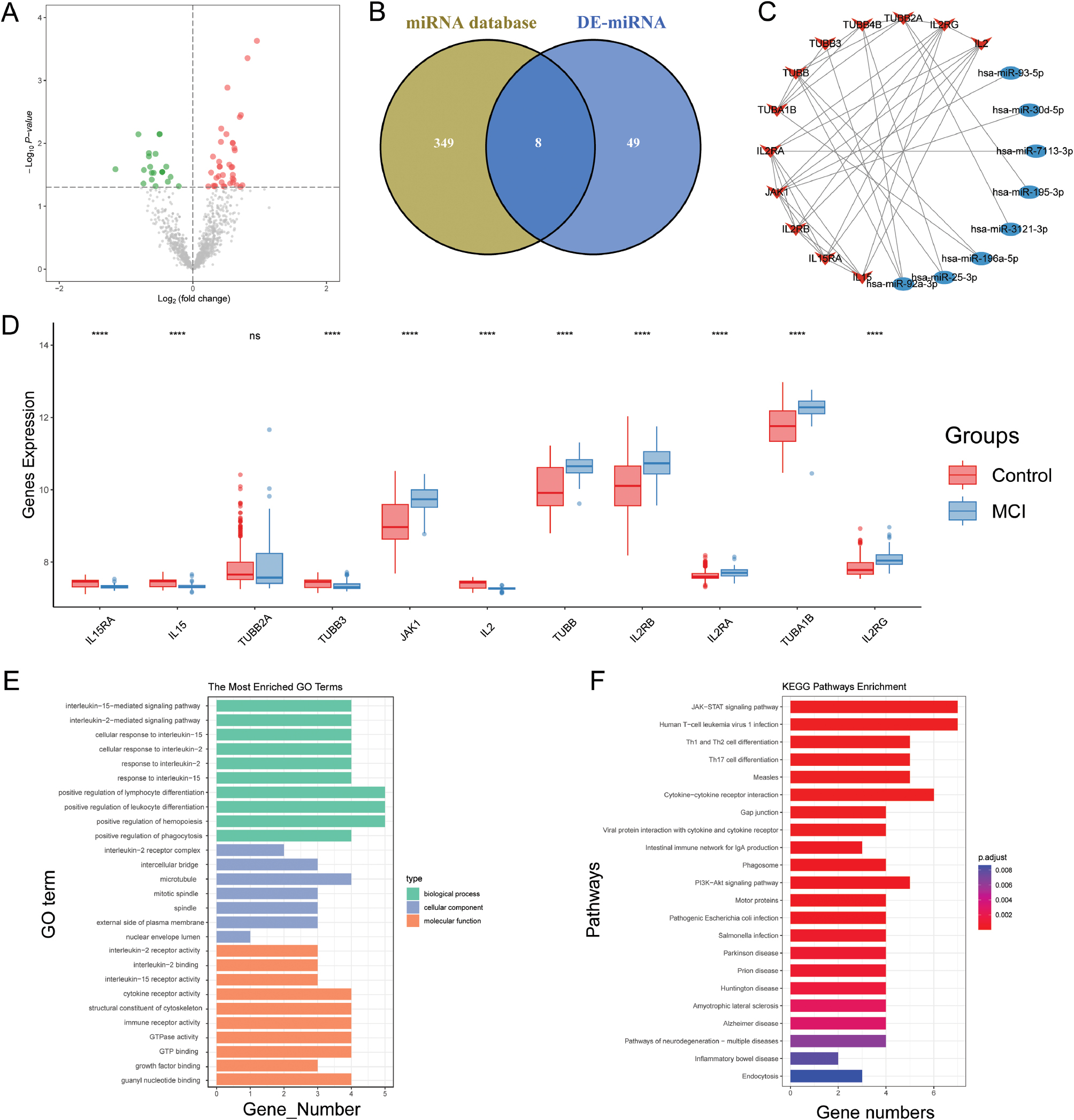
Our differential analysis of the GSE115071 dataset identified 57 differentially expressed microRNAs (DE-miRNAs): 20 downregulated and 37 upregulated (Figure 3A). Cross-referencing against the miRTarBase database led to the identification of 357 miRNAs with regulatory relationships to 11 mRNAs. Subsequently, a Venn diagram highlighted eight overlapping miRNAs (Figure 3B). These findings facilitated the construction of an mRNA-miRNA network comprising 11 mRNAs and eight miRNAs (Figure 3C).
Figure 3 Construction of mRNA-miRNA network. (A) Volcano plot of differentially expressed miRNAs (DE-miRNAs) in the GSE115071 dataset, with significantly upregulated miRNAs shown in red and significantly downregulated miRNAs shown in green. (B) Venn diagram comparing the overlap of DE-miRNAs identified with a pre-existing miRNA database. (C) Network diagram illustrating the mRNA-miRNA interactions in the study. (D) Differential expression analysis of 11 key genes from the protein-protein interaction (PPI) network between control and mild cognitive impairment (MCI) groups, presented in box plots. (E) GO enrichment analysis for the 11 PPI network genes, categorized by biological process, cellular component, and molecular function. (F) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of the 11 PPI network genes, with pathways ranked by p-value significance.
Differential and enrichment analyses
We observed significant differential expression for genes other than TUBB2A between the mild cognitive impairment (MCI) and control groups, a finding supporting the critical role of the PPI network centered around IL2RB and TUBA1B in MCI progression (Figure 3D). Further functional regulatory analysis via Gene Ontology (GO) enrichment revealed key roles of immune-related signaling pathways, notably the interleukin-15-mediated signaling pathway, the interleukin-2-mediated signaling pathway, and the interleukin-2 receptor complex (Figure 3E). Concurrently, KEGG pathway enrichment analysis highlighted the JAK-STAT signaling pathway, human T-cell leukemia virus 1 infection, and Th1/Th2 cell differentiation as central pathways (Figure 3F), thus suggesting regulatory effects of IL2RB and TUBA1B on immune function in MCI.
Elucidation of drug-target interactions in disease modulation
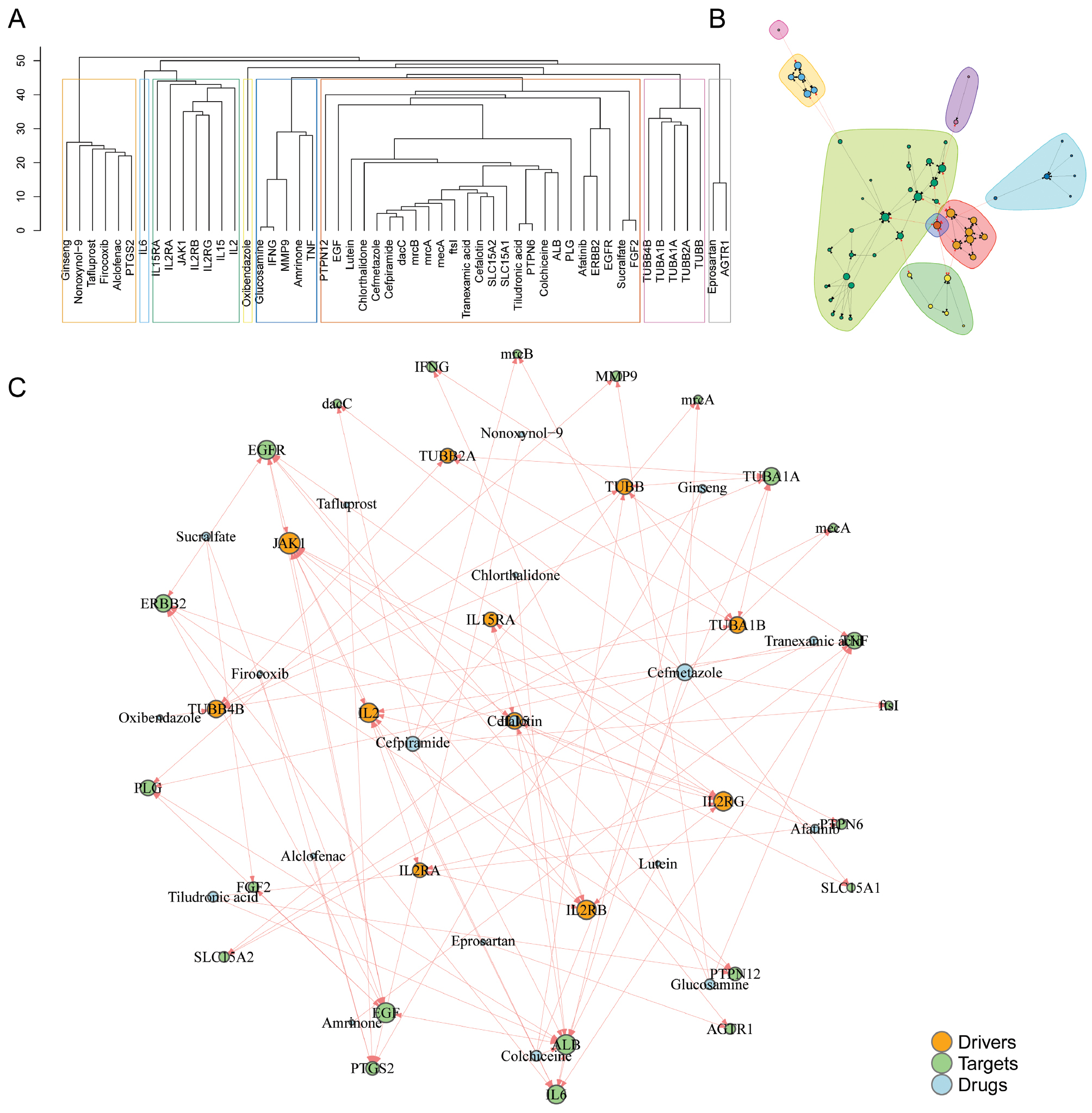
We used a multifaceted approach to unravel the intricate network of drug-target interactions relevant to disease modulation. Cluster analysis revealed a hierarchically structured dendrogram indicating potential connections among various compounds and genes within the human gene interaction network (Figure 4A). Distinct hierarchical clusters highlighted the proximity and possible interactions between entities that might play critical roles in disease pathways. Further analysis using a clustergram from network pharmacology predictions identified distinct groups of compounds or genes and consequently suggested subgroup-specific interactions essential for targeted therapeutic strategies (Figure 4B). Our analysis culminated in the construction of a detailed network pharmacology model illustrating the vast interconnectivity among genes, drugs, and disease drivers (Figure 4C). In this network, node size reflected the number of connections to a gene, whereas edge thickness indicated the strength of correlation, thus emphasizing key drivers and potential drug targets. This intricate network model provided valuable insights into complex regulatory interactions and might pave the way to the identification of critical nodes serving as promising therapeutic targets.
