Single-Cell Omics Analysis Reveals Critical Cell Subtypes and their Functional Attributes in Myocardial Ischemia-Reperfusion Injury
1First Clinical Medical College, Shandong University of Traditional Chinese Medicine, 16369 Jingshi Rd, Jinan 250014, China
2Xinxiang Medical University, Xinxiang 453003, China
3Department of Cardiovascular Diseases, Affiliated Hospital of Shandong University of Traditional Chinese Medicine, 16369 Jingshi Rd, Jinan 250014, China
aequal contributions as cofirst authors: Wenyang Nie, Junhao Yan and Zibo Xie.
*Correspondence to: Yong Wang, Department of Cardiovascular Diseases, Affiliated Hospital of Shandong University of Traditional Chinese Medicine, 16369 Jingshi Rd, Jinan, China, E-mail: 71002050@sdutcm.edu.cn; Zhen Wang, Department of Cardiovascular Diseases, Affiliated Hospital of Shandong University of Traditional Chinese Medicine, 16369 Jingshi Rd, Jinan, China, E-mail: 71000799@sdutcm.edu.cn; Zhenzhen Zhao, First Clinical Medical College, Shandong University of Traditional Chinese Medicine, 16369 Jingshi Rd, Jinan, China, E-mail: 2023110795@sdutcm.edu.cn
Received: September 3 2025; Revised: November 28 2025; Accepted: January 6 2026; Published Online: February 5 2026
Cite this paper:
Nie W, Yan J, Xie Z et al. Single-Cell Omics Analysis Reveals Critical Cell Subtypes and their Functional Attributes in Myocardial Ischemia-Reperfusion Injury. BIO Integration 2026; 7: 1–15.
DOI: 10.15212/bioi-2025-0159. Available at: https://bio-integration.org/
Download citation
© 2026 The Authors. This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See https://bio-integration.org/copyright-and-permissions/
Abstract
Cardiac cell death and myocardial fibrosis after ischemia-reperfusion (I/R) injury are the primary causes of impaired cardiac function. Despite advances in therapies targeting ischemia, fibrosis, and angiogenesis, the early molecular and cellular mechanisms driving I/R injury remain poorly defined, particularly regarding interactions between specific cardiomyocyte (CM) subtypes and stromal cells. Extensive signaling interactions have been demonstrated between myocardial cells and fibroblasts during injury repair and remodeling, but the dynamic characteristics and molecular pathways involved in early I/R stages remain to be fully elucidated. Herein, we used single-cell transcriptomics to identify a key subtype of CMs, termed C0 Atcayos+ CMs, that are activated in I/R. Components of the Bmp6–(Bmpr1a+Bmpr2) and Fgf1–Fgfr1 signaling axes were highly expressed and mediated interactions between the C0 Atcayos+ CM subtype and fibroblasts. These pathways are known to promote angiogenesis and regulate endothelial homeostasis, and to be crucial in inhibiting myocardial fibrosis. Ligand-receptor interaction network visualization suggested that communication between C0 Atcayos+ CMs and fibroblasts might be a critical link in the transition from myocardial cell death to fibrosis after I/R. Targeting these signaling axes might therefore offer new strategies to impede fibrosis progression and improve cardiac function after I/R. This research provides a potential reference for inhibiting the progression of diseases such as myocardial fibrosis after I/R.
Keywords
Cardiomyocytes, ischemia/reperfusion, myocardial fibrosis, scRNA-seq, transcriptional regulatory.
Introduction
I/R injury, a common complication after reperfusion therapy for acute myocardial infarction and other coronary artery diseases [1], substantially diminishes the clinical benefits of revascularization [2]. The pathological process of I/R injury involves multiple forms of cell death, including apoptosis, necrosis, ferroptosis, and pyroptosis. These processes, driven by reactive oxygen species (ROS) production, calcium overload, and inflammatory cascades, lead to CM loss and adverse remodeling [3]. NLRP3 inflammasomes are activated early during reperfusion by CMs and vascular endothelial cells, and subsequently trigger pyroptosis and amplify inflammation [4]. Mitochondrial dysfunction and impaired mitophagy, particularly in the PINK1/Parkin pathway, worsen oxidative stress and cell death [5]. Collectively, I/R injury is not merely an acute damage event but a complex pathological process involving multiple cell types and interconnected signaling networks.
After myocardial injury, quiescent cardiac fibroblasts are activated and transdifferentiate into myofibroblasts, which secrete large amounts of collagen that form fibrotic scars and help maintain structural integrity [6]. However, persistent fibroblast activation decreases ventricular compliance, impairs electrical conduction, and increases arrhythmia risk [7]. During the acute inflammatory phase, specific immune cells, such as S100a9hi macrophages, promote fibroblast-to-myofibroblast transition via the TGF-β/Smad3 pathway and facilitate macrophage-to-myofibroblast transition, thereby exacerbating fibrosis [8]. Fibroblasts exhibit substantial heterogeneity; for example, subtypes originating from the endocardium display high proliferative ability post-injury and can drive pathological fibrosis via the Wnt/β-catenin pathway [9]. Moreover, activation of the TGF-β-Smad2/3 signaling axis acts as a central driver of fibrosis, by markedly enhancing collagen deposition and extracellular matrix remodeling [10]. Therefore, post-I/R fibrosis is orchestrated by specific fibroblast subtypes and their associated signaling pathways.
Although fibroblasts play a central role in I/R-induced fibrosis, CMs remain the key cell type for maintaining and restoring cardiac function. Notably, microRNAs regulate hypertrophy, cell death, and cardiovascular disease progression, and consequently influence the development of I/R injury [11]. Furthermore, alterations in electrical coupling between CMs and fibroblasts increase excitability and heighten arrhythmia risk [7]. Therefore, we reasoned that integrating studies of fibroblast pro-fibrotic activity with CMs’ functional characteristics might facilitate precise identification of critical injury mechanisms at the cellular level in I/R injury. Herein, we applied single-cell transcriptomics to comprehensively characterize the functional features, subtype heterogeneity, and interactions between CMs and fibroblasts after I/R injury. The novelty of this work lies in the single-cell–level identification of distinct CM subtypes in the early phase of I/R injury, elucidation of their functional states and potential regulatory networks, and proposal of new therapeutic targets to prevent CM death and fibrotic transformation.
Methods
Data source and processing
We analyzed a single-cell RNA-sequencing (scRNA-seq) (10× Genomics) dataset from the GEO database (GSE227088, https://www.ncbi.nlm.nih.gov/geo/). Our re-analysis of the data was restricted to the sham (n=3) and ischemia/reperfusion (I/R, n=3) groups, whereas the pharmacologic Shexiang Baoxin pill cohort was excluded to avoid treatment confounding. In the sham group, mice received 7 days of vehicle gavage and underwent needle passage without ligation; in the I/R group, mice received 7 days of vehicle gavage, 30-min ligation of the left anterior descending coronary artery, and 24-h reperfusion before tissue collection. Raw data were processed in R (v4.1.3) with Seurat (v4.1.1). Putative doublets were removed with DoubletFinder (v2.0.3). Low-quality nuclei were filtered with standard thresholds: 300–4,000 detected genes (nFeature_RNA), 500–15,000 UMIs (nCount_RNA), mitochondrial gene fraction <20%, and erythrocyte-associated gene fraction <5%.
Dimensionality reduction, clustering, and cell type annotation
Gene expression levels were normalized with log (x + 1) transformation, with counts scaled to transcripts per 10,000 per cell. Highly variable genes (top 2,000) were identified, and principal component analysis was subsequently performed. The Harmony algorithm was applied to mitigate batch effects across samples. The top 30 principal components were selected for downstream clustering, and the results were visualized in two dimensions with UMAP.
For cell type annotation, cluster identities were assigned according to the expression of canonical markers (for CMs: TNNT2 and ACTC1; for fibroblasts: COL1A1, and PDGFRA; for endothelial cells: PECAM1 and KDR/FLT1; for smooth muscle cells: ACTA2 and TAGLN; for macrophages: LST1a and C1QA/B). Cross-validation was conducted against the CellMarker database (v2.0) and curated literature. Clusters were labeled only when most of their top differentially expressed genes were concordant with the expected lineage markers. Ambiguous clusters were left unlabeled until they were validated on the basis of additional evidence (http://xteam.xbio.top/CellMarker/), previously published studies, and established consensus. Identified CMs were further sub-clustered, and CM subtypes were defined according to differentially expressed marker genes.
Regarding gene nomenclature, we note that in mice, Atcayos is the official MGI symbol for an antisense long non-coding RNA transcribed opposite from Atcay (caytaxin). In humans, the protein-coding ortholog is ATCAY, a conserved neuronal gene implicated in Cayman ataxia; recent clinical genetics findings further support conserved locus relevance across mammals [12]. Antisense lncRNAs are an abundant class of regulatory RNAs that often act in cis on neighboring genes, and are increasingly being resolved through long-read and strand-specific transcriptomics [13, 14].
Differential expression and enrichment analysis
Differentially expressed genes (DEGs) for each cell cluster were identified with the Find Markers function in Seurat, on the basis of the Wilcoxon rank-sum test with default parameters. DEGs were defined as genes with logFC > 0.25 that were expressed in >25% of cells in each cluster. Functional enrichment analysis of DEGs, performed with the cluster Profiler (v4.6.2) and SCP (v0.4.8) packages, included Gene Ontology (GO) biological process (BP) analysis and gene set enrichment analysis (GSEA). In addition, AUCell was applied to estimate the activity of predefined gene sets within individual cells.
