Combating Multi-drug Resistant Mycobacterium Species by Efflux Pump Inhibitors: Contemporary Trends
1Department of Pharmacy, Birla Institute of Technology and Science Pilani, Pilani Campus, Rajasthan 333031, India
2Department of Natural Products, National Institute of Pharmaceutical Education and Research-Hyderabad (NIPER-H), Telangana, India
3Department of Pharmacology and Toxicology, National Institute of Pharmaceutical Education and Research-Hyderabad (NIPER-H), Telangana, India
4Department of Pharmacology, Nims Institute of Pharmacy, Nims University Rajasthan, Jaipur, Rajasthan, 303121, India
5Department of Pharmaceutics, Nims Institute of Pharmacy, Nims University Rajasthan, Jaipur, Rajasthan, 303121, India
6Department of Pharmacy Practice, Nims Institute of Pharmacy, University Rajasthan, Jaipur, Rajasthan, 303121, India
*Correspondence to: Dr. Gautam Kumar, Assistant Professor, Department of Pharmacy, Birla Institute of Technology and Science Pilani, Pilani Campus, Rajasthan 333031, India, E-mail: gautam.kumar@pilani.bits-pilani.ac.in; kumargautamgalaxy@gmail.com; Dr. Sandeep Kumar, Associate Professor, Nims Institute of Pharmacy, Nims University Rajasthan, Jaipur, Rajasthan, 203031, India, E-mail: ssandeepkatarea@gmail.com; sandeep.kumar2@nimsuniversity.org
Received: February 6 2025; Revised: April 1 2025; Accepted: May 3 2025; Published Online: June 2 2025.
Cite this paper:
Kumar G, Mondal P, Kadam A et al. Combating Multi-drug Resistant Mycobacterium Species by Efflux Pump Inhibitors: Contemporary Trends. BIO Integration 2025; 6: 1–32.
DOI: 10.15212/bioi-2025-0034. Available at: https://bio-integration.org/
Download citation
© 2025 The Authors. This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See https://bio-integration.org/copyright-and-permissions/
Abstract
Tuberculosis (TB) is caused by Mycobacterium tuberculosis (M.tb) complex (MTBC), which includes M.tb as one of the causative bacteria. In contrast, non-tuberculous mycobacteria (NTM) refers to Mycobacterium spp. that do not cause TB or leprosy (MTBC spp., M. leprae, and M. lepromatosis). Mycobacterium spp. are responsible for the deadliest infections and remain a significant challenge in diagnosis and treatment. Mycobacterium spp. have developed multiple complementary mechanisms to defend against antibiotics. Specifically, the mechanisms include modifying the drug target sites, enzymatically inactivating the drugs, and lowering intracellular antibiotic concentrations by overexpressing efflux pumps. These adaptations contribute to the emergence of multi-drug resistant pathogens. This review provides an overview of antibiotic resistance in Mycobacterium spp. with a focus on several key factors, such as enzyme-mediated antibiotic deactivation, gene expression, biofilm formation, and the role of efflux pumps. A critical objective of this review includes Mycobacterium efflux pumps, the significant role in antibiotic resistance, and compounds that act against these efflux pumps.
Keywords
ABC superfamily, efflux pump, MFS superfamily, Mycobacterium tuberculosis, non-tuberculous mycobacteria, RND superfamily, SMR superfamily, tuberculosis.
Introduction
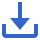
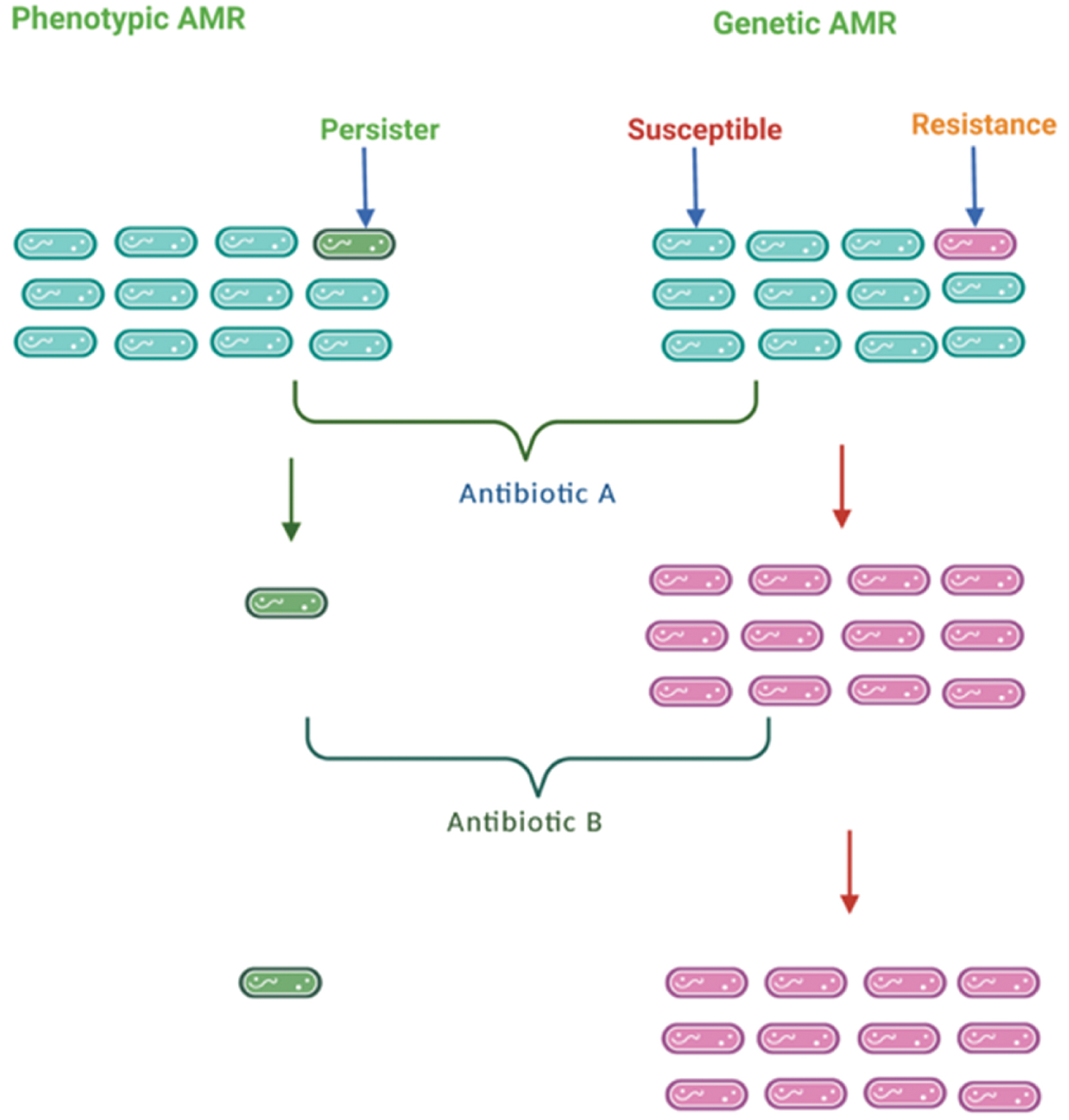
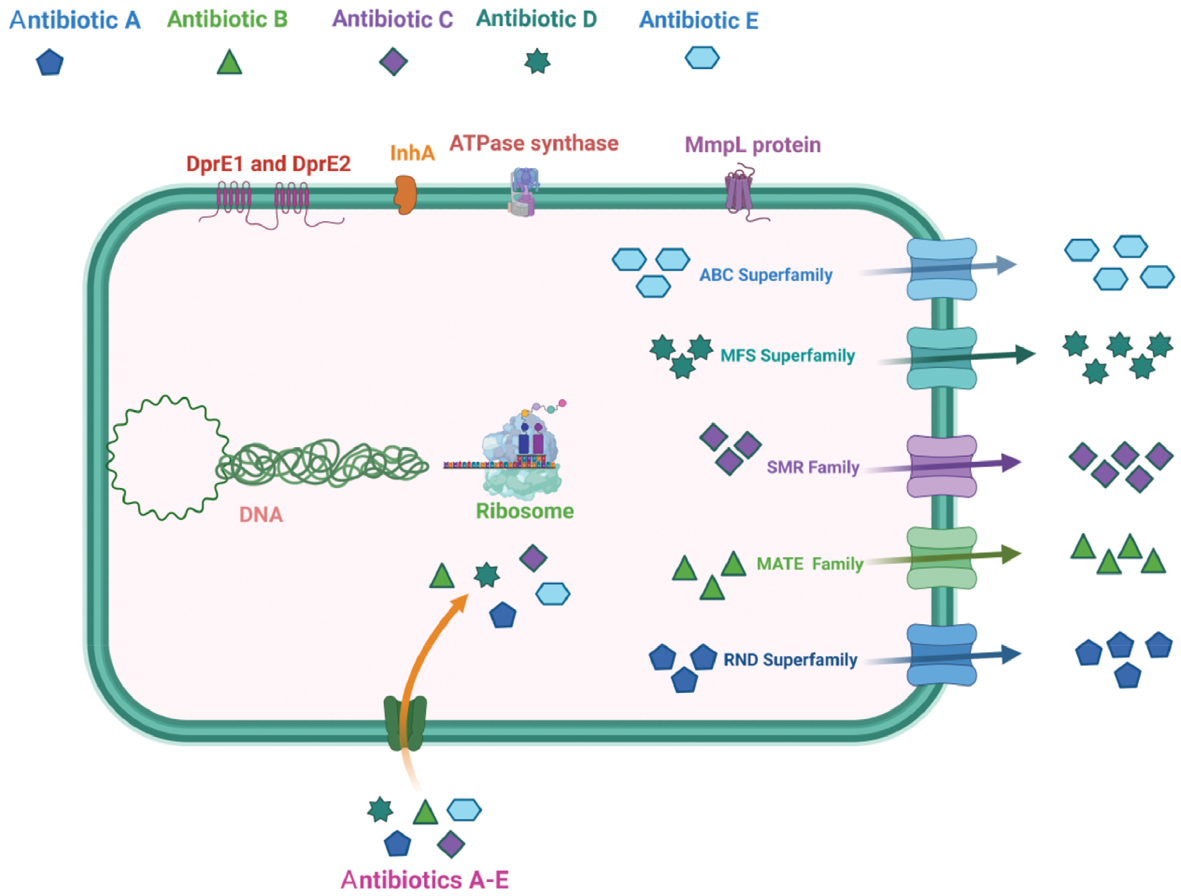