Figure 4 Analysis of the shortest pathway between drug targets and disease-related genes. (A) Dendrogram from cluster analysis depicting the hierarchical relationships among compounds or genes, indicating potential connections according to proximity within the human gene interaction network. (B) Clustergram associated with network pharmacology predictions, highlighting distinct groups of compounds or genes. (C) Network pharmacology analysis illustrating the interconnectivity between genes. Node size is proportional to the number of connections, and edge thickness represents correlation strength.
Integrative pathway and regulatory network analysis in cognitive impairment
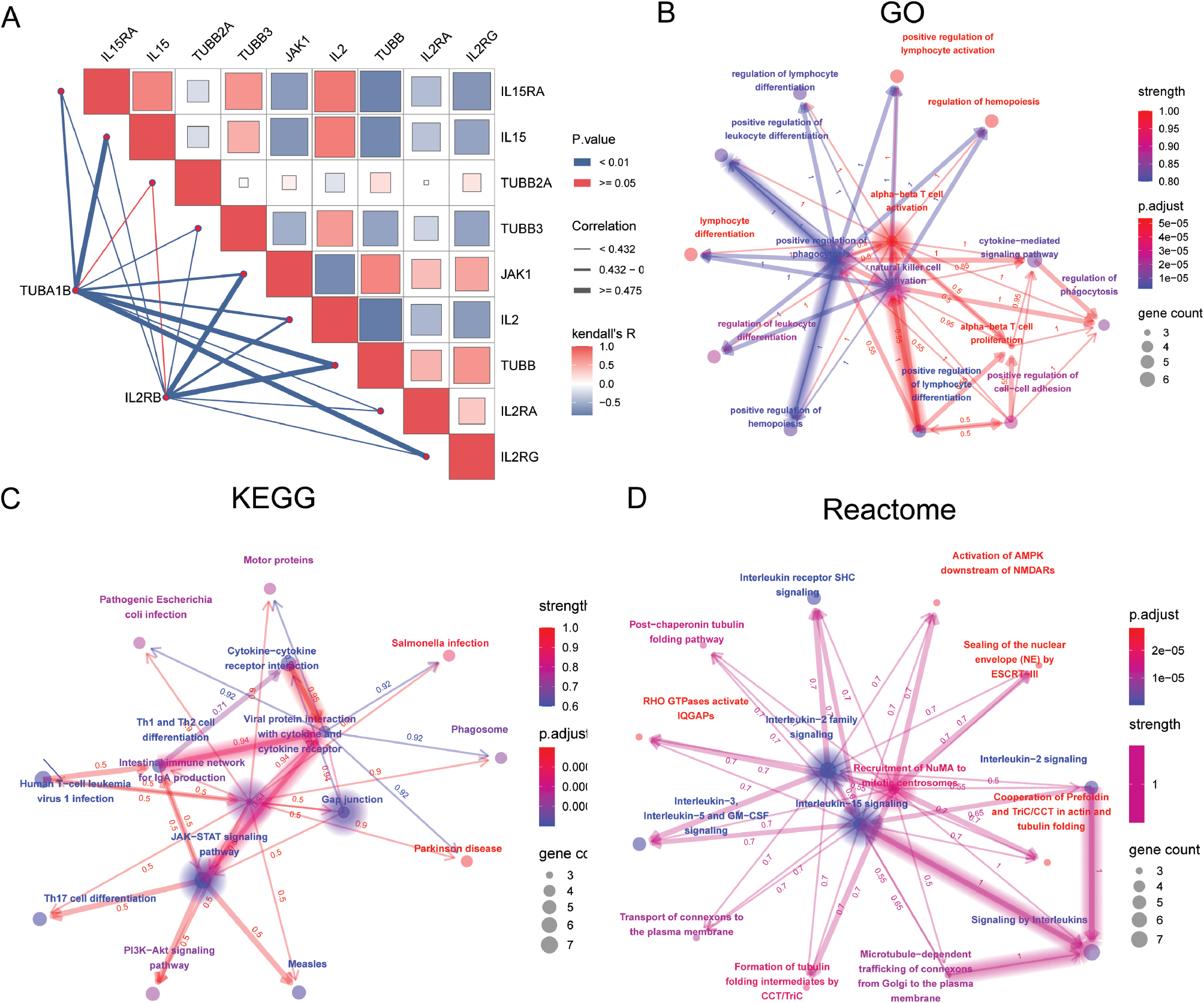
To elucidate the molecular mechanisms underlying cognitive impairment, we constructed a correlation matrix, which revealed a nuanced interplay among key genes (Figure 5A). The red and blue hues within the matrix delineated positive and negative correlations, respectively, and the color intensity was proportional to correlation strength. Kendall’s tau coefficient, represented by line thickness, reinforced the robust associations among the identified genes. GO enrichment analysis elucidated the biological processes, cellular components, and molecular functions involved in cognitive impairment (Figure 5B). The resulting network of enriched GO terms, connected by shared genes, suggested a complex regulatory landscape, with node color intensity reflecting p-value significance and node size indicating gene count. We further performed KEGG pathway analysis, which highlighted critical biological pathways characterized by varying gene counts and differential adjusted p-value significance levels (Figure 5C). Reactome pathway analysis additionally identified significant pathways, with edge thickness signifying interaction strength and node color corresponding to the significance according to the adjusted p value (Figure 5D).
Figure 5 Pathway and regulatory analysis in cognitive impairment. (A) Correlation analysis matrix showing the relationships among key genes, with red indicating positive correlation and blue indicating negative correlation. Correlation strength is indicated by color saturation, and line thickness represents Kendall’s tau coefficient. (B) Gene Ontology (GO) enrichment analysis in cognitive impairment. Nodes represent enriched GO terms connected by their shared genes, color indicates p-value significance, and node size reflects gene count. (C) Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis, highlighting pathways. Nodes are sized according to gene count, and edges are colored according to adjusted p-value significance levels. (D) Reactome pathway analysis depicting interactions among significant pathways involved in cognitive impairment. Edge thickness represents association strength, and node color represents the adjusted p-value significance level.