Trajectory analysis with slingshot and CytoTRACE
To investigate lineage dynamics and differentiation states of CM subtypes after I/R injury, we reconstructed developmental trajectories with the Slingshot package (v2.6.0). Lineage hierarchies and pseudotime curves were inferred with the getLineages and getCurves functions. Differentiation states were further validated and ranked with CytoTRACE, to provide insights into CM subtype developmental heterogeneity.
Pseudotime analysis of CM subtypes
We applied Monocle2 (v2.24.0) to reconstruct pseudotime trajectories of CMs, to capture dynamic transcriptional changes during differentiation after I/R injury. This analysis was aimed at revealing temporal cellular transitions along CM developmental pathways.
Cell–cell communication analysis
Cell–cell interactions were inferred from scRNA-seq data with the CellChat package (v1.6.1). Global intercellular communication networks were visualized, with a focus on interactions of key CM subtypes with fibroblasts, endothelial cells, neutrophils, and macrophages. CellChatDB.human served as the reference ligand–receptor interaction database. Statistical significance was assessed with a p-value threshold of 0.05.
SCENIC analysis
To reconstruct transcriptional regulatory networks and identify stable cell states, we conducted SCENIC analysis. Specifically, we used PySCENIC (v0.10.0) in Python (v3.7) with default parameters. AUCell matrices were generated to evaluate transcription factor enrichment and regulatory activity across cell subtypes.
Results
Cell clustering and functional annotation of the I/R and sham groups
After batch correction of cells from the I/R and sham groups in the GSE227088 dataset (Figure 1), 32,008 high-quality cells were obtained. On the basis of previous studies and established cell type–specific differential expression genes, we performed dimensionality reduction clustering and projected the data onto UMAP. The cells were classified into CMs; fibroblasts; ECs; T or NK cells; conventional dendritic cells (cDC1); neutrophils; Schwann cells; macrophages; and B or plasma cells. The top five marker genes for each cell type are shown in Figure 2A and B.
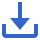
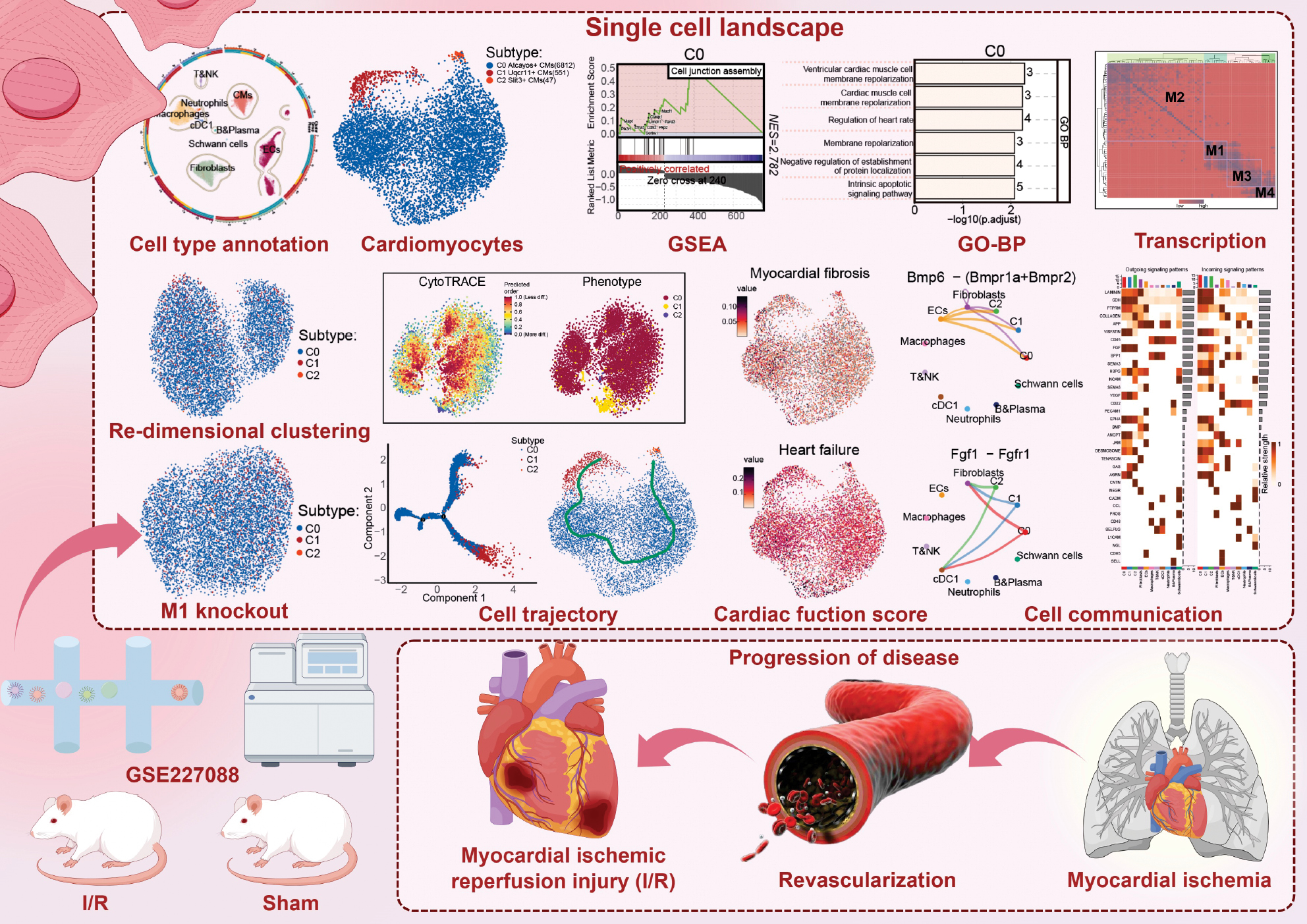
Figure 1 Overview of the research.
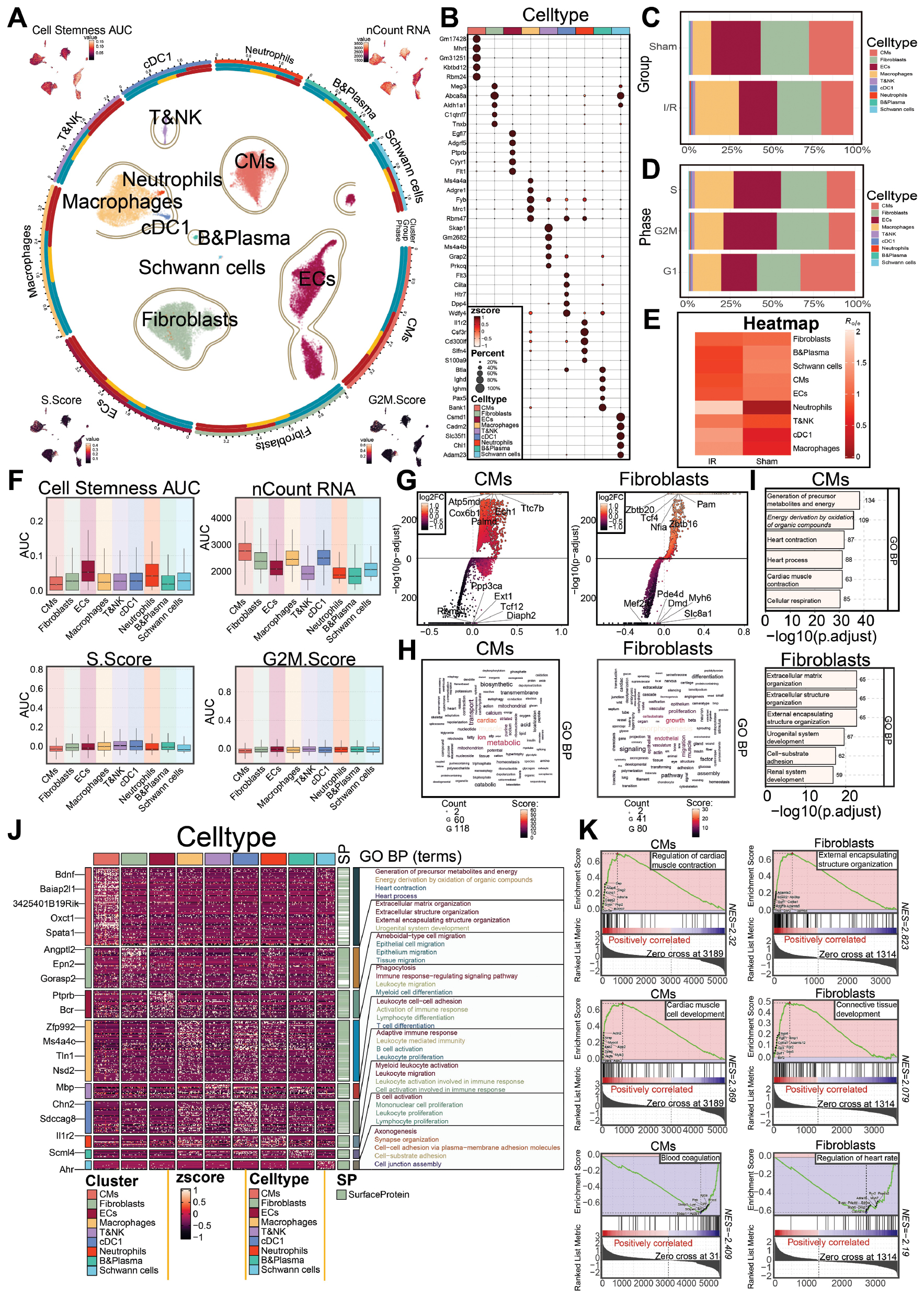
Figure 2 Cell subtype analysis and characterization of the I/R and sham groups. A: Loop graph (left) illustrates the clustering of various cell types after I/R. The four UMAP plots (arranged clockwise starting from the upper left corner) depict the distribution of cell stemness, nCount RNA, G2/MG2/M.Score, and S.Score across all cell types. B: Bubble plot shows the proportions and expression levels of the top five marker genes across cell types. C, D: Proportional bar charts display the distribution of cell types between the sham and I/R groups (C), and between S, G2/MG2/M, and G1 phases (D). E: Heatmap illustrates the Ro/e values of cell types in the sham and I/R groups. F: Boxplots compare the differences among all cell types, according to four metrics: cell stemness AUC, nCount RNA, S.Score, and G2/M.Score. G: Volcano plots display DEGs between CMs and fibroblasts in the I/R and sham groups. H: Word cloud plots from GO enrichment analysis depict the major biological processes involving DEGs in CMs and fibroblasts. I: Bar charts show enriched GO-BP terms for DEGs in CMs and fibroblasts. J: Heatmap shows the expression levels of key surface protein genes associated with cardiovascular function and immune responses across cell types, alongside GO functional annotations. K: GSEA showed the enrichment trends of differentially expressed gene sets in CMs and fibroblasts. Both positively and negatively enriched pathways are indicated.