Tuberculosis (TB) remains the deadliest infection from single infectious organisms and continues to hold centre stage among bacterial infections and remains a significant challenge in terms of diagnosis and treatment [1, 2]. The WHO Report 2023 stated that 7.5 million individuals were diagnosed with TB in 2022 and approximately 1.6 million people died from TB, including 187,000 co-infected with HIV and TB [3]. Macrophages phagocytose most internalized Mycobacterium tuberculosis (M.tb) bacilli that reach the physiologic system and eliminate significant bacilli populations. Some bacteria can escape the phagosome compartment and remain inactive, allowing patients to remain asymptomatic [4, 5]. The M.tb bacilli subpopulation is known as persisters, which are phenotypically tolerant to antibiotics. Also, persisters remains metabolically inactive and do not grow like resistant mutant M.tb bacilli [6]. A schematic representation depicting genetic and phenotypic resistance is shown in Figure 1.
Figure 1 Schematic representation depicting antimicrobial resistance (phenotypic and genetic) is displayed above. Mycobacterium persisters do not have the effects of antibiotics A and B. The genetically mutated Mycobacterium survives and multiplies in the proximity of antibiotics A and B.
Non-tuberculous mycobacteria (NTM) are species that belong to the same genus (Mycobacterium) but refers to bacteria other than the M.tb complex (MTBC) that do not cause leprosy or TB. NTM are ubiquitous and can survive in a wide range of environmental conditions. Most likely, drinking water or other materials are sources of NTM. Moreover, NTM can survive chlorine treatment and enter tap water [7–9]. NTM species are classified according to the Runyon classification based on growth rates in solid culture medium and pigment formation. Notably, Runyon types I, II, and III NTM are slow-growing Mycobacterium spp. and are characterised based on pigment formation or colour development. Runyon type IV Mycobacterium spp. are rapidly growing bacteria that can grow in <7 d. Pigments are categorized as Runyon type I and are considered photochromogens when developed after exposure to light. Mycobacteria classified as Runyon type II develop pigments, even in the dark; these bacteria are known as scotochromogens. Mycobacteria are categorized as Runyon type III if the pigments are not intense and these mycobacteria are known as non-photochromogens [10]. Runyon classification of NTM and their species are given in Table 1.
Table 1 Non-Tuberculous Mycobacteria (NTM) Runyon Classification and the Mycobacterium spp
| Runyon Classification | NTM Species | Ref. |
|---|---|---|
| Runyon type I (photochromogens) | M. kansasii | [13, 14] |
| M. simiae | [15] | |
| M. marinum | [16, 17] | |
| Runyon type II (scotochromogens) | M. gordonae | [18, 19] |
| M. szulgai | [20] | |
| M. scrofulaceum | [21] | |
| Runyon type III (non-photochromogens) |
M. avium complex (M. avium and M. intracellulare) |
[22] |
| M. malmoense | [23] | |
| M. xenopi | [23] | |
| M. terrae complex | [214] | |
| M. genavense | [23] | |
| M. haemophilum | [23] | |
| M. ulcerans | [23] | |
| Rapid-growing Runyon type IV | M. abscessus | [23, 24] |
| M. chelonae | [23, 215, 216] | |
| M. smegmatis | [217] | |
| M. fortuitum | [25] |
NTM can infect humans and are responsible for severe infections in immune-compromised and -competent individuals [11, 12]. NTM species, including M. kansasii, M. simiae, M. gordonae, M. szulgai, M. scrofulaceum, M. avium complex (MAC), and M. malmoense, are responsible for the majority of pulmonary infections [13–23]. NTM species, such as M. marinum, M. ulcerans, M. chelonae, M. fortuitum, and M. abscessus, are responsible for skin infections. In addition, M. chimaera and M. abscessus are the most common NTM species responsible for soft tissue infections [24–26].