Functional pathway regulation in immune response
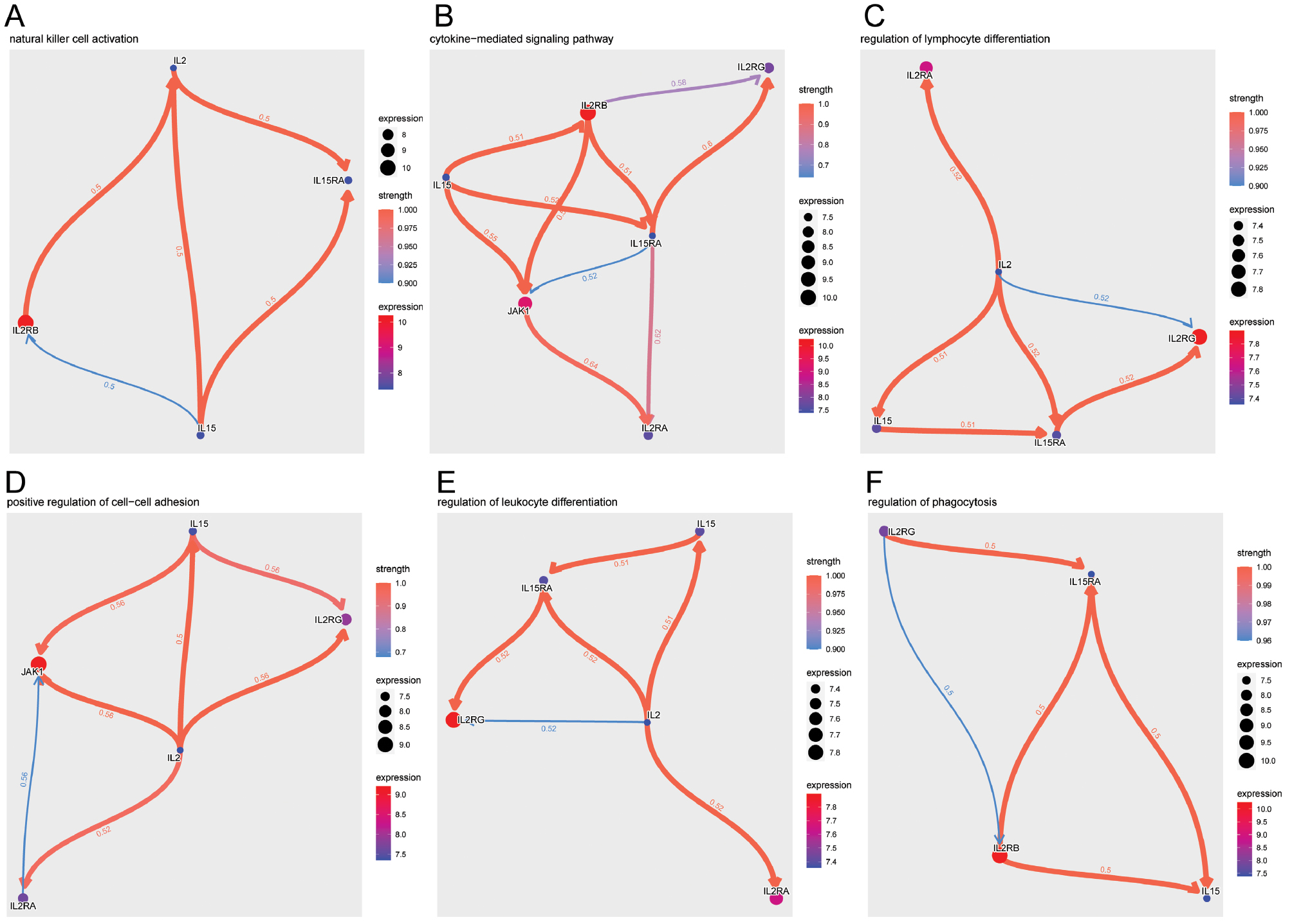
The natural killer cell activation pathway was prominent, and IL-2 and IL-15 were found to play substantial roles in regulating immune responses, particularly in the modulation of inflammation and immune cell activation (Figure 6A). The cytokine-mediated signaling pathway also emerged as significant; IL2RB was central to its critical regulatory role in immune modulation, as evidenced by multiple connections reflecting its regulatory function in the immune response (Figure 6B). Lymphocyte differentiation, an essential aspect of the immune response, displayed a network of regulatory interactions, particularly involving IL2RA and IL2RG, which were key modulators in this process (Figure 6C). The positive regulation of cell-cell adhesion, which is essential for immune synapse formation, was highlighted by interactions involving JAK1 and IL-15 (Figure 6D). Furthermore, the regulation of leukocyte differentiation demonstrated the influence of IL-2 on the immune landscape, particularly in the differentiation of various leukocyte subtypes (Figure 6E). Finally, the regulation of the phagocytosis pathway revealed a noteworthy negative correlation between IL2RG and IL-15RA, a finding suggesting a complex interplay in the phagocytic process (Figure 6F).
Figure 6 GO Pathway regulation analysis. (A) Natural killer cell activation pathway. (B) Cytokine-mediated signaling pathway. (C) Regulation of lymphocyte differentiation pathway. (D) Positive regulation of cell-cell adhesion pathway. (E) Regulation of leukocyte differentiation pathway. (F) Regulation of phagocytosis pathway. Nodes indicate individual genes, and their expression levels are denoted by node size. Edge color and width represent regulatory strength and direction: red for upregulation and blue for downregulation in the context of cognitive impairment.
Delineation of immune pathway dynamics in cognitive impairment via KEGG analysis
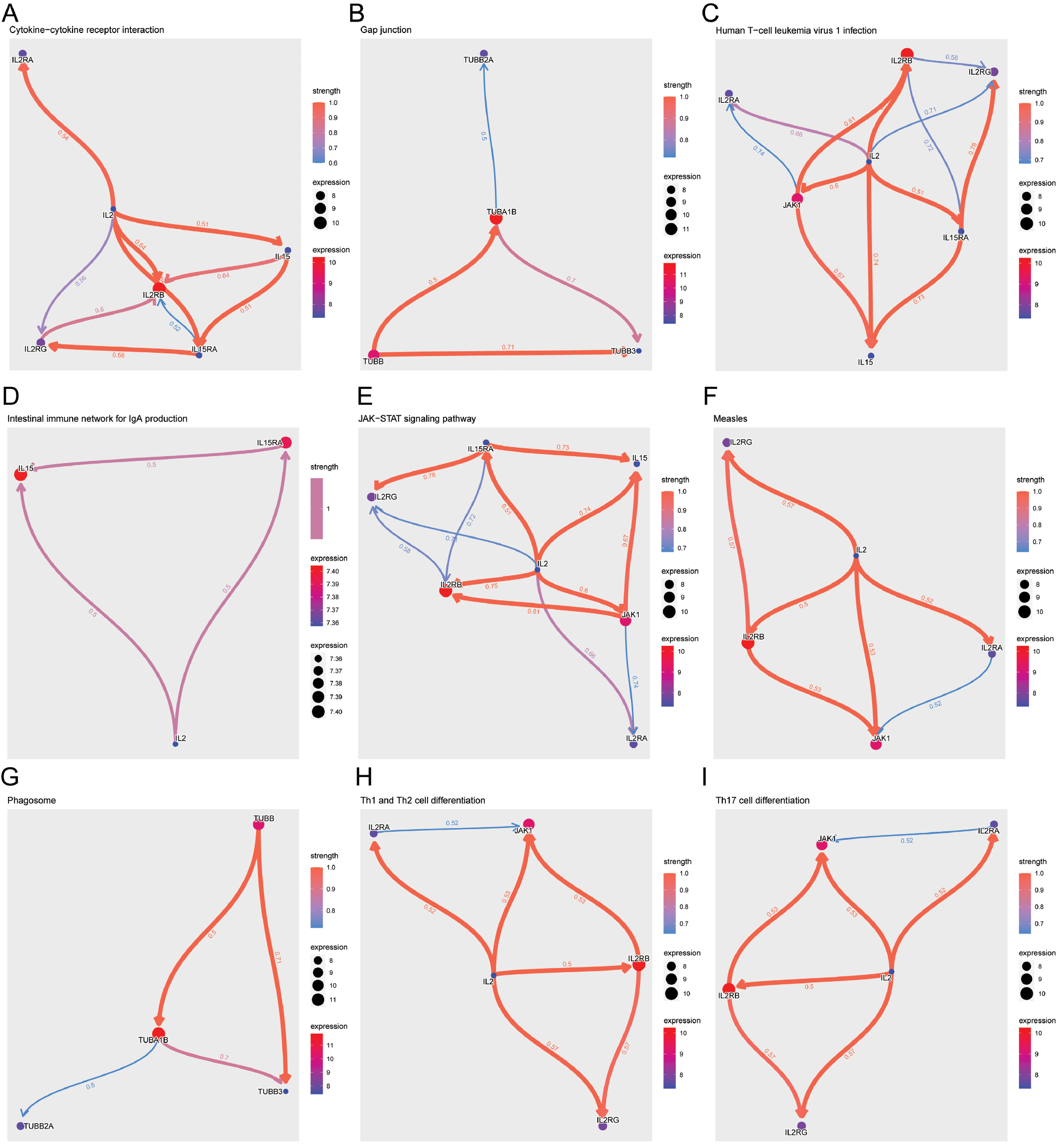
KEGG analysis of immune-related pathways revealed significant regulatory patterns in cognitive impairment. The cytokine-cytokine receptor interaction pathway’s considerable regulatory strength, particularly in the upregulation of IL-2, suggested enhanced immune communication (Figure 7A). Gap junction pathways, which are crucial for intercellular communication, exhibited a trend toward TUBA1B downregulation potentially reflecting altered neuronal connectivity (Figure 7B). Infection-related pathways, such as those for human T-cell leukemia virus 1 and measles, were implicated, thereby indicating potential viral effects on immune responses in cognitive impairment (Figure 7C and 7F). The significant downregulation of the intestinal immune network for IgA production suggested a diminished response from gut-associated lymphoid tissue (Figure 7D). The JAK-STAT signaling pathway, which is essential for cytokine signaling, demonstrated mixed regulatory patterns reflecting the complex role of this pathway in cognitive function (Figure 7E). Moreover, the analysis of Th1/Th2 and Th17 cell differentiation pathways revealed a potent regulatory network in which IL2RG was consistently implicated, a finding highlighting its versatile role in T cell differentiation (Figure 7H and 7I). The phagosome pathway, involved in pathogen clearance, displayed a balanced regulatory pattern indicating preserved phagocytic function (Figure 7G).