We further visualized the proportions of each cell type in the I/R and sham groups with UMAP. Among the cell types, T or NK cells, cDC1, neutrophils, and macrophages were more prevalent in the I/R group, whereas ECs, Schwann cells, and B or plasma cells were more abundant in the sham group (Figure 2C). Heatmap analysis revealed higher neutrophil gene expression in the I/R group than the sham group (Figure 2E). The higher prevalence of neutrophils and other inflammatory cells in the I/R group suggested that leukocyte-mediated inflammatory responses contribute to I/R injury.
Compared with the sham group, CMs, fibroblasts, ECs, and macrophages showed a higher proportion of cells distributed across the G1, S, and G2/M phases. In contrast, compared with the sham group, all cell types exhibited lower S.Score and G2/M.Score values, likely reflecting leukocyte infiltration and inflammation-associated ischemia, which impair energy metabolism and lead to cellular injury [15] (Figure 2A, D, F). Among non-leukocyte cell types, fibroblasts displayed relatively higher stemness scores compared with several other cell types. In addition, fibroblasts showed a higher nCountRNA score than most other cell types, except CMs (Figure 2A and F).
We then presented the top five upregulated and downregulated genes for CMs and fibroblasts, which had high stemness and nCountRNA scores (Figure 2G). Gene expression enrichment analysis revealed that fibroblasts were associated with morphogenesis, growth, and endothelial terms, whereas CMs were associated with muscle, cardiac, and metabolic terms (Figure 2H). GO-BP analysis indicated that CMs were enriched primarily in generation of precursor metabolites and energy derivation by oxidation of organic compounds, whereas fibroblasts were enriched in extracellular matrix organization and extracellular structure organization (Figure 2I and J). GSEA revealed that upregulated genes in CMs were involved primarily in regulation of cardiac muscle contraction and cardiac muscle cell development, whereas fibroblasts were enriched in external encapsulating structure organization and connective tissue development (Figure 2K).
Therefore, during the development of I/R injury, CMs are involved primarily in cardiac contraction and development, whereas fibroblasts play a role in tissue remodeling.
Single-cell profiling of CMs reveals subtype heterogeneity and functional enrichment during I/R injury
We performed batch correction and sequential processing of 7,410 CMs from both the I/R and sham groups, followed by dimensionality reduction, clustering, and projection onto UMAP. Three distinct and heterogeneous CM subtypes were identified and designated as C0 Atcayos+ CMs, C1 Uqcr11+ CMs, and C2 Slit3+CMs, according to their specific marker gene expression (Figure 3A). Among these, C0 Atcayos+ CMs were the predominant subtype in both groups and had a higher relative proportion in the I/R group than the sham group. In contrast, the proportions of C1 Uqcr11+ CMs and C2 Slit3+ CMs were consistently lower in the I/R group than the sham group (Figure 3B).
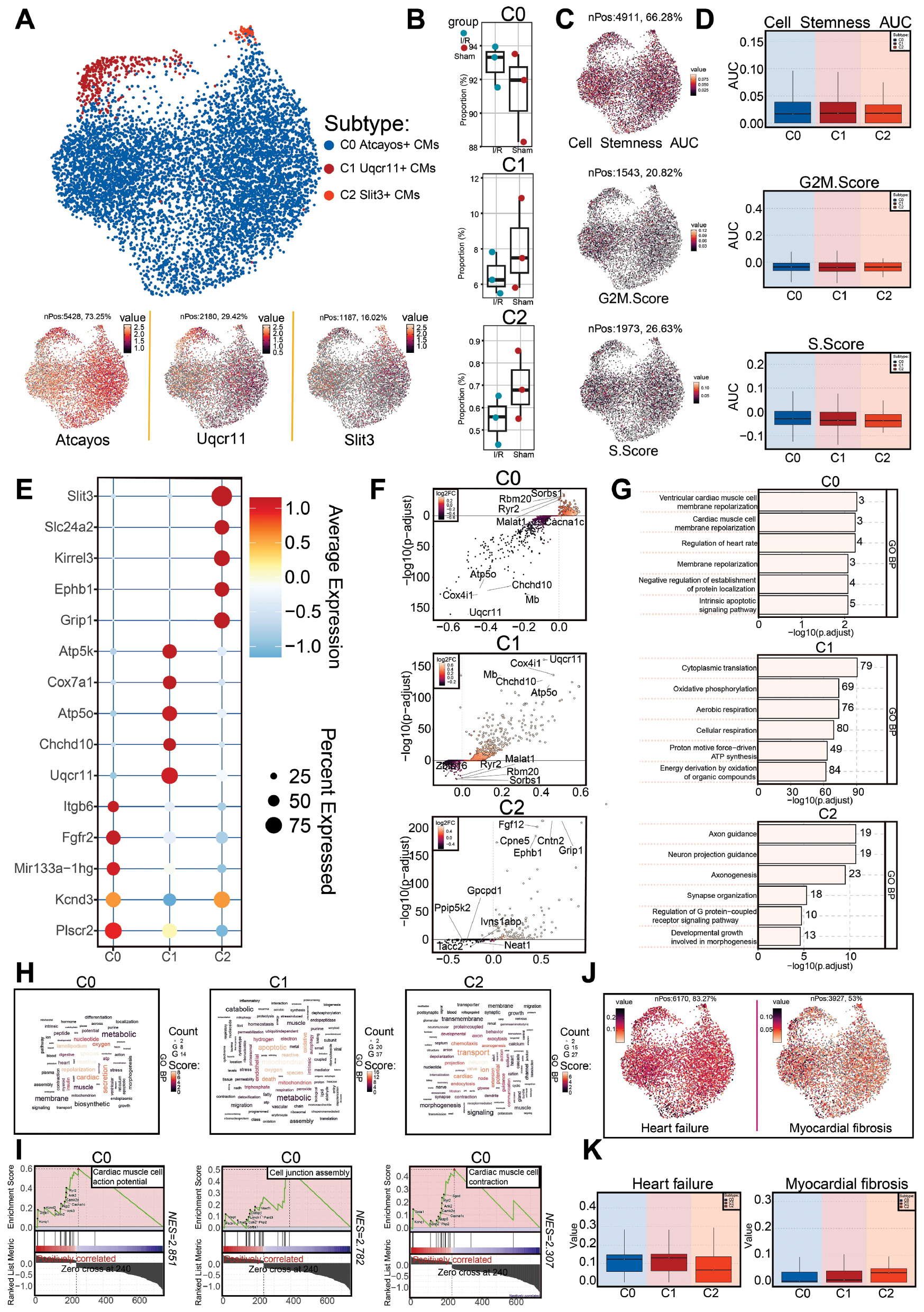
Figure 3 Heterogeneity analysis of CM subtypes and functional annotation of differentially expressed genes. A: CMs were clustered into the following subtypes: C0 (high expression of Atcayos), C1 (high expression of Uqcr11), and C2 (high expression of Slit3). UMAP plots show their spatial distribution and key gene expression. B: Box plots compare the distribution of the three subtypes in the sham and I/R groups. C, D: UMAP plot (A) and box plot (B) show the expression levels of cell stemness (cell stemness AUC), G2/M score (G2/M.Score), and S phase score (S.Score) for the three subtypes. E: Bubble chart shows the expression levels and proportions of five marker genes for the three subtypes. F: Volcano plot displays the differentially expressed genes (DEGs) within each subtype. G: Bar chart shows the enrichment status of different functional pathways in C0, C1, and C2 subtypes, according to GO enrichment analysis. H: Word cloud displays the major biological processes in GO enrichment analysis for C0, C1, and C2 subtypes. I: GSEA reveals the positively and negatively correlated pathways enriched in C0, C1, and C2 subtypes. J: Distribution of cell scores associated with heart failure and myocardial fibrosis in each CM subtype. K: Box plots further compare the differences in heart failure and myocardial fibrosis scores among C0, C1, and C2 subtypes.
Next, we evaluated stemness, G2/M phase, and S phase scores across the three subtypes (Figure 3C and D). We also identified the top five marker genes for each subtype (Figure 3E). For C1 Uqcr11+ CMs, these included Atp5k, Cox7a1, Atp5o, Chchd10, and Uqcr11. DEGs (upregulated and downregulated) in the three subtypes were further visualized with volcano plots (Figure 3F).