Because M.tb and NTM species react favourably to the standard smear acid-fast staining process, clinical detection of M.tb and NTM species poses a considerable problem and frequently results in incorrect interpretations. As a result, the frequency of NTM has been underestimated due to the prevalence of TB in several countries [27].
Treatment of TB includes a cocktail of first-line anti-TB drugs, including isoniazid (INH), rifampicin (RIF), ethambutol (EMB), and pyrazinamide (PZA) [28]. Second-line anti-TB drugs (groups 2–4) are prescribed to treat drug-resistant TB. Group 2 drugs include injectable aminoglycosides (streptomycin [STR], kanamycin, and amikacin) and polypeptides (capreomycin and viomycin). Group 3 drugs include oral and injectable fluoroquinolones [FQs] (gatifloxacin, moxifloxacin, and levofloxacin). Group 4 drugs include the following oral medications: p-aminosalicylic acid; cycloserine; teridazone; ethionamide; prothionamide; thioacetazone; and linezolid [LZD] [28]. Third-line anti-TB drugs (group 5) have unclear efficacy or an undefined role. Group 5 drugs include clofazimine, linezolid, amoxicillin plus clavulanate, imipenem plus cilastatin, and clarithromycin [29]. In recent years, several compounds have been identified as potential anti-TB agents and are summarised in Table 2 [30–32].
Table 2 Anti-TB Agents or Drugs under Clinical Evaluation
| Anti-TB Agents/Drugs | Chemical Class | Mode of Action | Ref. |
|---|---|---|---|
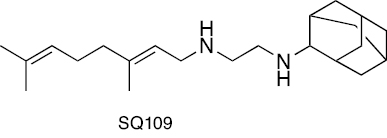 |
Ethylenediamine | MmpL3 transporter inhibition. Additionally, SQ109 disrupts the proton motive force of the M.tb membrane. | [218–221] |
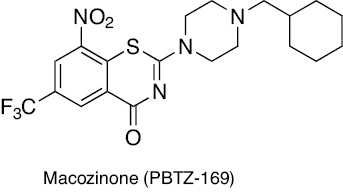 |
Benzothiazones | Inhibition of membrane DprE1 enzyme | [222] |
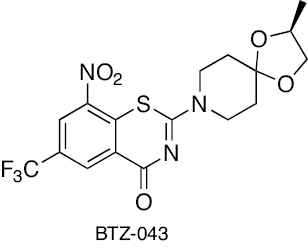 |
Benzothiazones | Inhibition of membrane DprE1 enzyme | [223, 224] |
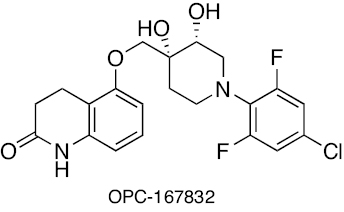 |
Carbostyril | Inhibition of membrane DprE1 enzyme | [225, 226], |
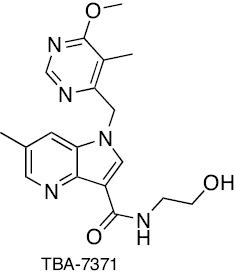 |
Azaindole | Inhibition of membrane DprE1 enzyme | [227–230] |
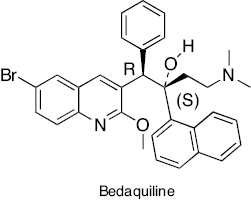 |
Diarylquinoline | ATP synthesis inhibition | [231, 232] |
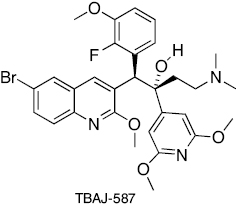 |
Diarylquinoline | ATP synthesis inhibition | [233, 234] |
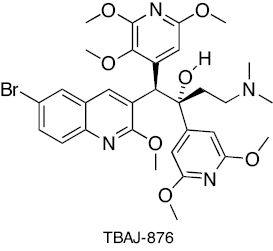 |
Diarylquinoline | ATP synthesis inhibition | [233, 234] |
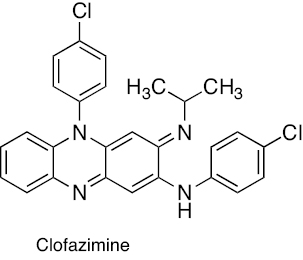 |
Riminophenazine | Clofazimine is a competitive inhibitor of menaquinone (MK-4) and checks the point of entry of electrons into the respiratory chain, which leads to energy depletion. | [235–240] |
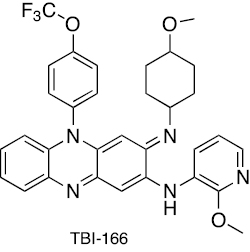 |
Riminophenazine | TBI-166 presumably competes with menaquinone (MK-4), an essential cofactor in the mycobacterial ETC, and checks the electron’s entry point into the respiratory chain | [241, 242] |