Figure 7 Kyoto encyclopedia of genes and genomes pathway regulatory analysis. (A) Cytokine-cytokine receptor interaction pathway. (B) Gap junction pathway. (C) Human T-cell leukemia virus 1 infection pathway. (D) Intestinal immune network for IgA production pathway. (E) JAK-STAT signaling pathway. (F) Measles infection pathway. (G) Phagosome pathway. (H) Th1 and Th2 cell differentiation pathway. (I) Th17 cell differentiation pathway. Nodes indicate individual genes, and their expression levels are denoted by node size. Edge color and width represent regulatory strength and direction: red for upregulation and blue for downregulation in the context of cognitive impairment.
Reactome pathway regulatory analysis in cognitive impairment
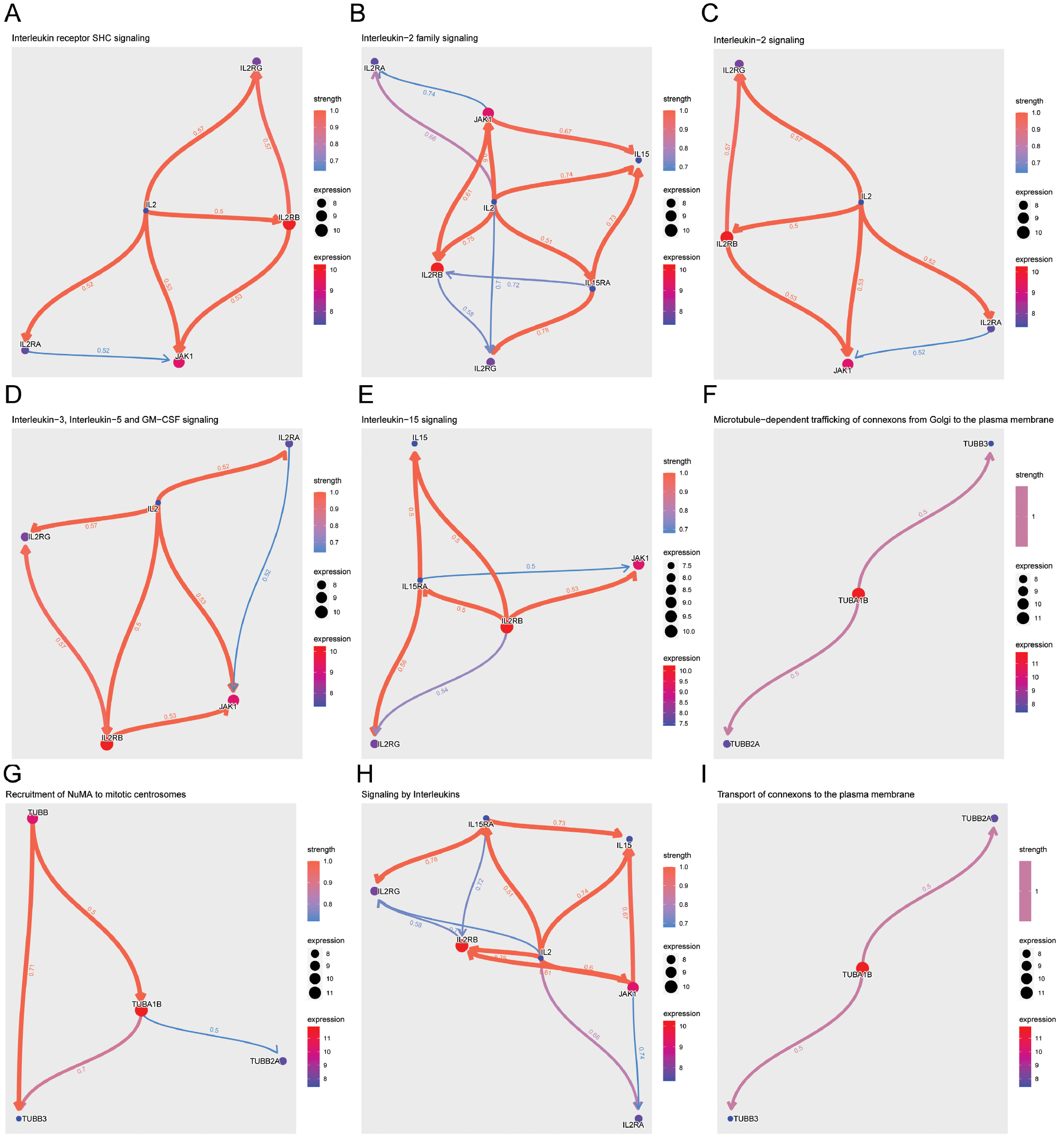
Our reactome pathway regulatory analysis uncovered notable modulation of the interleukin signaling cascade, with implications in cognitive impairment. The interleukin receptor SHC signaling pathway (Figure 8A) showed pronounced upregulation, evidenced by enlarged nodes correlating with heightened gene expression. Similarly, the interleukin-2 family (Figure 8B) and interleukin-2 signaling pathways (Figure 8C) revealed robust red edges suggesting upregulation potentially contributing to disease pathology. Additionally, the combined signaling pathways of interleukins-3, -5, and GM-CSF (Figure 8D) were predominantly upregulated, whereas the interleukin-15 signaling pathway (Figure 8E) exhibited a mixture of regulatory influences suggesting a nuanced role in cognitive impairment. Notably, the analysis of microtubule-dependent trafficking of connexons from the Golgi to the plasma membrane (Figure 8F) highlighted TUBA1B as a central upregulated node and suggested its involvement in the cellular communication disruptions observed in cognitive decline. The recruitment of NuMA to mitotic centrosomes (Figure 8G) showed similar upregulation, particularly in TUBB3, thus hinting at its contribution to cell division anomalies associated with neurodegeneration. Additionally, interleukin signaling (Figure 8H) underscored complex immune responses; IL2RB was notably upregulated, thereby emphasizing its regulatory importance. Ultimately, we identified connexon delivery to the plasma membrane (Figure 8I) as an important process that is subject to significant regulatory modulation by TUBB2A. Our results provide a broad understanding of changes in gene expression in cognitive impairment that might be explored as therapeutic targets in interleukin and microtubule signaling pathways.
Figure 8 Reactome pathway regulatory analysis. (A) Interleukin receptor SHC signaling pathway. (B) Interleukin-2 family signaling pathway. (C) Interleukin-2 signaling pathway. (D) Interleukin-3, -5, and GM-CSF signaling pathway. (E) Interleukin-15 signaling pathway. (F) Microtubule-dependent trafficking of connexons from the Golgi to the plasma membrane. (G) Recruitment of NuMA to mitotic centrosomes. (H) Interleukin signaling. (I) Transport of connexons to the plasma membrane. Nodes represent individual genes and are sized according to their expression levels. Edges show the direction and strength of regulation: red for upregulation and blue for downregulation in the context of cognitive impairment. Edge width indicates the degree of regulatory influence.