To gain deeper insight into the biological functions and differences among subtypes, we conducted enrichment analyses. GO-BP analysis revealed that C0 Atcayos+ CMs were enriched primarily in terms such as ventricular cardiac muscle cell membrane repolarization, regulation of heart rate, negative regulation of protein localization, and intrinsic apoptotic signaling pathway, and showed strong associations with cardiac function and repolarization (Figure 3G and H). GSEA further indicated that C0 Atcayos+ CMs were enriched in cardiac muscle cell action potential, cell junction assembly, and cardiac muscle cell contraction (Figure 3I). These findings suggested that C0 Atcayos+ CMs might regulate heart rate, apoptotic signaling, and myocardial contraction, thereby contributing to myocardial injury [3, 16, 17].
Finally, to investigate the critical role of this key subtype in I/R-associated heart failure and myocardial fibrosis, we assessed heart failure and fibrosis scores across the three subtypes. C0 Atcayos+ CMs, together with C1 Uqcr11+ CMs, exhibited higher heart failure scores than C2 Slit3+ CMs, suggesting their potential involvement in heart failure. In contrast, fibrosis scores were relatively lower and were comparable across all three subtypes (Figure 3J and K).
Transcription factor regulatory differences in CM subtypes
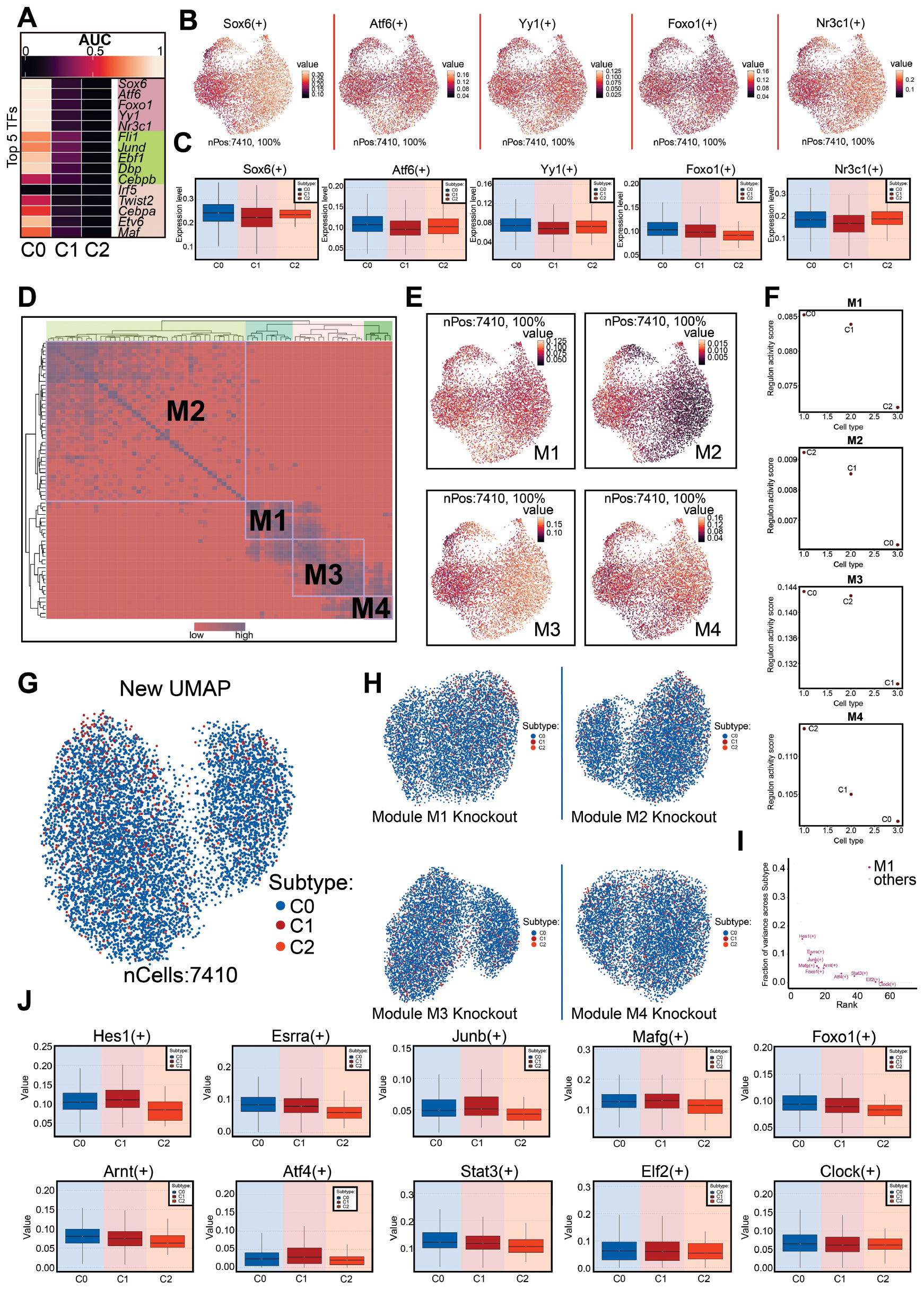
To explore transcriptional regulatory differences among subtypes, we performed pySCENIC analysis and identified the top five transcription factors (TFs) for each subtype (Figure 4A). The expression patterns of the top five TFs in C0 Atcayos+ CMs across the subtypes were further illustrated with UMAP and box plots. (Figure 4B and C). We used SCENIC to identify the transcriptional regulatory modules. The CM transcriptional regulatory network was divided into four major modules (M1, M2, M3, and M4) (Figure 4D). The average activity scores of each module were mapped to the UMAP plot (Figure 4E). Scatter plots revealed that the TFs in C0 Atcayos+ CMs were associated primarily with the M1 and M3 modules (Figure 4F). To further assess the contributions of these modules, we performed a second round of dimensionality reduction and projected the data onto a new UMAP after selectively removing each module. Removal of the M1 module markedly altered the clustering structure, whereas removal of the M3 module had only a minor effect (Figure 4G and H). We then examined the TFs with higher expression in the M1 module than the other three modules, and observed that their expression levels remained consistently high in C0 Atcayos+ CMs (Figure 4I and J). Together, these findings suggested that the molecular features of the M1 module at least partly represented the transcriptional characteristics of C0 Atcayos+ CMs.
Figure 4 TF regulatory network and modular dynamics analysis. A: Heatmap shows the expression of the top five TFs in the three subtypes. B, C: UMAP plot (B) and box plot (C) display the expression distribution of high-expression TFs (Sox6, Atf6, Yy1, Foxo1, and Nr3c1) in the C0 subtype. D: Mutual information heatmap reveals co-expression modules (M1–M4) in CMs, with colors indicating similarity. E: UMAP plot shows the distribution and expression levels of four modules (M1–M4) in the three subtypes. F: Schematic diagram of the average activity scores of modules M1–M4 across subtypes. G, H: New UMAP plot (G) after re-clustering of CM subtypes and differential clustering analysis with sequential removal of the M1, M2, M3, and M4 modules, thus generating four UMAP plots (H). I: Visual comparison of differential expression of M1 module characteristic TFs with other modules. J: Box plot shows the distribution of M1 module activity scores compared with those for other modules (M2, M3, and M4) in different subtypes (Hes1, Esrra, Junb, Mafg, Foxo1, Arnt, Atf4, Stat3, Eif2, and Clock).