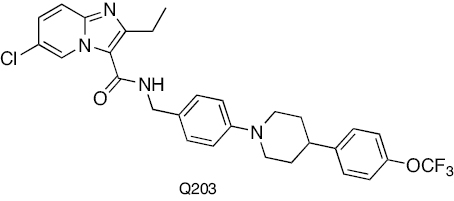 |
Imidazopyridine amide | Cytochrome bc1 complex inhibition | [243–246] |
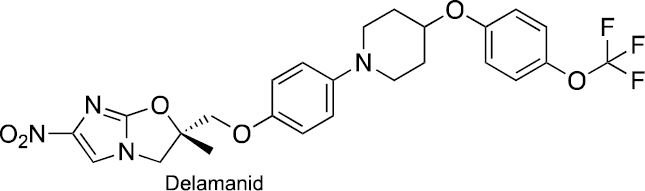 |
Nitro-dihydro-imidazooxazole | Delamanid exhibits anti-mycobacterial activity by inhibiting methoxy-mycolic acid and ketomycolic acid. | [102, 247, 248] |
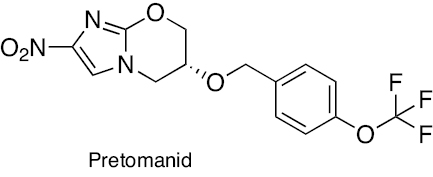 |
Nitroimidazooxazine | Pretomanid inhibits cell wall lipids and proteins. | [248–250] |
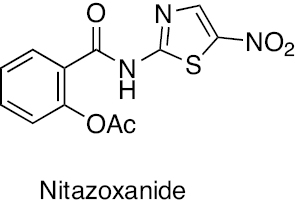 |
Nitrothiazolyl salicylamide | Nitzoxanide displays activity against M.tb by disrupting membrane potential and pH homeostasis. Moreover, Nitzoxanide also targets human quinone oxidoreductase (NQO1)-mediated mTORC1 signalling that promotes autophagy of M.tb inside host cells. | [251–254] |
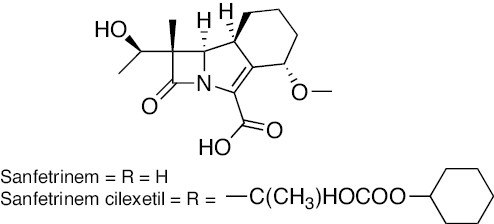 |
Tricyclic β- lactam | Cell wall inhibitors | [255–258] |
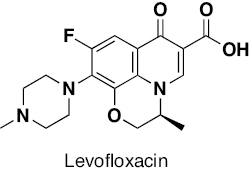 |
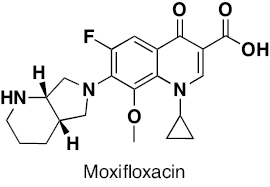
Fluoroquinolone | Inhibition of DNA replication by binding to topoisomerase II (DNA gyrase). | [259, 260] |
 |
[259, 260] | ||
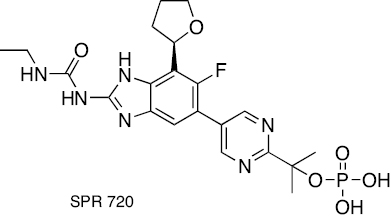 |
Aminobenzimidazole | Inhibition of gyrase B | [44, 96] |
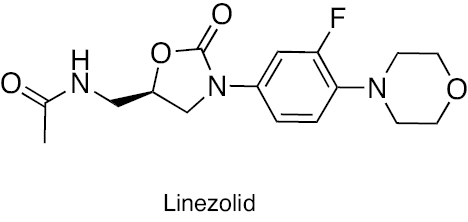 |
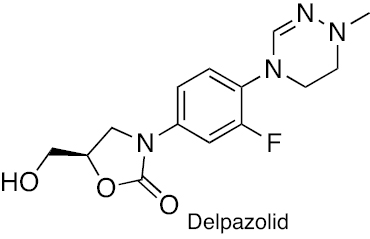
Oxazolidinone | Inhibition of protein synthesis by binding to the 50S ribosome subunit | [261] |
 |
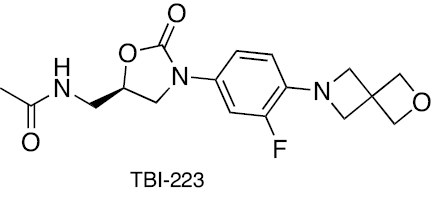
[262] | ||
 |
[263–265] | ||
 |
[266] | ||
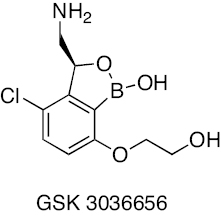 |
Benzoxaborole | Leucyl-tRNA synthetase (LeuRS) | [36] |
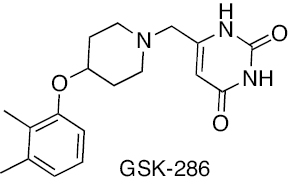 |
Pyrimidine-2,4-dione | Cholesterol catabolism | [36] |
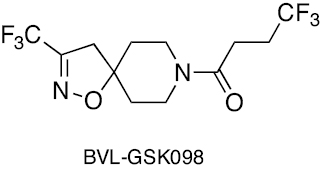 |
Spiroisoxazoline | Antagonizes the resistance mechanism of ethionamide | [267] |
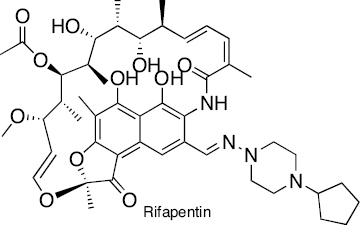 |
Macrolactum | Inhibition of DNA-dependent RNA polymerase activity | [268–271] |
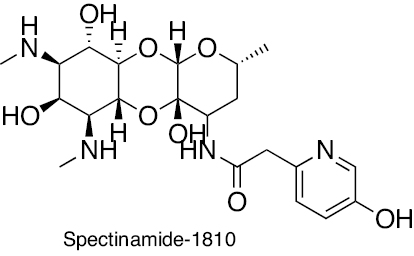 |
Aminoglycoside | Ribosome protein synthesis inhibition | [272–275] |
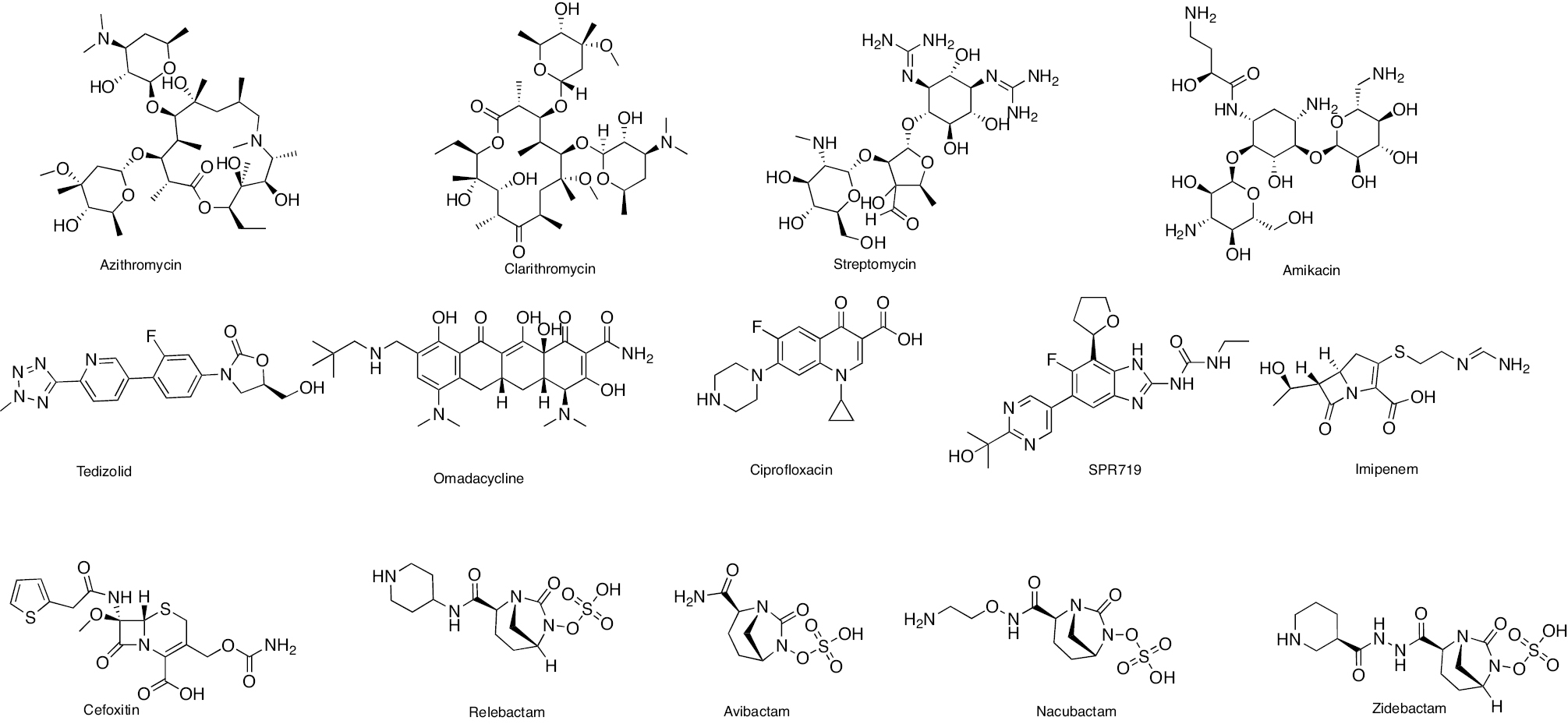
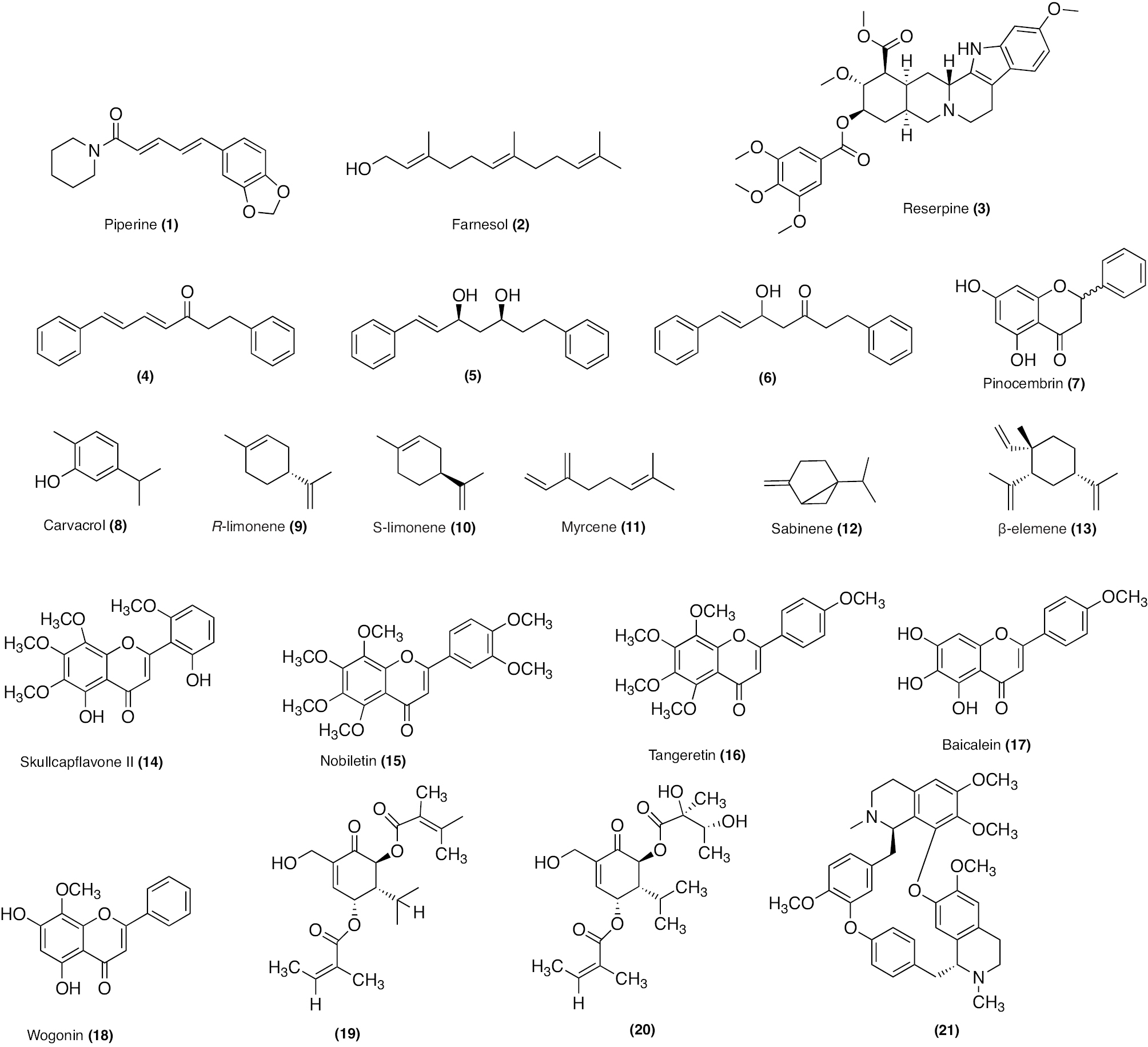
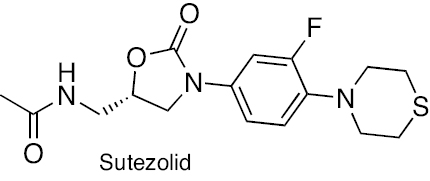
Treatment regimens for fast-growing NTM include macrolides, aminoglycosides, fluoroquinolones, oxazolidinones, tigecycline, carbapenems, and cephalosporins. In contrast, RIF, macrolides, EMB, and amikacin are front-line agents for slow-growing bacilli NTM species [12, 33]. The antibacterial activity of macrolides, such as azithromycin and clarithromycin, inhibits the 50S ribosomal subunit protein synthesis. It possesses activity against MAC, M. kansasii, M. malmoense, and M. abscessus [34, 35]. The antibacterial activity of