Discussion
The roles of big data and bioinformatics technologies in biomarker identification and applications are becoming increasingly important. These technologies are essential for diagnosing and predicting diseases [31, 32]. The integration of multi-omic data with bioinformatics analysis and novel drug development strategies holds promise for diagnostics and therapeutic interventions for complex diseases, and provides insights for guiding future research directions [33–37]. This study examined the relationships among DEGs, immunity, and therapeutic genes in AD and related cognitive impairments. Through a detailed analysis, we mapped the relationships between DEGs and several drug targets. This work expands the molecular understanding of the mechanisms underlying these complex disorders.
Studying the immune microenvironment has explained interactions between immune and metabolic pathways in disease and consequently provided important references for precision immunotherapy [38, 39]. Extensive studies on PPI networks and their regulatory mechanisms have revealed important roles in signaling and functional regulation of cells. Our investigation indicated a notable potential DEG signature. Among the identified DEGs, IL2RB and TUBA1B were found to play key roles in the regulation of immunity and cognition with respect to AD. Studies have implicated TUBB2A and IL2RB in interleukin-15 and interleukin-2 signaling pathways. Moreover, the JAK-STAT pathway is involved in HTLV-1 infection and in Th1/Th2 differentiation, an area of interest in investigations of HTLV-1. The inflammation from interleukins might accelerate AD progression. When the inflammatory response is triggered, various neuroinflammatory mediators are synthesized and secreted by astrocytes and microglia, including complement activators/inhibitors, chemokines, cytokines, free radicals, and inflammatory enzymes. Sufficient inflammation can kill neurons and lead to further inflammation. Additionally, subsets of immune cells might play major roles in AD. Studies have shown that TEMRA, a subset of immune cells from healthy individuals, people with AD, and people with MCI, disrupt neuronal activity and form neural protrusion spheres, a hallmark of AD-associated neuronal damage. Therefore, TEMRA cells not only secrete toxic cytokines but also directly damage neurons. The identification of these genes not only enriches current understanding but also might open avenues for novel therapeutic strategies. The observed pathway enrichment, particularly in glycolysis and the TGF-beta signaling cascade, underscores the complex biological processes disrupted in AD. These pathways have well-documented roles in immune modulation and synaptic plasticity, both of which are crucial for maintenance of cognitive function. Consequently, the interplay between these pathways highlights the role of the immune microenvironment in the diagnosis and prognosis of various diseases [40, 41]. In-depth studies in transcriptomics and proteomics have not only revealed key pathways and molecular mechanisms, but also provided a theoretical basis for the development of efficient targeted therapeutic agents [42, 43].
Music therapy is known to ameliorate cognitive function in patients with Alzheimer’s disease (AD), primarily by enhancing memory recall and retention [17, 44]. It achieves this by stimulating neural pathways associated with autobiographical memory, with studies confirming its role in supporting memory processes [45, 46]. As the disease progresses, the activities of sporting clubs, which encourage social interaction and a sense of belongingness, are diminished [47]. Music therapy encourages socialization and a sense of belonging, which tend to be lost as the disease progresses [48]. This stimulus encourages increased social interactions and a feeling of belongingness, which tends to decline with the progression of the disease [49, 50]. The immune microenvironment has been a key area of recent research. A transcriptomic analysis has indicated immune cell activity associated with the disease. The immune microenvironment plays a crucial role in studying immune evasion and the progression of Alzheimer’s disease. The findings from these studies have also contributed to the identification of novel immunotherapeutic strategies against cancer [51]. Targeting and altering protein interaction networks has been found to potentially interfere with signaling pathways that are essential in regulating cell functions. Music therapy appears particularly promising for the treatment of other diseases due to its demonstrated benefits in enhancing cognitive and motor skills [52, 53].
Music therapy’s demonstrated benefits extend beyond cognitive and emotional functions, and can improve motor skills and coordination [54]. Beyond its cognitive and emotional benefits, music therapy can enhance motor skills and coordination [55]. Being physically impaired may also present additional benefits, such as improving the therapeutic effects of music therapy, which can help patients with AD manage symptoms like agitation and tension. In general, AD worsens over time. By treating common symptoms such as tension and agitation, music therapy can help create a pleasant and soothing atmosphere [56]. Soothing music can calm behaviors and assist caregivers. The medical world is starting to understand the biological mechanisms underlying the efficacy of music therapy in AD. Music therapy in dementia relies on the ability of music to activate a wide array of brain areas, many of which are compromised in dementia. This involvement might stimulate synaptic plasticity and the formation of new neural connections, which could help ameliorate some of the degenerative effects associated with the disease. [57]. Moreover, the stimulation provided by music, combined with emotional and cognitive factors, might contribute to the release of neuroprotective factors and growth factors, and consequently enhance brain health [58].
Our mRNA-miRNA network analysis revealed the intricate regulatory mechanisms underlying cognitive impairment. The important network components identified, as well as their relationships, might play essential roles in eliciting the effects of exercise and music therapy on cognition. Our findings validate and expand on earlier research. The immune-regulatory function of IL2RB and the synaptic-stabilizing action of TUBA1B are consistent with prior hypotheses regarding immune-inflammatory aspects of AD. They also suggest that synapse health is essential for cognition. In addition, the enrichment in glycolysis and TGF-beta signaling pathways observed herein suggests that metabolic and immune dysregulation might be interlinked in AD pathogenesis. The consistent findings across studies suggest a need to exclude other mechanisms before identifying the cause of a disease or a therapeutic target [59]. Analysis of immune infiltration further indicated the roles of immune cells in disease progression. The immunological functions of DEGs suggested that targeted genetic engineering of specific genes might shift the disease course. This systems biology approach underscores the multifaceted nature of gene regulation and its profound effects on disease progression. Moreover, studies on neuroinflammation in AD have highlighted the importance of immunomodulation in disease management, thereby reinforcing our findings regarding the regulatory roles of IL2RB and TUBA1B in immune pathways [60, 61]. The development of novel targeted therapeutic strategies has promise in markedly enhancing treatment efficacy while decreasing adverse effects, and consequently advancing the field of precision medicine [62]. Comparative studies on different treatment approaches and their effects on patient prognosis have been extensively examined through systematic reviews and meta-analyses [63]. Computer-aided drug design trends have led to the ongoing emergence of research hotspots in this field and offered new possibilities for drug development [64]. Moreover, the development of highly antifungal wood preservatives from traditional herbal root extracts has offered a new perspective in materials science [65]. Transcriptomics is critical in revealing the immune microenvironment, and can provide valuable insights regarding the diagnosis and prognosis of various diseases [66, 67]. Extensive studies examining the key roles of PPIs and their regulation have shed light on their importance in cell signaling and functional regulation within biological systems [68–70].
The use of big data and bioinformatics in biomarker identification has become increasingly important in disease diagnosis and prognostication [71, 72]. Through the integration of data-driven approaches and experimental validation, researchers have made significant advancements in several key areas, including the identification of novel biomarkers, improved diagnostic methods, and the development of potential therapeutic strategies [71].
Despite the valuable contributions of this study, certain limitations should be acknowledged. The reliance on gene expression data derived from autopsy brain samples limited our ability to capture dynamic temporal modifications in gene expression. Future studies should incorporate longitudinal observation designs with real-time patient tracking to validate our findings and explore their clinical applicability. Moreover, the central modulators identified, such as IL2RB and TUBA1B, and the proposed mRNA-miRNA network, were hypotheses generated from in silico analysis. Although the drug-target interaction evaluation provided useful insights, it was based on computational predictions and requires experimental validation. Subsequent research should focus on in vitro cell experiments and the development of AD animal models for in vivo validation, to fully evaluate the therapeutic potential of the identified drug targets.
Understanding the roles of cytokines in immune responses provides valuable insights for developing effective therapeutic strategies for a range of diseases [73, 74]. With the growing body of research on exercise and music therapy, particularly their applications in neurodegenerative diseases such as Alzheimer’s disease, scientists are increasingly focusing on how these interventions modulate the interactions between immune microenvironments and metabolic pathways, thereby offering new perspectives for precision therapy. Exercise and music therapy not only positively influence the immune microenvironment but also might enhance overall patient health by affecting gene expression, protein interactions, and related signaling pathways; consequently, these interventions have potential for treating chronic diseases and improving quality of life [35, 62].