Pseudotime trajectory of CM subtypes and crosstalk with fibroblasts
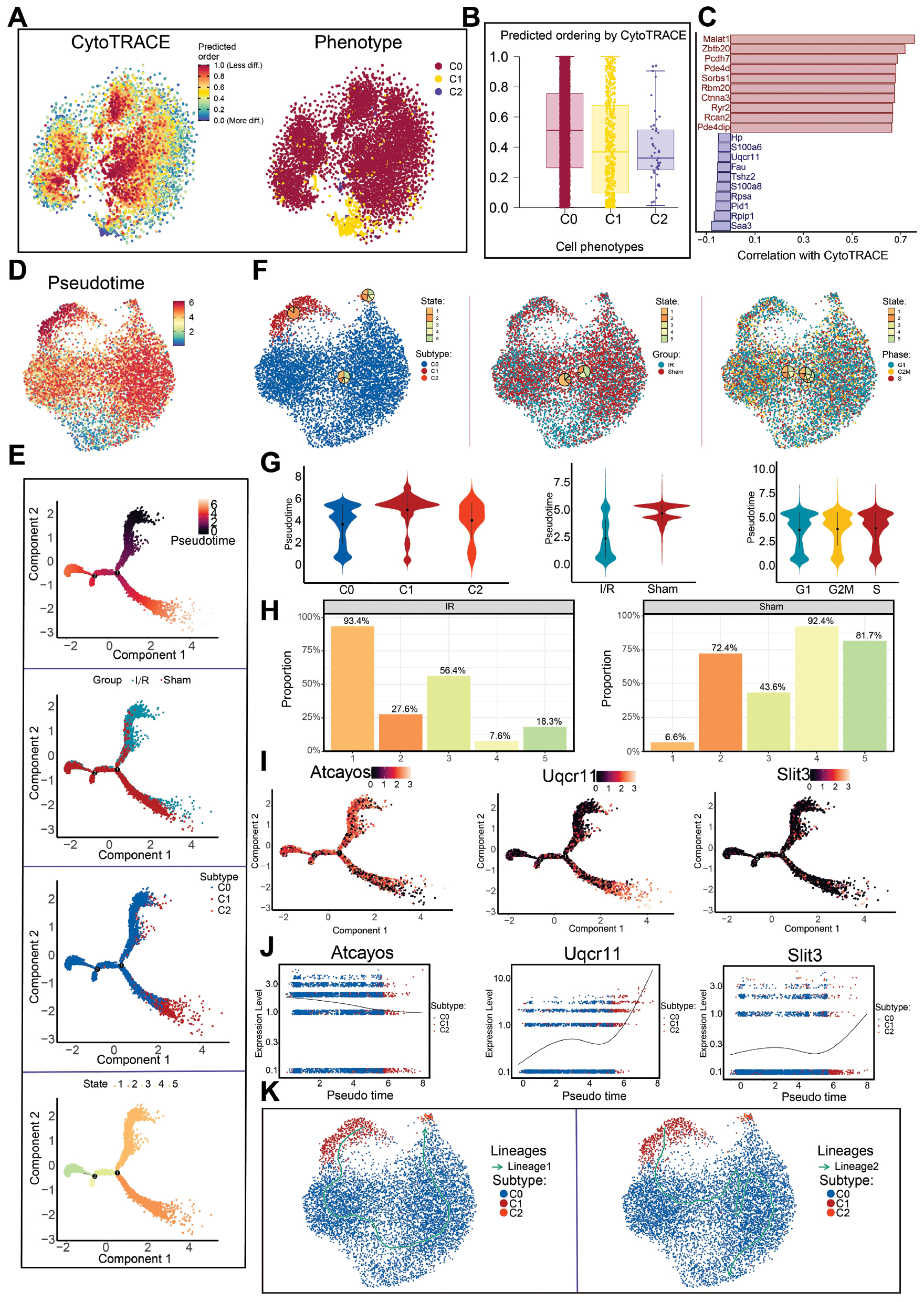
We first applied CytoTRACE to evaluate the differentiation potential of the three CM subtypes. The analysis revealed higher CytoTRACE scores in C0 Atcayos+ CMs than in C1 Uqcr11+ CMs and C2 Slit3+ CMs (Figure 5A and B), thus indicating a less differentiated transcriptional state of C0 Atcayos+ CMs. Notably, C0 Atcayos+ CMs displayed the lowest differentiation potential and therefore might potentially promote the early phase of I/R injury. Differential expression analysis demonstrated marked activation of cardiac structural and functional gene programs, whereas genes associated with metabolic and stress-response pathways were selectively downregulated (Figure 5C).
Figure 5 Single-cell trajectory inference reveals the differentiation dynamics of CM subtypes. A: CytoTRACE prediction results and cell phenotype distribution. Left: cell differentiation potential; right: cell phenotype distribution. B: Box plot presents the relationship between CytoTRACE predicted cell ordering and cell phenotype. C: Bar chart displays the expression of upregulated and downregulated differential genes in CytoTRACE prediction. D: Monocle2 pseudotime trajectory analysis, with a UMAP plot showing the developmental paths from the starting state toward different differentiation directions, with colors representing pseudotime progression. E: Pseudotime trajectory plot showing the pseudotime analysis results based on groups, subtypes, and states. F: Distribution of CM cells across subtypes, groups, and phases (G1, G2/M, and S). G: Violin plot displays the pseudotime distribution differences across subtypes, groups, and phases (G1, G2/M, and S). H: Proportions of CMs in the I/R and sham groups at the indicated time points. I: Expression of Atcayos Uqcr11 and Atcayos Slit3 genes across cell lineages, subtypes, and pseudotime stages. J: Expression distribution of developmental-associated genes (Atcayos, Uqcr11, and Slit3) along the pseudotime trajectory, revealing their dynamic expression patterns during subtype transitions. K: Developmental trajectory of subtypes, with Lineage1 on the left and Lineage2 on the right.
To elucidate the developmental continuum, we reconstructed a pseudotime trajectory with Monocle2. The results indicated a progression from top to bottom, featuring two branch points (nodes 1 and 2) that delineated five transcriptional states: state 1 (early); state 3 (intermediate); and states 2, 4, and 5 (late) (Figure 5D and E). UMAP and violin plots showed that C0 Atcayos+ CMs were located predominantly in the early stage, and were followed by C2 Slit3+ CMs, whereas C1 Uqcr11+ CMs were enriched in the later stages (Figure 5F and G). In agreement with these observations, cells from the I/R group were concentrated in the early stage, whereas those from the sham group were found predominantly in the later stages (Figure 5G and H).
Pseudotime analysis further revealed that the signature gene Atcayos was highly expressed in C0 Atcayos+ CMs during the earliest state, and its expression gradually decreased along the trajectory (Figure 5E, I, J). A small subtype of C1 Uqcr11+ CMs appeared at the initial segment, although most of these cells were distributed in later stages, whereas C2 Slit3+ CMs minimally contributed to the early states. Slingshot-based trajectory reconstruction identified C1 Uqcr11+ CMs as the computational starting point, according to global transcriptional similarity patterns. Notably, although C0 Atcayos+ CMs were not positioned at the trajectory origin, they represented the predominant early responding population during I/R, as evidenced by their high CytoTRACE scores and enrichment in early stress-response programs (Figure 5K). Therefore, the trajectory origin reflects algorithmic initialization within transcriptional manifolds rather than a true developmental lineage starting point. This temporal sequence highlights the biological relevance of selecting C0 Atcayos+ CMs as a key focus for investigating the early events in I/R injury.
Cell–cell communication networks highlight functional interactions between C0 Atcayos+ CMs and fibroblasts
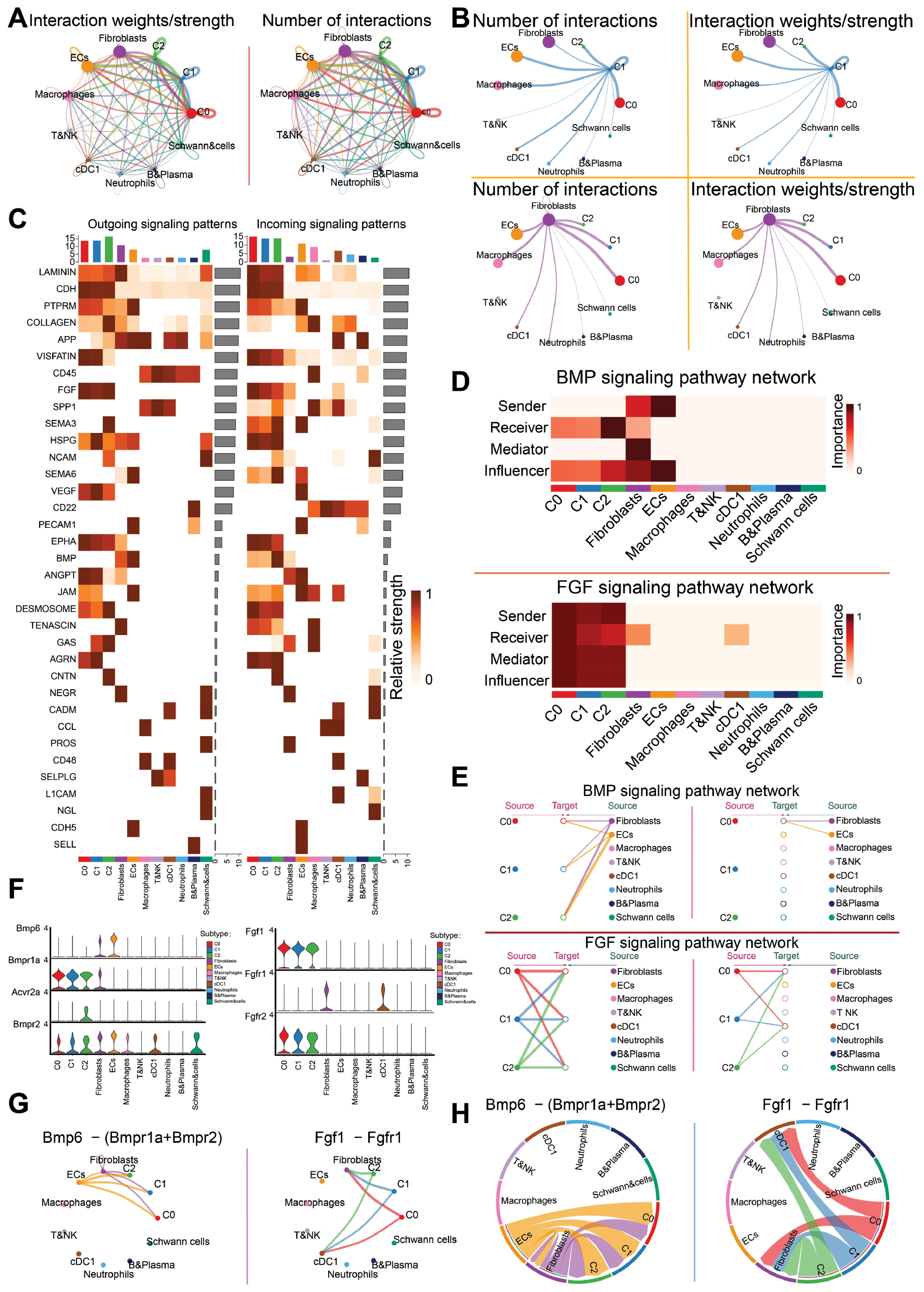
To investigate intercellular communication in the I/R and sham groups, we first conducted circle graph analysis to visualize the overall interaction strength and frequency among all cell types (Figure 6A). C0 Atcayos+ CMs exhibited extensive communication networks with fibroblasts, macrophages, endothelial cells, and neutrophils. Among these, interactions between C0 Atcayos+ CMs and fibroblasts were particularly prominent, in agreement with their close functional association with cardiac remodeling and repair after I/R injury. Given the critical roles of fibroblasts in extracellular matrix regulation and structural remodeling, subsequent analyses were focused on C0 Atcayos+ CM–fibroblast communication (Figure 6B). Receptor expression in receiver cells and ligand expression in sender cells were then visualized as heatmaps to delineate the signaling landscape (Figure 6C).