aminoglycosides, including streptomycin and amikacin, inhibits protein synthesis by acting on the 30S ribosomal subunit. Aminoglycosides efficiently cure patients with cavitary MAC lung disease [36, 37]. The antibacterial activity of oxazolidinone class antibiotics inhibits binding of 23S rRNA to the catalytic site of the 50S ribosome, which in turn prevents the formation of a functional 70S initiation complex. Delpazolid has antibacterial activity against in vitro and in vivo models of macrophage and murine M. abscessus infections [38]. Tedizolid exhibits antibacterial activity against M. fortuitum, M. chelonae, and M. abscessus subsp. massiliense, M. marinum, M. kansasii, MAC, and M. simiae [38, 39]. Tetracycline, doxycycline, and tigecycline effectively treat NTM infections. The tetracycline analog, omadacycline, has inhibitory activity against M. abscessus subsp. abscessus, M. abscessus subsp. massiliense, M. chelonae, M. immunogenum, M. fortuitum, and M. mucogenicum, including resistant strains [40]. The fluoroquinolones, such as ciprofloxacin and moxifloxacin, are active against MAC and M. intracelluare [41, 42]. SPR719 is a gyrase B inhibitor that effectively reduces the mycobacterial burden in lungs of infected mice [43]. SPR719 is active against multiple clinical strains of NTM, including MAC and M. abscessus [44]. SPR720 is a phosphate prodrug of SPR719 that effectively treats human NTM and pulmonary TB infections [45]. The antibacterial activity inhibits DNA-directed RNA polymerase (RNAP). Rifabutin is an RIF derivative active against M. abscessus subsp. abscessus. Rifabutin also inhibits other NTM, including MAC, M. kansasii, M. malmoense, and M. xenopi [46, 47]. The anti-mycobacterial activity of bedaquiline (BDQ) inhibits ATP synthesis and growth of MAC, M. abscessus subsp. abscessus, M. abscessus subsp. massiliense, and M. kansasii [48]. Clofazimine has been reported to be potent in patients with pulmonary NTM, M. abscessus group, and MAC infections [49]. Cell wall biosynthesis inhibitors disrupt the bacterial cells, leading to a compromise in bacteria viability [50]. β-lactams, including imipenem and cefoxitin, exhibit antibacterial activity against M. abscessus. β-lactamase inhibitors, including clavulanate, tazobactam, avibactam, relebactam, nacubactam, and zidebactam, in combination with other β-lactams have potential inhibitory activity against NTM species [51–55]. The chemical structures of compounds acting against NTM species are shown in Figure 2. In this review the diverse mechanisms underlying antibiotic resistance in Mycobacterium spp. are discussed and effective solutions are explored, including natural and synthetic compounds, to tackle the pressing challenge of antibiotic resistance in these organisms.