Conclusion
Our findings notably advance basic understanding of DEGs, immune-related genes, and therapy-associated genes in AD, as well as CI in AD. This study identified IL2RB and TUBA1B as potential therapeutic targets and highlighted key pathways and regulatory networks that might enhance targeted therapy in cancer. However, further studies are necessary to validate these findings and demonstrate their potential for clinical translation, with the ultimate goal of ameliorating cognitive impairment and AD.
Data availability statement
The data used in this study are publicly available from the Gene Expression Omnibus (GEO) database (datasets GSE140831, GSE48624, and GSE115071). The raw data and analysis scripts are available from the corresponding author upon reasonable request.
Ethics statement
No direct interactions with human or animal subjects were involved. Therefore, ethical approval and informed consent were not required.
Author contributions
Ke Wu, Jianling Liu, Yusheng Su, and Yisheng Chen contributed equally to this work. Ke Wu, Jianling Liu, and Yisheng Chen were primarily responsible for the study design, data analysis, and interpretation of results. Yusheng Su, Jiansheng Yang, and Jianzhong Xiao contributed to the investigation, data collection, and initial manuscript drafting. Jia Lina, Zhengzhou Chen, and Yunshen Ge were involved in the experimental design and data validation. Wei Luo and Zhong He provided critical revisions to the manuscript and facilitated access to key resources. Yanjin Wu and Chengwan Shen assisted with the statistical analysis and manuscript refinement. Haojun Shi contributed to the integration of music therapy in the study, whereas Bin Huang provided expertise in molecular techniques. John H Zhang played a major role in manuscript revisions and final approval. All authors reviewed and approved the final manuscript.
Funding
This research was funded by the National Scholarship for Studying Abroad (202306100231) and the 2024 Research Project of Ningde Normal University (Project No. 000059091501).
Acknowledgements
We sincerely thank the reviewers for their careful evaluation and valuable comments on this article.
Conflict of interest
Yisheng Chen is a board member of BIO Integration. He was not involved in the peer-review or handling of the manuscript. The other authors declare that there are no conflicts of interest.
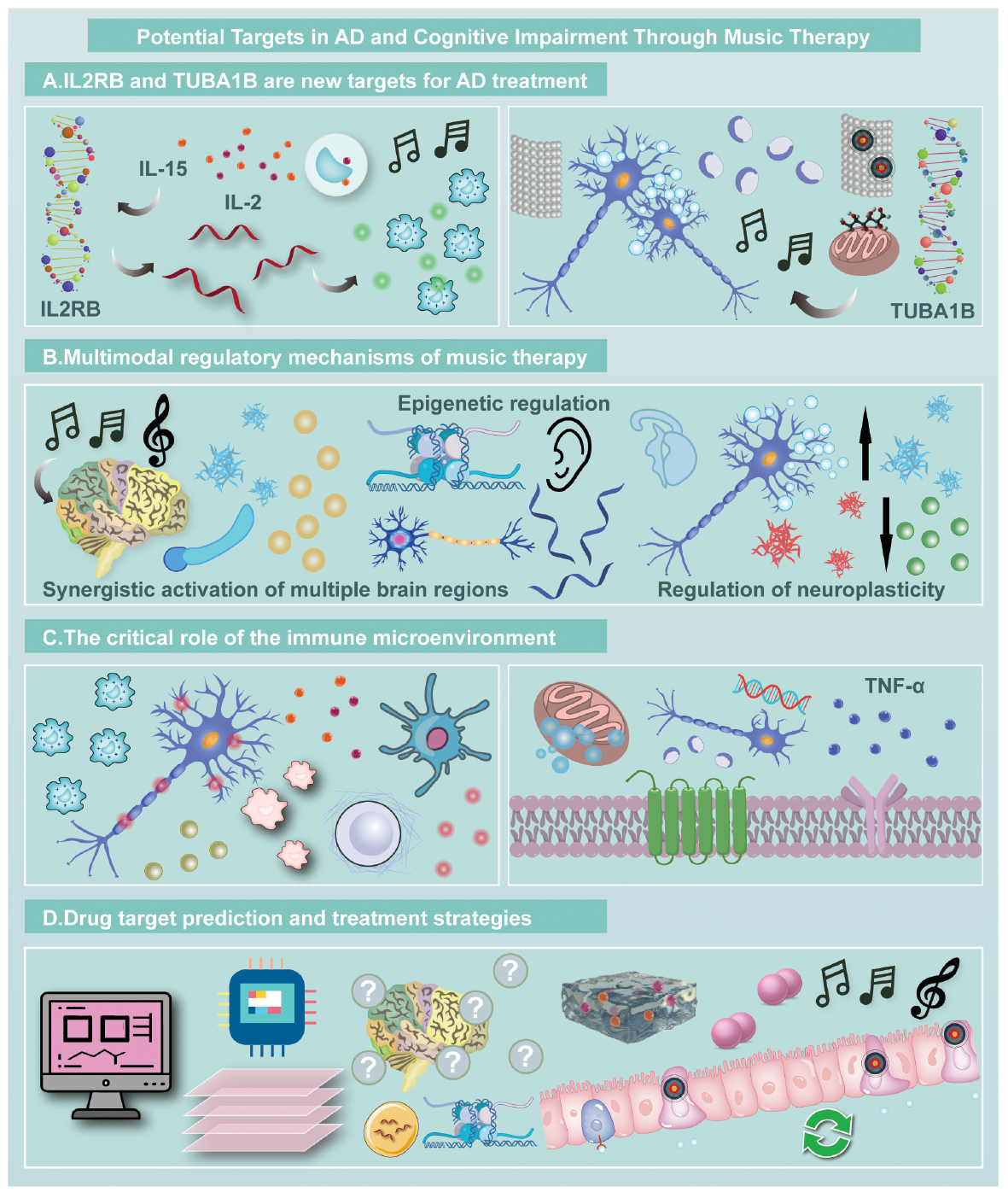
Graphical abstract
Highlights
- IL2RB and TUBA1B as are important immune and metabolic regulatory genes in Alzheimer’s disease and cognitive impairment.
- Music therapy demonstrates potential therapeutic prospects by promoting neuroplasticity, immune balance, and gene expression regulation.
In brief
This study explores the potential of IL2RB and TUBA1B as therapeutic targets for Alzheimer’s disease and cognitive impairment, and for the first time incorporates music therapy into the analysis of immune and metabolic pathways, providing new insights for precision medicine.