Figure 6 Signaling pathway interaction modes and functional regulation among CM subtypes. A: Total cell communication network among subtypes, showing the interaction strength and number of interactions for each cell type. B: Intercellular interactions. Left top and left bottom show interaction quantity between the C1 subtype and fibroblasts. Right top and right bottom show interaction weights between the C1 subtype and fibroblasts. C: Heatmap depicts the relative strengths of incoming (receiving) and outgoing (sending) signaling activities of different pathways across CM subtypes. D, E: BMP and FGF signaling networks. Panel D illustrates the roles of different cell types as signal senders and receivers, as well as their relative contributions to pathway-level signal transduction and downstream functional effects. Panel E shows the BMP-associated (top) and FGF-associated (bottom) signaling networks. F: Representative ligand-receptor genes in the BMP and FGF pathways in various cell types. G: Interactions among cells in the Bmp6–(Bmpr1a+Bmpr2) and Fgf1−Fgfr1 signaling pathways. Left: Bmp6-associated pathways; right: Fgf1-associated pathways. H: Chord diagram further visualizes the communication directions and strengths of Bmp6 and Fgf1 signaling pathways among cell types.
Cell–cell communication network analysis identified two key signaling pathways mediating this interaction. In the BMP signaling network, fibroblasts were predicted to act as the source, whereas C0 Atcayos+ CMs served as the target. In contrast, within the FGF signaling network, C0 Atcayos+ CMs functioned as the source, and fibroblasts served as the target (Figure 6D and E). To further characterize these pathways, we examined ligand–receptor expression patterns. In the Bmp6–(Bmpr1a+Bmpr2) axis, fibroblasts expressed predominantly Bmp6, whereas C0 Atcayos+ CMs expressed high levels of Bmpr1a and Bmpr2. These findings suggested a potential regulatory influence of fibroblast-derived BMP signals on this CM subtype. In contrast, in the Fgf1–Fgfr1 axis, C0 Atcayos+ CMs exhibited elevated Fgf1 expression, whereas fibroblasts expressed Fgf1 at high levels, thus indicating a reciprocal regulatory interaction mediated by FGF signaling (Figure 6F–H). Together, these results highlighted a bidirectional communication between C0 Atcayos+ CMs and fibroblasts orchestrated through BMP and FGF signaling pathways.
Discussion
I/R injury is a common and severe pathological process in coronary artery disease, and CM death is among its core features. Beyond classical apoptosis and necrosis, newly identified forms of regulated cell death, including ferroptosis, necroptosis, and pyroptosis, have been found to be closely associated with I/R injury. These mechanisms exacerbate CM loss by promoting ROS generation, inducing calcium stress, and activating inflammatory responses, thereby driving adverse myocardial remodeling, functional impairment, and heart failure [3, 18–21]. The extensive loss of CMs and the accompanying inflammatory milieu create a pathological environment that triggers subsequent myocardial remodeling and fibrotic responses. After I/R injury, the transition of the myocardium from acute inflammation to fibrosis is a key determinant of prognosis [8]. Myocardial fibrosis not only is a final common pathway in various cardiac diseases but also is tightly associated with the activation of cardiac fibroblasts. Under physiological conditions, fibroblasts participate in extracellular matrix homeostasis; however, under injury, they transdifferentiate into myofibroblasts, thus promoting excessive collagen deposition, chamber dilation, CM hypertrophy, and further cell death, and ultimately leading to heart failure. Despite the importance of fibrosis in cardiovascular disease, incomplete understanding of the precise mechanisms underlying fibroblast-mediated fibrosis in I/R injury has limited the development of targeted therapeutic strategies [6, 22, 23]. Moreover, therapeutic approaches targeting signaling pathways broadly involved in CM death and fibrosis have yet to achieve substantial clinical translation [4, 24]. Therefore, we aimed to identify key pathways and potential therapeutic targets capable of preventing CM death and fibrosis after I/R injury through single-cell analysis.
We first observed that the proportion of CMs was reduced in the I/R group, and cells were mainly distributed in the G1 phase, indicating a reduced proliferative capacity. Simultaneously, neutrophils, key effector cells in acute inflammation, had Ro/e values markedly greater than 1 in the I/R group but less than 1 in the sham group, thus suggesting an unexpectedly high level of neutrophil enrichment in I/R injury. This cellular finding established a basis for the transition from inflammation to fibrosis after I/R injury. In I/R injury, inflammatory cells exert complex and diverse effects: neutrophils and monocytes exacerbate local tissue injury by activating pro-inflammatory responses, whereas dendritic cells and regulatory T cells alleviate excessive inflammation and promote tissue repair through the secretion of anti-inflammatory cytokines such as IL-10 [25–27]. Therefore, precise modulation of inflammatory cell functions is crucial for achieving a balance between protection and injury, and mitigating the myocardial damage caused by I/R.
Further enrichment analysis revealed significant CM enrichment in regulation of cardiac muscle contraction. Under ischemic conditions, CMs enter a viable but non-contractile protective state by lowering contractile energy demands, whereas after reperfusion, their transcriptional profile shifts toward contraction-regulating genes; these observations reflect a metabolic adaptation process to maintain myocardial function [23, 28]. Fibroblasts were enriched in connective tissue development. After cardiac injury, fibroblasts modulate excitability by depositing extracellular matrix and regulating electrical coupling, and consequently affect cardiac rhythm [7]. Notably, Meg3 was highly expressed in fibroblasts and was closely associated with cardiac remodeling and matrix metalloproteinase-2 (MMP-2) regulation. Silencing Meg3 suppresses Mmp-2 transcription and subsequently alleviates cardiac fibrosis [29]. Additionally, CMs exhibited high expression of Mhrt, a cardiac-specific lncRNA that antagonizes the chromatin remodeler Brg1, inhibits aberrant gene expression, and protects the heart against hypertrophy and failure under stress conditions [30, 31].
To further investigate the role of CM metabolism in I/R injury, we focused on the identified C0 Atcayos+ CM subtypes. Differentially expressed genes in C0 Atcayos+ CMs included Sorbs1, Rbm20, Ryr2, Malat1, Cacna1c, Chchd10, Atp5o, Mb, Cox4i1, and Uqcr11, which were enriched primarily in metabolic process terms such as regulation of heart rate and intrinsic apoptotic signaling pathway. Previous studies have shown that heart rate and its variability reflect disease status and can predict the risk of progression from post-perfusion injury to heart failure, including adverse outcomes such as mortality [30, 32]. We therefore hypothesized that C0 Atcayos+ CMs were associated with heart rate regulation and apoptosis in I/R injury. GSEA further revealed positive correlations with the terms cell junction assembly and cardiac muscle cell contraction. Notably, gap junction channels are critical for normal cardiac impulse propagation, and their dysfunction increases arrhythmia risk [33]. Therefore, C0 Atcayos+ CMs might influence apoptosis in I/R injury by modulating heart rate and cell–cell coupling. These findings, combined with the above results, suggested that C0 Atcayos+ CMs might enter a viable but non-contractile protective state that decreases contractile energy demands, and consequently plays a potential role in post-I/R adaptation and CM death.
Detailed scoring for heart failure and myocardial fibrosis further validated the selection of this key subtype. Although the I/R group exhibited lower myocardial fibrosis scores, because of substantial inflammatory infiltration and milder fibrosis, C0 Atcayos+ CMs showed higher heart failure scores, thus supporting the rationale for their selection.
PySCENIC analysis and visualization identified Sox6, Atf6, and Yy1 as highly expressed TFs in C0 Atcayos+ CMs. Sox6 overexpression is closely associated with cardiomyopathy and heart failure, and this TF plays a key role in CM differentiation and cardiac function regulation [34, 35]. ROS overproduction after I/R injury exacerbates myocardial infarction [36], and Atf6 contributes to this process by promoting ROS generation through endoplasmic reticulum stress responses [37]. Yy1 facilitates cardiac hypertrophy and heart failure by regulating the transcription and splicing stability of downstream genes [38]. These findings suggest that Sox6, Atf6, and Yy1 might be critical intervention points for preventing CM death and fibrotic transformation after I/R injury.
C0 Atcayos+ CMs exhibited high CytoTRACE scores, in agreement with a less differentiated (earlier-like) transcriptional state. This state should not be interpreted to reflect active proliferation, because adult CMs are generally post-mitotic; instead, it reflects stress-induced transcriptional plasticity during the early response to I/R injury. Monocle2 trajectory analysis confirmed that the early stage was composed primarily of C0 Atcayos+ CMs, and the I/R group showed a predominant presence in state 1. Trajectory analysis of the characteristic gene Atcayos showed high expression in early stages, which gradually declined over pseudotime. Similarly, Slingshot lineage analysis confirmed that C0 Atcayos+ CMs were distributed primarily in early to mid-developmental stages, a finding further supporting our hypothesis.