Figure 2 Chemical structures of compounds acting against NTM species.
Antibiotic resistance in Mycobacterium spp.
Antibiotic resistance in pathogenic microorganisms poses a global threat. Antibiotic resistance is a condition in which conventional antibiotics cannot effectively cure a disease [56]. In most cases, when patients do not recover from TB, second-line anti-TB drugs are given in combination for 18–24 months to treat multidrug-resistant (MDR)-TB [28]. M.tb has demonstrated resistance to most first- and second-line anti-TB drugs. Current research has documented the development of drug-resistant TB (DR-TB), MDR-TB, extensively drug-resistant TB (XDR-TB), and totally drug-resistant TB (TDR-TB) over the past several decades [57]. M.tb strains exhibit various levels of drug resistance. The I-R strain is resistant to INH, while the R-R strain has resistance to RIF. The MDR-TB strain is resistant to INH and RIF. Pre-extensively drug-resistant (Pre-XDR) TB strains are resistant to RIF and fluoroquinolones. XDR TB strains are resistant to RIF, fluoroquinolones, and BDQ or linezolid. Finally, TDR TB strains are resistant to all first- and second-line anti-TB drugs [58]. MDR-TB poses a significant challenge that has made conventional TB treatment ineffective, which warrants immediate attention and innovative solutions. A list of drug-resistant M.tb strains is given in Table 3.
Table 3 Major Drug-Resistant M.tb Strains
| Type of Drug-resistant M.tb Strains | Resistant to Drug |
|---|---|
| I-R | M.tb shows resistance to isoniazid (INH) |
| RR | M.tb shows resistance to rifampicin (RIF) |
| MDR | M.tb shows resistance to INH + RIF |
| Pre-XDR | M.tb shows resistance to RIF + fluoroquinolone (a class of second-line anti-TB drugs) |
| XDR | M.tb shows resistance to RIF + any fluoroquinolone, + at least one of bedaquiline or linezolid |
| TDR | M.tb shows in vitro resistance to all first- and second-line drugs. |
MDR in M.tb strains primarily evolved due to the initial ineffectiveness of TB treatments, including first- and second-line antibiotics and exposure of bacilli to several antibiotics [56]. Antibiotic resistance primarily arises from factors, such as inadequate antibiotic concentration, extended periods of monotherapy, poor prescribing practices, and variations in the pharmacogenomics, pharmacodynamics, and pharmacokinetics of the administered drugs [59, 60].
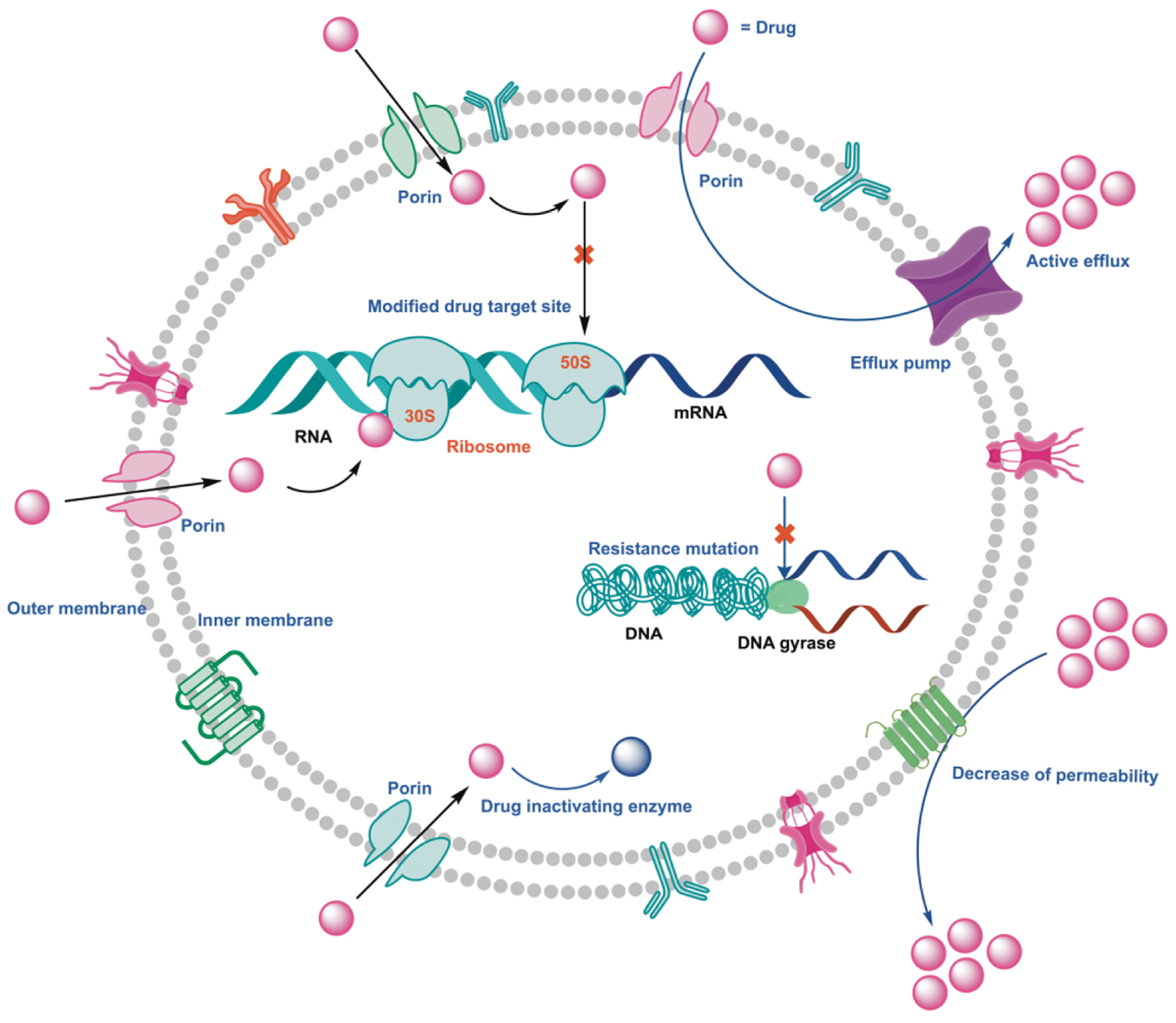
Antibiotic resistance in mycobacteria can be divided into two categories: acquired resistance; and intrinsic resistance. Intrinsic resistance refers to the natural ability of bacteria to resist antibiotics, whereas acquired resistance develops through specific chromosomal mutations or the horizontal transfer of genes [61]. There are no reports of horizontal gene transfer (HGT) of resistance genes through plasmids or transposable elements in Mycobacterium spp. This indicates that the acquired resistance results from chromosomal mutations [61, 62]. Single-nucleotide polymorphisms (SNPs), mutations involving changes in nucleotide sequences, insertions of new genetic material, and deletions cause changes in the genomic DNA of M.tb [12, 63, 64]. Mycobacterium bacilli exhibit several modes of antibiotic resistance, including low permeability of the cell wall, overactive efflux pumps, enzymatic degradation of antibiotics, and modifications at the target site (Figure 3) [57, 65]. Recent data have revealed that M.tb and NTM exhibit acquired resistance against most antibiotics and develop complementary mechanisms to counteract their effects [66]. Therefore, there is an urgent need to control MTBC and NTM infections and address Mycobacterium resistance issues [28].