References
- Reiman EM, Mattke S, Kordower JH, Khachaturian ZS, Khachaturian AS. Developing a pathway to support the appropriate, affordable, and widespread use of effective Alzheimer’s prevention drugs. Alzheimers Dement 2022;18:7-9. [PMID: 35103395 DOI: 10.1002/alz.12533]
- Wei C. The risk and the prevention of Alzheimer’s disease and its challenges within these thirty years. Highl Sci Eng Technol 2022;8:163-8. [DOI: 10.54097/hset.v8i.1123]
- Bird TD. Genetic factors in Alzheimer’s disease. N Engl J Med 2005;352:862-4. [DOI: 10.1056/NEJMp058027]
- Ku C-S, Vasiliou V, Cooper DN. A new era in the discovery of de novo mutations underlying human genetic disease. Hum Genomics 2012;6:27. [PMID: 23232122 DOI: 10.1186/1479-7364-6-27]
- Ahmad SR, Zeyaullah M, AlShahrani AM, Dawria A, Ali H, et al. Deciphering the enigma of Neuron-Glial interactions in neurological disorders. Front Biosci (Landmark Ed) 2024;29:142. [PMID: 38682185 DOI: 10.31083/j.fbl2904142]
- Li Y-Y, Guo D-D, Duan R-N, Li Y. Interactions between beta-amyloid and pericytes in Alzheimer’s disease. Front Biosci (Landmark Ed) 2024;29:136. [PMID: 38682184 DOI: 10.31083/j.fbl2904136]
- Chen Y, Luo Z, Sun Y, Li F, Han Z, et al. Exercise improves choroid plexus epithelial cells metabolism to prevent glial cell-associated neurodegeneration. Front Pharmacol 2022;13:1010785. [PMID: 36188600 DOI: 10.3389/fphar.2022.1010785]
- Huang J, Lin W, Sun Y, Wang Q, He S, et al. Quercetin targets VCAM1 to prevent diabetic cerebrovascular endothelial cell injury. Front Aging Neurosci 2022;14:944195. [PMID: 36118693 DOI: 10.3389/fnagi.2022.944195]
- Klay D, Grutters JC, van der Vis JJ, Platenburg MGJP, Kelder JC, et al. Progressive Disease with low survival in adult patients with pulmonary fibrosis carrying surfactant-related gene mutations: an observational study. Chest 2023;163:870-80. [PMID: 36370864 DOI: 10.1016/j.chest.2022.11.002]
- Leggieri M, Thaut MH, Fornazzari L, Schweizer TA, Barfett J, et al. Music intervention approaches for Alzheimer’s disease: a review of the literature. Front Neurosci 2019;13:132. [PMID: 30930728 DOI: 10.3389/fnins.2019.00132]
- Hobeika L, Samson S. Why do music-based interventions benefit persons with neurodegenerative disease? Music and the Aging Brain, Elsevier; 2020. p. 333-49. [DOI: 10.1016/B978-0-12-817422-7.00013-4]
- Koelsch S. A neuroscientific perspective on music therapy. Ann N Y Acad Sci 2009;1169:374-84. [PMID: 19673812 DOI: 10.1111/j.1749-6632.2009.04592.x]
- Benavides JA, Caparrós C, Da Silva RM, Lembo T, Tem Dia P, et al. The power of music to prevent and control emerging infectious diseases. Front Med 2021;8:756152. [PMID: 34901067 DOI: 10.3389/fmed.2021.756152]
- de Quadros A. Music, the arts, and global health: in search of sangam, its theory and paradigms. J Folk Res 2017;54:15. [DOI: 10.2979/jfolkrese.54.2.02]
- Altenmuller E, Schlaug G. Neurobiological aspects of neurologic music therapy. Music Med 2013;5:210-6. [DOI: 10.1177/1943862113505328]
- Chan MMY, Han YMY. The functional brain networks activated by music listening: a neuroimaging meta-analysis and implications for treatment. Neuropsychology 2022;36:4-22. [PMID: 34636582 DOI: 10.1037/neu0000777]
- Peck KJ, Girard TA, Russo FA, Fiocco AJ. Music and memory in Alzheimer’s disease and the potential underlying mechanisms. J Alzheimers Dis 2016;51:949-59. [PMID: 26967216 DOI: 10.3233/JAD-150998]
- Matrone C, Brattico E. The power of music on Alzheimer’s disease and the need to understand the underlying molecular mechanisms. J Alzheimers Dis Parkinsonism 2015;5:196. [DOI: 10.4172/2161-0460.1000196]
- Särkämö T. Cognitive, emotional, and neural benefits of musical leisure activities in aging and neurological rehabilitation: a critical review. Ann Phys Rehabil Med 2018;61:414-8. [PMID: 28461128 DOI: 10.1016/j.rehab.2017.03.006]
- Ridder HM, Bøtker JØ. Music therapy and skill sharing to meet psychosocial needs for persons with advanced dementia. music and dementia. Oxford University Press; 2019. pp. 225-41. [DOI: 10.1093/oso/9780190075934.003.0011]
- Wu J-F, Chen Y-S, Xie Y-C. Single-cell transcriptomic profiling reveals ZEB1-mediated regulation in microglial subtypes and the impact of exercise on neuroinflammatory responses. Tradit Med Res 2025;10:11. [DOI: 10.53388/TMR20240423002]
- Wan R, Chen Y, Feng X, Luo Z, Peng Z, et al. Exercise potentially prevents colorectal cancer liver metastases by suppressing tumor epithelial cell stemness via RPS4X downregulation. Heliyon 2024;10:e26604. [PMID: 38439884 DOI: 10.1016/j.heliyon.2024.e26604]
- Yan C, Chen Y, Sun C, Ahmed MA, Bhan C, et al. Does proton pump inhibitor use lead to a higher risk of coronavirus disease 2019 infection and Progression to Severe Disease? a Meta-analysis. Jpn J Infect Dis 2022;75:10-5. [PMID: 34053958 DOI: 10.7883/yoken.JJID.2021.074]
- Chauleau J-Y, Trassin M. Sensing multiferroic states non-invasively using optical second harmonic generation. Microstructures 2024;4:2024005. [DOI: 10.20517/microstructures.2023.50]
- Jang J, Choi S-Y. Reduced dimensional ferroelectric domains and their characterization techniques. Microstructures 2024;4:2024016. [DOI: 10.20517/microstructures.2023.67]
- Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 2015;43:e47. [PMID: 25605792 DOI: 10.1093/nar/gkv007]
- Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinformatics 2013;14:7. [PMID: 23323831 DOI: 10.1186/1471-2105-14-7]
- Yu G, Wang L-G, Han Y, He Q-Y. clusterProfiler: an R Package for Comparing Biological Themes Among Gene Clusters. OMICS 2012;16:284-7. [DOI: 10.1089/omi.2011.0118]
- Sato N, Tamada Y, Yu G, Okuno Y. CBNplot : Bayesian network plots for enrichment analysis. Bioinformatics 2022;38:2959-60. [PMID: 22455463 DOI: 10.1093/bioinformatics/btac175]
- Guney E, Menche J, Vidal M, Barábasi A-L. Network-based in silico drug efficacy screening. Nat Commun 2016;7:10331. [PMID: 26831545 DOI: 10.1038/ncomms10331]
- Ou S, Xu Y, Liu Q, Yang T, Chen W, et al. Analysis of large datasets for identifying molecular targets in intestinal polyps and metabolic disorders. Biocell 2024;48:415-29. [DOI: 10.32604/biocell.2024.046178]
- Wu J, Zhi Z, Xu W, Li D, Li Q, et al. LIM1863 is useful to explore collective cancer cell migration, and the group of heterogeneous cells undergoing collective migration behaves like a supracellular unit. Biocell 2023;47:2671-80. [DOI: 10.32604/biocell.2023.043494]
- Li M, Liu X, Jiang M, Lei Y, Li Z, et al. Prognostic capability of clinical SYNTAX score in patients with complex coronary artery disease and chronic renal insufficiency undergoing percutaneous coronary intervention. Rev Cardiovasc Med 2024;25:18. [PMID: 39077637 DOI: 10.31083/j.rcm2501018]
- Figueredo VM. The heart renaissance. Rev Cardiovasc Med 2024;25:91. [DOI: 10.31083/j.rcm2503091]
- Chen L, Yang T, Wu J, Cheng G, Zhao M, et al. Multi-omics strategy reveals that Cordyceps sinensis ameliorates sepsis-associated acute kidney injury via reprogramming of mitochondrial energy metabolism and macrophage polarization. Acta Mater Med 2024;3:269-88. [DOI: 10.15212/AMM-2024-0018]
- Luo X, Liu J, Du X, Yang J, Jiang X, et al. A comparative analysis of vaccine lists, prices, and candidates, and the national immunization program between China and the United States. Acta Mater Med 2024;3:46-56. [DOI: 10.15212/AMM-2023-0033]
- Wang M, Xu S, Shi X, Da Silva-Júnior EF, Zhan P. Solving the mysteries of urate transport: structural insights into GLUT9 and URAT1. Acta Mater Med 2024;3:345-8. [DOI: 10.15212/AMM-2024-0048]
- Mei K, Chen Z, Huang L, Wang J, Wei Y. Correlation between the immune microenvironment and bladder cancer based on a prognostic miRNA risk model. CI 2024;3:37-48. [DOI: 10.58567/ci03020002]
- Mei K, Chen Z, Wang Q, Ali A, Huang Y, et al. A prognostic aging-related lncRNA risk model correlates with the immune microenvironment in HCC. CI 2024;3:37-48. [DOI: 10.58567/ci03020003]