On the basis of these results, we speculated that C0 Atcayos+ CMs might be highly active after I/R injury and engage in specific intercellular interactions. Analysis of high-expression signaling pathways in the I/R group revealed the following: (1) In ECs, VEGF signaling promotes angiogenesis under hypoxia [39]. (2) BMP signaling is involved in CM proliferation, angiogenesis, and fibrosis-related remodeling in cardiovascular injury and regeneration [40–42]. (3) FGF signaling is associated with inflammation in heart failure, endothelial activation, and mechanotransduction, and BMP–FGF synergy drives lineage diversification of cardiac valve progenitors [43]; (4) In neutrophils, laminin signaling supports endothelial proliferation and repair post-injury [44]. All these pathways are tightly associated with angiogenesis and fibrosis regulation.
Circular and chord diagram visualizations confirmed the strong potential biological significance of BMP and FGF signaling networks between C0 Atcayos+ CMs and fibroblasts. BMP signaling promotes angiogenesis and homeostatic endothelial behavior [45], whereas FGF signaling enhances endothelial glycolysis and supports proliferation and migration during vascular development [46]. At the molecular level, we identified two key communication axes: Bmp6–(Bmpr1a+Bmpr2) and Fgf1–Fgfr1. Notably, Bmp6 is highly expressed in fibroblasts and ECs, and has been shown to inhibit myocardial fibrosis [47], and recombinant Fgf1 (rFgf1) also decreases myocardial fibrosis [48].
The hypoxic environment in early I/R injury directly triggers immune cell infiltration–driven inflammation and initiates angiogenesis [49]. This finding might explain the pro-angiogenic effects of Bmp6- and Fgf1-associated communication observed between C0 Atcayos+ CMs and fibroblasts. We therefore propose that these two communication networks are critical regulatory pathways linking CM death to fibrotic transformation after I/R injury. Given their importance in early repair, enhancing Bmp6–(Bmpr1a+Bmpr2) and Fgf1–Fgfr1 signaling might provide a promising therapeutic strategy to delay CM death, attenuate fibrosis, and ameliorate cardiac function after I/R injury.
Conclusion
Herein, we delineated a CM subtype, C0 Atcayos+ CMs, that shows early transcriptional responses to I/R injury and engages in putative crosstalk with fibroblasts through the established Bmp6–(Bmpr1a/Bmpr2) and Fgf1–Fgfr1 signaling axes. These interactions align with pro-angiogenic and fibrosis-modulating programs, and provide a mechanistic link between CM injury and subsequent remodeling. Rather than proposing new targets, our results define the cell type and temporal context in which these pathways might be leveraged to modulate early CM–fibroblast communication. Our analyses offer hypothesis-generating insights that invite confirmation with orthogonal approaches and have potential to inform more precise therapeutic strategies.
Data availability statement
Study-related original data and other data can be obtained from the corresponding author (2023110795@sdutcm.edu.cn).
Ethics statement
This analysis of data from a database does not present any ethical issues.
Author contributions
Wenyang Nie, Junhao Yan, Zibo Xie, Zhen Wang, and Zhenzhen Zhao conceived and designed the research. The data were downloaded and collected by Wenyang Nie, Yuhang Liu, and Jingwen Zhang. The article was written by Wenyang Nie and Junhao Yan, after data analysis. Quality control was performed on the articles by Yong Wang, Zhen Wang, and Zhenzhen Zhao. Zhenzhen Zhao led the submission. The final manuscript was read and approved by all authors. Wenyang Nie, Junhao Yan, and Zibo Xie are co-first authors. Yong Wang, Zhen Wang, and Zhenzhen Zhao are co-corresponding authors.
Funding
This study was funded by the Young Scientists Fund of the National Natural Science Foundation of China (No. 82104797), Natural Science Foundation of Shandong Province general project (ZR2022MH307), and Jinan Science and Technology Plan Project (202225004).
Acknowledgements
Graphical abstract and Figure 1 were created with Figdraw (ID:WISTUf6110, PIPPPdd282).
Conflict of interest
The authors declare that there are no conflicts of interest.
Graphical abstract
Highlights
Ischemia-reperfusion (I/R) injury, a frequent complication of reperfusion therapy, is a complex pathological process rather than a transient acute insult. It is driven by excessive ROS generation, calcium overload, and inflammatory signaling, and it leads to multiple forms of cell death, including apoptosis, necrosis, ferroptosis, and pyroptosis. Early NLRP3 inflammasome activation in cardiomyocytes (CMs) and endothelial cells, together with mitochondrial dysfunction and impaired PINK1/Parkin-mediated mitophagy, amplifies oxidative stress and inflammation, and ultimately results in CM loss and adverse cardiac remodeling. In this study, we used single-cell transcriptomic profiling to systematically dissect CM and fibroblast heterogeneity after I/R injury; define their dynamic functional states and intercellular interactions; and identify key regulatory networks and potential therapeutic targets involved in CM death and fibrotic remodeling during the early phase of I/R injury.
In brief
Single-cell transcriptomic analysis reveals an early-responsive cardiomyocyte subtype (C0 Atcayos+) in myocardial ischemia-reperfusion injury. This subtype exhibits prominent bidirectional communication with fibroblasts, mediated primarily through the Bmp6–(Bmpr1a+Bmpr2) and Fgf1-Fgfr1 signaling axes, providing mechanistic insight into the transition from cardiomyocyte injury to fibrotic remodeling .
References
- Wu X, Reboll MR, Korf-Klingebiel M, Wollert KC. Angiogenesis after acute myocardial infarction. Cardiovasc Res 2021;117(5):1257-73. [PMID: 33063086 DOI: 10.1093/cvr/cvaa287]
- Wang K, Li Y, Qiang T, Chen J, Wang X. Role of epigenetic regulation in myocardial ischemia/reperfusion injury. Pharmacol Res 2021;170:105743. [PMID: 34182132 DOI: 10.1016/j.phrs.2021.105743]
- Xiang Q, Yi X, Zhu XH, Wei X, Jiang DS. Regulated cell death in myocardial ischemia-reperfusion injury. Trends Endocrinol Metab 2024;35(3):219-34. [PMID: 37981501 DOI: 10.1016/j.tem.2023.10.010]
- Welt FGP, Batchelor W, Spears JR, Penna C, Pagliaro P, et al. Reperfusion injury in patients with acute myocardial infarction: JACC scientific statement. J Am Coll Cardiol 2024;83(22):2196-213. [PMID: 38811097 DOI: 10.1016/j.jacc.2024.02.056]
- Zhao X, Wang Z, Wang L, Jiang T, Dong D, et al. The PINK1/Parkin signaling pathway-mediated mitophagy: a forgotten protagonist in myocardial ischemia/reperfusion injury. Pharmacol Res 2024;209:107466. [PMID: 39419133 DOI: 10.1016/j.phrs.2024.107466]
- Travers JG, Kamal FA, Robbins J, Yutzey KE, Blaxall BC. Cardiac fibrosis: the fibroblast awakens. Circ Res 2016;118(6):1021-40. [PMID: 26987915 DOI: 10.1161/CIRCRESAHA.115.306565]
- Wang Y, Li Q, Tao B, Angelini M, Ramadoss S, et al. Fibroblasts in heart scar tissue directly regulate cardiac excitability and arrhythmogenesis. Science 2023;381(6665):1480-7.