Figure 3 A graphical representation depicting bacterial modes of resistance, including antibiotic degrading enzyme, antibiotic altering enzyme, antibiotic decrease to permeability, and efflux pump.
Genes-mediated antibiotic resistance in Mycobacterium spp.
MTBC acquires antibiotic resistance due to spontaneous gene alterations. For example, the antibiotic-dependent spontaneous mutation rate that leads to resistance varies, as follows: RIF, approximately 1 in 108 bacilli; and INH, STR, and EMB, approximately 1 in 106 bacilli [67]. Sequential changes in additional genes make the strains unresponsive to various targets [63, 68]. MspA, the M. smegmatis membrane porin, transfers hydrophilic substances via the outer membrane to the cytoplasm. Additionally, MspA is responsible for introducing small hydrophilic antibiotics into M. smegmatis. Studies have shown that deleting MspA porin results in resistance to various hydrophilic antibiotics. The porin mutant of M. smegmatis, ML10 (a deletion of MspA and MspC) exhibits β-lactam antibiotic resistance [69]. Similarly, CpnT is an outer membrane protein in M. bovis BCG and M.tb that is involved in food absorption. Strains with a CpnT mutation are unresponsive to antibiotics, such as RIF, EMB, clarithromycin, tetracycline, and ampicillin, and to nitric oxide [70].
INH is a prodrug used in treating TB that requires metabolic activation by catalase-peroxidase. INH resistance has been observed in M.tb with katG and inhA gene mutations. Mycobacterium strains with katG gene mutations have decreased INH activation because there is a pronounced decrease or loss in catalase-peroxidase activity. inhA gene mutations may influence construction of the bacillus cell wall. The minimum inhibitory concentration (MIC) of INH is significantly higher than M.tb isolates with katG mutations. Moreover, structural alterations in the INH target subsequently results in INH resistance [71].
Additional mutational genes in other clinical INH-resistant isolates have been identified, which include furA, ndh (encodes NADH dehydrogenase), ahpC, Rv0340-0343, fadE24, efpA, and kasA [72, 73]. M.tb exposed to INH shows variance in ManLam-related gene expression. The hspX gene in increased in RIF-resistant and MDR-TB strains in response to INH, whereas tgs1 and sigE are strongly upregulated in MDR-TB and drug-sensitive TB strains, respectively [74]. RIF exhibits anti-mycobacterial activity by acting on the β-subunit of DNA-dependent RNA polymerase. The Escherichia coli model was used to study RIF resistance, which demonstrated that the rpoB gene mutations are the primary cause of RIF resistance [75]. It has been shown that rpoB gene mutations are responsible for cross-resistance to several rifamycins. In addition, rpoB gene mutations are present in the 81 bp RIF-resistance determining region (RDDR), which affects the RNA polymerase β-subunit. These modifications prevent rifamycins from binding to the RNA polymerase β-subunit, leading M.tb to be less susceptible to the RIF, and ultimately RIF-resistant strains emerged [76]. The most common mutated codons in the RDDR are 531, 526, and 516 [77]. Moreover, homoplastic SNP analysis revealed that the compensatory effects of rpoC and rpoA significantly contribute to RIF resistance [78].
PZA is a prodrug that requires pyrazinamidase for conversion to the metabolically active form. A phylogenetic analysis of the MTBC tree revealed that PZA resistance could be attributed to the pncA H57D mutation [79]. Along with pncA, other genes, such as rpsA and panD, contribute significantly to PZA resistance in M.tb. Mutations, such as NK11K, P69T, and D126N, are present in the pncA gene. According to a recent study, pncA gene mutations reduce the active site volume, making it more difficult for PZA to bind to the active site. Additionally, these modifications lead to a low binding affinity for PZA [80, 81]. Interestingly, M. bovis, which resists PZA therapy, carried the pncA H57D mutation [79]. Notably, mutations in the pncA gene do not always result in PZA resistance and PZAse critical failure. Tt has been verified that pncA gene mutations lead to a limited degree of PZA resistance by partially decreasing PZAse activity. In addition to the above-mentioned mutations other mutations e.g. D12A, T135P, and D49N are implicated in PZA resistance [82].
EMB exhibits anti-mycobacterial activity by acting on the target Emb proteins responsible for cell wall biosynthesis. The Emb proteins include EmbA, EmbB, and EmbC, and EMB resistance mutations are mainly located near the Emb binding site [83]. The mutations are primarily present in the embCAB operon that encodes arabinosyl transferases, which are responsible for EMB resistance. Among the arabinosyl transferases, alterations are most frequently noted in the embB gene and less often in the embC, embA, embR, and ubiA genes [84]. Moreover, ubiA gene mutations lead to enhanced production of decaprenylphosphoryl-D-arabinose, which is the building block used in cell wall synthesis and is the likely cause of EMB resistance [85]. Most mutations have been noted in the 576 bp embB region, which is known as the ETB resistance-determining region (ERDR). The canonical mutations of embB codons are 306, 406, and 497 [86, 87]. Interestingly, it has been shown that EMB resistance in M.tb isolates have slight variations according to geographic regions [86, 87]. STR hinders translational proofreading and stops protein synthesis in Mycobacterium spp. rrs, rpsL, and gidB gene mutations encode 16S rRNA, protein S12, and 7-methylguanosine methyltransferase, which are linked to STR resistance. gidB inactivation methylates 16S rRNA, resulting in the development of STR resistance [88]. The three rpsL gene mutations are present in Lys43Arg, Lys88Arg, and Lys88Thr residues, and two gidB mutations are mainly observed in Trp45Ter and Gly69Asp residues. The most frequent STR mutation among rpsL gene mutations is Lys43Arg. Notably, Mycobacterium spp. devoid of rpsL and rrs mutations do not exhibit whiB7-mediated STR resistance [89]. Insignificant resistance to kanamycin and STR has been reported to be due to eight different mutations in the 5′ untranslated region of the transcriptional activator whiB7 [90]. There is resistance to kanamycin and STR due to acquisition of unlinked chromosomal mutations. Notably, mutations increased whiB7 transcripts by a factor of 23–145, which enhances eis (Rv2416c) and tap (Rv1258c) expression. Increased expression of tap mediates STR resistance via overexpression of the efflux pump and eis is responsible for kanamycin resistance in mutated Mycobacterium spp. [90].