- Tang Y, Tang R. Health neuroscience–how the brain/mind and body affect our health behavior and outcomes. J Integr Neurosci 2024;23:69. [PMID: 38682228 DOI: 10.31083/j.jin2304069]
- Wu S, Fu Z, Wang S, Zheng F, Qiu W, et al. Disrupted functional brain network architecture in sufferers with boxing-related repeated mild traumatic brain injury: a resting-state EEG study. J Integr Neurosci 2024;23:102. [PMID: 38812391 DOI: 10.31083/j.jin2305102]
- Zhang G, Zhang Y, Chen L, Liu L, Gao X. E3 ubiquitin ligase-dependent regulatory mechanism of TRIM family in carcinogenesis. CI 2023;2:37-48. [DOI: 10.58567/ci02020005]
- Qin H, Luo Z, Sun Y, He Z, Qi B, et al. Low-intensity pulsed ultrasound promotes skeletal muscle regeneration via modulating the inflammatory immune microenvironment. Int J Biol Sci 2023;19:1123-45. [PMID: 36923940 DOI: 10.7150/ijbs.79685]
- Moreira SV, Justi FRDR, Moreira M. Can musical intervention improve memory in Alzheimer’s patients? Evidence from a systematic review. Dement Neuropsychol 2018;12:133-42. [PMID: 29988347 DOI: 10.1590/1980-57642018dn12-020005]
- Quintin E-M. Music-evoked reward and emotion: relative strengths and response to intervention of people with ASD. Front Neural Circuits 2019;13:49. [PMID: 31619969 DOI: 10.3389/fncir.2019.00049]
- Zhang L. Effects of music therapy on well-being of people with Alzheimer’s disease and related dementia. J Stud Res 2020;9. [DOI: 10.47611/jsrhs.v9i2.1224]
- Reschke-Hernández AE, Gfeller K, Oleson J, Tranel D. Music therapy increases social and emotional well-being in persons with dementia: a randomized clinical crossover trial comparing singing to verbal discussion. J Music Ther 2023;60:314-42. [PMID: 37220880 DOI: 10.1093/jmt/thad015]
- Rio R. A community-based music therapy support group for people with Alzheimer’s disease and their caregivers: a sustainable partnership model. Front Med 2018;5:293. [PMID: 30460236 DOI: 10.3389/fmed.2018.00293]
- Thomson BR, Schwendinger N, Beckmann K, Gentinetta T, Couto D, et al. Haptoglobin attenuates cerebrospinal fluid hemoglobin-induced neurological deterioration in sheep. Transl Stroke Res 2025;16:728-32. [PMID: 38652234 DOI: 10.1007/s12975-024-01254-9]
- Wang X, Wen D, Xia F, Fang M, Zheng J, et al. Single-cell transcriptomics revealed white matter repair following subarachnoid hemorrhage. Transl Stroke Res 2025;16:800-16. [PMID: 38861152 DOI: 10.1007/s12975-024-01265-6]
- Shen L, Jiang H. Pan-cancer and single-cell analysis of actin cytoskeleton genes related to disulfidptosis. Open Med 2024;19:20240929. [PMID: 38584831 DOI: 10.1515/med-2024-0929]
- Wang H, Zhao W, Wang D, Chen J. ANO6 (TMEM16F) inhibits gastrointestinal stromal tumor growth and induces ferroptosis. Open Med 2024;19:20240941. [PMID: 38756246 DOI: 10.1515/med-2024-0941]
- Cao Y, Yang H, Huang Y, Lu J, Du H, et al. Mesenchymal stem cell-derived exosomal miR-26a induces ferroptosis, suppresses hepatic stellate cell activation, and ameliorates liver fibrosis by modulating SLC7A11. Open Med 2024;19:20240945. [PMID: 38756248 DOI: 10.1515/med-2024-0945]
- Fujioka T, Dawson DR, Wright R, Honjo K, Chen JL, et al. The effects of music-supported therapy on motor, cognitive, and psychosocial functions in chronic stroke. Ann N Y Acad Sci 2018;1423:264-74. [PMID: 29797585 DOI: 10.1111/nyas.13706]
- Forte R, Tocci N, De Vito G. The impact of exercise intervention with rhythmic auditory stimulation to improve gait and mobility in parkinson disease: an umbrella review. Brain Sci 2021;11:685. [PMID: 34067458 DOI: 10.3390/brainsci11060685]
- De La Rubia Ortí JE, García-Pardo MP, Iranzo CC, Madrigal JJC, Castillo SS, et al. Does music therapy improve anxiety and depression in Alzheimer’s patients? J Altern Complement Med 2018;24:33-6. [PMID: 28714736 DOI: 10.1089/acm.2016.0346]
- King JB, Jones KG, Goldberg E, Rollins M, MacNamee K, et al. Increased functional connectivity after listening to favored music in adults with Alzheimer dementia. J Prev Alzheimers Dis 2019;6:56-62. [PMID: 30569087 DOI: 10.14283/jpad.2018.19]
- Angelucci F, Fiore M, Ricci E, Padua L, Sabino A, et al. Investigating the neurobiology of music: brain-derived neurotrophic factor modulation in the hippocampus of young adult mice. Behav Pharmacol 2007;18:491-6. [PMID: 17762517 DOI: 10.1097/FBP.0b013e3282d28f50]
- Finsterer J. Before blaming SARS-CoV-2 for intra-cerebral aneurysm formation and rupture, alternative mechanisms need to be ruled out. Brain Hemorrhages 2022;3:214-5. [PMID: 36097507 DOI: 10.1016/j.hest.2022.09.001]
- Chen H, Deng C, Meng Z, Meng S. Research progress of targeting neuro-immune inflammation in the treatment of Alzheimer’s disease. Front Biosci (Landmark Ed) 2022;27:312. [PMID: 36472107 DOI: 10.31083/j.fbl2711312]
- Chen Y, Chen X, Luo Z, Kang X, Ge Y, et al. Exercise-induced reduction of IGF1R sumoylation attenuates neuroinflammation in APP/PS1 transgenic mice. Adv Res 2025;69:279-97. [PMID: 38565402 DOI: 10.1016/j.jare.2024.03.025]
- Li J, Ma J, Feng Q, Xie E, Meng Q, et al. Building osteogenic microenvironments with a double-network composite hydrogel for bone repair. Research (Wash D C) 2023;6:0021. [PMID: 37040486 DOI: 10.34133/research.0021]
- Yuan Y, Pan B, Tang E, Mo H, Zhu J, et al. Surgical methods of total thyroidectomy for differentiated thyroid cancer: a systematic review and Bayesian network meta-analysis. Int J Surg 2024;110:529-40. [PMID: 37916941 DOI: 10.1097/JS9.0000000000000819]
- Wu Z, Chen S, Wang Y, Li F, Xu H, et al. Current perspectives and trend of computer-aided drug design: a review and bibliometric analysis. Int J Surg 2024;110:3848-78. [PMID: 38502850 DOI: 10.1097/JS9.0000000000001289]
- Pan M, Li S, Huang T, Wang X, Dong X, et al. An advanced wood preservative based on the extract of Stellera chamaejasme root with high antifungal activity. Adv Compos Hybrid Mater 2024;7:74. [DOI: 10.1007/s42114-024-00885-8]
- Banchereau R, Cepika A-M, Banchereau J, Pascual V. Understanding human autoimmunity and autoinflammation through transcriptomics. Annu Rev Immunol 2017;35:337–70. [DOI: 10.1146/annurev-immunol-051116-052225]
- Zhao L, Hu H, Zhang L, Liu Z, Huang Y, et al. Inflammation in diabetes complications: molecular mechanisms and therapeutic interventions. MedComm 2024;5:e516. [PMID: 38617433 DOI: 10.1002/mco2.516]
- Huo S, Xue J, Wang S, Shan H, Chen G, et al. A pilot trial of neoadjuvant pyrotinib plus trastuzumab, dalpiciclib, and letrozole for triple-positive breast cancer. MedComm 2024;5:e505. [PMID: 38469548 DOI: 10.1002/mco2.505]
- Xin Y, Gao C, Wang L, Liu Q, Lu Q. Lipopolysaccharide released from gut activates pyroptosis of macrophages via Caspase 11-Gasdermin D pathway in systemic lupus erythematosus. MedComm 2024;5:e610. [PMID: 38881675 DOI: 10.1002/mco2.610]
- Mohan CD, Rangappa KS, Sethi G. Transmembrane protein 25 abrogates monomeric EGFR-driven STAT3 activation in triple-negative breast cancer. MedComm 2024;5:e492. [PMID: 38532948 DOI: 10.1002/mco2.492]
- Wang J, Hu Y, Zhao K, Fan J, Zhu J, et al. Comprehensive analysis of the expression of cell adhesion molecules genes in hepatocellular carcinoma and their prognosis, and biological significance. Front Biosci (Landmark Ed) 2024;29:76. [PMID: 38420809 DOI: 10.31083/j.fbl2902076]
- Tragoonlugkana P, Chitchongyingcharoen N, Pruksapong C, Hassan S, Ngamkham K, et al. The use of human platelet lysate as a coating substance for adipose-derived stem cell expansion. Front Biosci (Landmark Ed) 2024;29:88. [PMID: 38420820 DOI: 10.31083/j.fbl2902088]
- Sheng T, Feng Q, Luo Z, Zhao S, Xu M, et al. Effect of phase clustering bias on phase-amplitude coupling for emotional EEG. J Integr Neurosci 2024;23:33. [PMID: 38419437 DOI: 10.31083/j.jin2302033]
- Guo L, Ma C, Wang M, Du Y. Tropisetron ameliorated cyclophosphamide-induced hemorrhagic cystitis via restraining TLR-4/NF-κB and JAK1/STAT3 signaling pathways. Arch Esp Urol 2023;76:56-64. [PMID: 36914420 DOI: 10.56434/j.arch.esp.urol.20237601.5]