- Shen S, Zhang M, Wang X, Liu Q, Su H, et al. Single-cell RNA sequencing reveals S100a9hi macrophages promote the transition from acute inflammation to fibrotic remodeling after myocardial ischemia‒reperfusion. Theranostics 2024;14(3):1241-59. [PMID: 38323308 DOI: 10.7150/thno.91180]
- Han M, Liu Z, Liu L, Huang X, Wang H, et al. Dual genetic tracing reveals a unique fibroblast subpopulation modulating cardiac fibrosis. Nat Genet 2023;55(4):665-78. [PMID: 36959363 DOI: 10.1038/s41588-023-01337-7]
- Khalil H, Kanisicak O, Prasad V, Correll RN, Fu X, et al. Fibroblast-specific TGF-β-Smad2/3 signaling underlies cardiac fibrosis. J Clin Invest 2017;127(10):3770-83. [PMID: 28891814 DOI: 10.1172/JCI94753]
- Carbonell T, Gomes AV. MicroRNAs in the regulation of cellular redox status and its implications in myocardial ischemia-reperfusion injury. Redox Biol 2020;36:101607. [PMID: 32593128 DOI: 10.1016/j.redox.2020.101607]
- Siavashani ES, Ashrafi MR, Ghabeli H, Heidari M, Garshasbi M. Novel homozygous frameshift variant in the ATCAY gene in an Iranian patient with Cayman cerebellar ataxia; expanding the neuroimaging and clinical features: a case report. BMC Med Genomics 2023;16(1):226. [PMID: 37752557 DOI: 10.1186/s12920-023-01643-3]
- Rojo-Carrillo JJ, Garrido-Rodríguez P, Llamas-López M, Cifuentes-Riquelme R, Padilla J, et al. Landscape of antisense genes in the human genome and identification of new human hepatic antisense RNAs by long-read sequencing. BMC Genomics 2024;25(1):1148. [PMID: 39604851 DOI: 10.1186/s12864-024-11017-3]
- Jiang B, Yuan Y, Yi T, Dang W. The roles of antisense long noncoding RNAs in tumorigenesis and development through cis-regulation of neighbouring genes. Biomolecules 2023;13(4):684. [PMID: 37189431 DOI: 10.3390/biom13040684]
- Algoet M, Janssens S, Himmelreich U, Gsell W, Pusovnik M, et al. Myocardial ischemia-reperfusion injury and the influence of inflammation. Trends Cardiovasc Med 2023;33(6):357-66. [PMID: 35181472 DOI: 10.1016/j.tcm.2022.02.005]
- Ammirati E, Moslehi JJ. Diagnosis and treatment of acute myocarditis: a review. J Am Med Assoc 2023;329(13):1098-113. [PMID: 37014337 DOI: 10.1001/jama.2023.3371]
- Sun X, Chen H, Gao R, Qu Y, Huang Y, et al. Intravenous transplantation of an ischemic-specific peptide-TPP-mitochondrial compound alleviates myocardial ischemic reperfusion injury. ACS Nano 2023;17(2):896-909. [PMID: 36625783 DOI: 10.1021/acsnano.2c05286]
- Li Y, Huang D, Jia L, Shangguan F, Gong S, et al. LonP1 links mitochondria-ER interaction to regulate heart function. Research (Wash D C) 2023;6:0175. [PMID: 37333972 DOI: 10.34133/research.0175]
- Morawin B, Tylutka A, Bielewicz F, Zembron-Lacny A. Diagnostics of inflammaging in relation to sarcopenia. Front Public Health 2023;11:1162385. [PMID: 37465171 DOI: 10.3389/fpubh.2023.1162385]
- Chen F, He Z, Wang C, Si J, Chen Z, et al. Advances in the study of S100A9 in cardiovascular diseases. Cell Prolif 2024;57(8):e13636. [PMID: 38504474 DOI: 10.1111/cpr.13636]
- Yuan J, Wang JM, Li ZW, Zhang CS, Cheng B, et al. Integration of ATAC-Seq and RNA-Seq identifies the key genes in myocardial ischemia. Genes Dis 2023;10(1):62-4. [PMID: 37013038 DOI: 10.1016/j.gendis.2022.05.013]
- Zou Y, Shan Z, Han Z, Yang J, Lin Y, et al. Regulating blood clot fibrin films to manipulate biomaterial-mediated foreign body responses. Research (Wash D C) 2023;6:0225. [PMID: 37719049 DOI: 10.34133/research.0225]
- Lu J, Ren J, Liu J, Lu M, Cui Y, et al. High-resolution single-cell transcriptomic survey of cardiomyocytes from patients with hypertrophic cardiomyopathy. Cell Prolif 2024;57(3):e13557. [PMID: 37766635 DOI: 10.1111/cpr.13557]
- Gogineni VS, Shah KB. High-risk percutaneous coronary intervention: challenges and considerations. Cardiovasc Innov Appl 2024;9(1). [DOI: 10.15212/CVIA.2024.0029]
- Eltzschig HK, Eckle T. Ischemia and reperfusion–from mechanism to translation. Nat Med 2011;17(11):1391-401. [PMID: 22064429 DOI: 10.1038/nm.2507]
- Peng Y, Zhou G, Guo M, Cheng Z, Luo S, et al. Inhibition of stimulator of interferon genes protects against myocardial ischemia-reperfusion injury in diabetic mice. Cardiovasc Innov Appl 2023;8(1). [DOI: 10.15212/CVIA.2023.0020]
- Ouyang Y, Hong Y, Mai C, Yang H, Wu Z, et al. Transcriptome analysis reveals therapeutic potential of NAMPT in protecting against abdominal aortic aneurysm in human and mouse. Bioact Mater 2024;34:17-36. [PMID: 38173843 DOI: 10.1016/j.bioactmat.2023.11.020]
- Ross JJ. Myocardial perfusion-contraction matching. Implications for coronary heart disease and hibernation. Circulation 1991;83(3):1076-83. [PMID: 1999010 DOI: 10.1161/01.cir.83.3.1076]
- Piccoli MT, Gupta SK, Viereck J, Foinquinos A, Samolovac S, et al. Inhibition of the cardiac fibroblast-enriched lncRNA Meg3 prevents cardiac fibrosis and diastolic dysfunction. Circ Res 2017;121(5):575-83. [PMID: 28630135 DOI: 10.1161/CIRCRESAHA.117.310624]
- Han P, Li W, Lin CH, Yang J, Shang C, et al. A long noncoding RNA protects the heart from pathological hypertrophy. Nature 2014;514(7520):102-6. [PMID: 25119045 DOI: 10.1038/nature13596]
- Wang L, Xu GE, Spanos M, Li G, Lei Z, et al. Circular RNAs in cardiovascular diseases: regulation and therapeutic applications. Research (Wash D C) 2023;6:0038. [PMID: 37040523 DOI: 10.34133/research.0038]
- Olshansky B, Ricci F, Fedorowski A. Importance of resting heart rate. Trends Cardiovasc Med 2023;33(8):502-15. [PMID: 35623552 DOI: 10.1016/j.tcm.2022.05.006]
- Kalcheva N, Qu J, Sandeep N, Garcia L, Zhang J, et al. Gap junction remodeling and cardiac arrhythmogenesis in a murine model of oculodentodigital dysplasia. Proc Natl Acad Sci U S A 2007;104(51):20512-6. [PMID: 18077386 DOI: 10.1073/pnas.0705472105]
- Cohen-Barak O, Yi Z, Hagiwara N, Monzen K, Komuro I, et al. Sox6 regulation of cardiac myocyte development. Nucleic Acids Res 2003;31(20):5941-8. [DOI: 10.1093/nar/gkg807]
- Li L, Zhang J, Zhang Q, Huang Y, Hu J. Cardiac proteomics reveals the potential mechanism of microtubule associated protein 4 phosphorylation-induced mitochondrial dysfunction. Burns Trauma 2019;7:8. [PMID: 30906793 DOI: 10.1186/s41038-019-0146-3]
- Hoerter J, Gonzalez-Barroso MD, Couplan E, Mateo P, Gelly C, et al. Mitochondrial uncoupling protein 1 expressed in the heart of transgenic mice protects against ischemic-reperfusion damage. Circulation 2004;110(5):528-33. [PMID: 15262832 DOI: 10.1161/01.CIR.0000137824.30476.0E]
- Rocca C, De Bartolo A, Granieri MC, Rago V, Amelio D, et al. The antioxidant selenoprotein T mimetic, PSELT, induces preconditioning-like myocardial protection by relieving endoplasmic-reticulum stress. Antioxidants (Basel) 2022;11(3):571. [PMID: 35326221 DOI: 10.3390/antiox11030571]
- Yu S, Sun Z, Ju T, Liu Y, Mei Z, et al. The m7G methyltransferase Mettl1 drives cardiac hypertrophy by regulating SRSF9-mediated splicing of NFATc4. Adv Sci (Weinh) 2024;11(29):e2308769. [PMID: 38810124 DOI: 10.1002/advs.202308769]
- Longchamp A, Mirabella T, Arduini A, MacArthur MR, Das A, et al. Amino acid restriction triggers angiogenesis via GCN2/ATF4 regulation of VEGF and H2S production. Cell 2018;173(1):117-29.e14. [PMID: 29570992 DOI: 10.1016/j.cell.2018.03.001]
- Wang W, Hu YF, Pang M, Chang N, Yu C, et al. BMP and notch signaling pathways differentially regulate cardiomyocyte proliferation during ventricle regeneration. Int J Biol Sci 2021;17(9):2157-66. [PMID: 34239346 DOI: 10.7150/ijbs.59648]
- Cao Y, Wang Y, Zhou Z, Pan C, Jiang L, et al. Liver-heart cross-talk mediated by coagulation factor XI protects against heart failure. Science 2022;377(6613):1399-406. [PMID: 36137043 DOI: 10.1126/science.abn0910]
- Dai Y, Zhao F, Chen Q, Zeng B, Gu W, et al. Microfluidic chip-integrated vascularized endometrial complexes: mitochondrial function and paracrine crosstalk enhance regenerative potential. Bioact Mater 2025;54:551-69. [DOI: 10.1016/j.bioactmat.2025.08.035]
- Lincoln J, Alfieri CM, Yutzey KE. BMP and FGF regulatory pathways control cell lineage diversification of heart valve precursor cells. Dev Biol 2006;292(2):292-302. [PMID: 16680829 DOI: 10.1016/j.ydbio.2005.12.042]
- Thurgur H, Penny J, Pinteaux E. Endothelial cell activation by interleukin-1 and extracellular matrix laminin-10 occurs via the YAP signalling pathway. J Neuroimmunol 2022;373:577993. [PMID: 36327619 DOI: 10.1016/j.jneuroim.2022.577993]
- Kulikauskas MR, Shaka X, Bautch VL. The versatility and paradox of BMP signaling in endothelial cell behaviors and blood vessel function. Cell Mol Life Sci 2022;79(2):77. [PMID: 35044529 DOI: 10.1007/s00018-021-04033-z]
- Yu P, Wilhelm K, Dubrac A, Tung JK, Alves TC, et al. FGF-dependent metabolic control of vascular development. Nature 2017;545:(7653):224-8. [DOI: 10.1038/nature22322]
- Jiang F, Tang J, Wei X, Pan H, Fan X, et al. BMP6, a potential biomarker of inflammatory fibrosis and promising protective factor for dilated cardiomyopathy. Chin Med 2025;20(1):12. [PMID: 39825396 DOI: 10.1186/s13020-025-01062-9]
- Li G, Shao Y, Guo HC, Zhi Y, Qiao B, et al. MicroRNA-27b-3p down-regulates FGF1 and aggravates pathological cardiac remodelling. Cardiovasc Res 2022;118(9):2139-51. [PMID: 34358309 DOI: 10.1093/cvr/cvab248]
- Kelly AG, Panigrahy D. Targeting angiogenesis via resolution of inflammation. Cold Spring Harb Perspect Med 2023;13(3):a041172. [PMID: 35817542 DOI: 10.1101/cshperspect.a041172]