D-cycloserine exhibits anti-mycobacterial activity by acting on the enzymes (D-alanine racemase and D-alanine:D-alanine ligase), which are involved in peptidoglycan metabolism. Notably, overexpression of D-alanine racemase and D-alanine:D-alanine ligase are responsible for D-cycloserine resistance in M. smegmatis. A point mutation in the cycA gene causes M. bovis BCG resistance to D-cycloserine [91]. A mutation in the alanine racemase-encoding gene (rv3423c) is responsible for cycloserine resistance in M.tb [92].
FQs exhibit anti-mycobacterial action by acting on DNA gyrase and topoisomerase IV [43]. Mutations causing resistance to quinolone antibiotics occur in a specific segment of gyrA known as the quinolone resistance-determining region (QRDR). Codons 90 and 94 of gyrase A are referred to as the QRDR [93]. Moreover, mutations in gyrA and gyrB may cause the development of cross-resistance to many FQs [94]. M.tb isolates exposed to ofloxacin had a primary target mutation in gyrA and a minor mutation in gyrB [94]. In addition, the ESX-5 membrane complex secretion system protein, Rv1783, exhibited a V762G mutation, which is responsible for ofloxacin resistance [94]. This V762G mutation increased the MIC of ofloxacin from 1 to 32 mg/L in the M.tb strain. Notably, the MIC of ofloxacin in the M.tb strain did not show any change in the use of efflux pump inhibitors (EPIs), carbonyl cyanide m-chlorophenylhydrazone (CCCP), and reserpine. This finding implies that the gyrA target mutation and the V762G mutation in Rv1783 reduced the efficacy of ofloxacin against M.tb [94]. Other studies have suggested that M. smegmatis, M.tb, and M. avium possess the mfpA gene responsible for quinolone resistance. The mfpA gene encodes a pentapeptide repeated pentapeptide protein. This peptide binds to DNA gyrase, preventing FQs from reaching the target [94]. SPR719 belongs to an aminobenzimidazole class, which exhibits anti-TB activity by blocking gyrase B enzyme activity [95]. SPR720, a prodrug of SPR719, demonstrates anti-mycobacterial activity against MDR-TB, drug-susceptible M.tb, and MDR-TB strains. Moreover, SPR720 is active against NTM, such as MAC and M. abscessus [43, 44, 96]. A recent investigation revealed that mutation in the gyrase B ATPase domain is associated with low-frequency resistance in M. avium and M. abscessus. Furthermore, frameshift mutations in the transcriptional repressor, MAB_4384, a regulator of MmpS5/MmpL5 expression, are responsible for SPR719 resistance in M. abscessus [95].
M.tb uses proton motive force (PMF) generated across the plasma membrane to synthesize ATP. Inhibiting ATP synthesis results in an imbalance in cellular pH and PMF. BDQ selectively inhibits M.tb ATP synthase [97]. The atpE gene encodes ATP synthase, which is responsible for BDQ resistance in M.tb strains. In addition, it has been shown that the development of BDQ resistance in M.tb leads to clofazimine cross-resistance. Rv0678 is a negative regulator of mmpL5 and mmpS5. This mutation is also responsible for BDQ resistance. Significantly, there was a > 4-fold increase in BDQ MIC due to this mutation. Verapamil and reserpine are well-known EPIs that reduce the MIC of BDQ and clofazimine against M.tb with the Rv0678 mutation. Additionally, reserpine enhanced the bactericidal effects of BDQ, whereas verapamil had no such effect on mice infected with M.tb. Therefore, reserpine may inhibit the Rv0678 gene mutation in vivo [98].
Another study showed that loss of mutations in pepQ (Rv2535c), corresponding to the putative Xaa-Pro aminopeptidase, is responsible for clofazimine cross-resistance in M.tb. Further analysis of mice infected with M.tb H37Rv (pepQ mutants) showed a 4-fold higher BDQ and clofazimine MIC. The concomitant use of verapamil and reserpine reduces and restores the BDQ and clofazimine MIC in mutant and parent M.tb strains. This result implies that pepQ function loss decreases M.tb susceptibility to BDQ and clofazimine [99]. The anti-TB program recently included BDQ for treating MDR-TB. Involvement of the mutation in Rv0678 and Rv2535c reduces the effectiveness of BDQ. Therefore, adjuvant therapy may be required to treat TB.
Delamanid is a prodrug belonging to the nitro-dihydro-imidazooxazole class used in treating MDR-TB [100, 101]. M.tb bacilli activate delamanid and catalyze reduction of the nitro to the active form. Delamanid activation requires reduced deazaflavin co-factor, F420, F420-dependent glucose-6-phosphate dehydrogenase, and the nitroreductase gene product, Rv3547 [102]. Pretomanid is also a prodrug belonging to the nitroimidazoles class. Mycobacterial glucose-6-phosphate dehydrogenase converts pretomanid into the metabolically active form. Delamanid and pretomanid resistance have been linked to gene mutations (ddn, fgd1, fbiA, fbiB, fbiC, and fbiD) involved in the F420-dependent bioactivation pathway [103, 104]. A structural investigation showed that several point mutations in the gene encoding the delamanid-activating enzyme impact the ability to bind with delamanid [105]. The gene, Rv2983 (fbiD), is required to synthesize the co-factor, F420. Notably, fbiB and Rv2983 mutants have minor alterations in delamanid sensitivity but substantial levels of pretomanid resistance. Additionally, nitroimidazole-resistant, F420 H2-deficient M.tb mutants are hypersensitive to malachite green [106].
DprE1 is an enzyme crucial for the biosynthesis of decaprenylphosphoryl-D-arabinofuranose (DPA). D-arabinofuranose (Araf) residues and arabinosyltransferase biosynthesis use DPA as a substrate. D-arabinofuranose is the building block for lipoarabinomannan and arabinogalactan [107]. Inhibition of the DprE1 enzyme prevents DPA biosynthesis. Therefore, the DprE1 enzyme is an essential anti-TB target [108]. BTZ043 and PBTZ169 have anti-mycobacterial activity by blocking the DprE1 enzyme, which causes arabinogalactan depletion. These two drugs have undergone clinical evaluation and completed phase II clinical studies. Interestingly, mutations in Rv0678 exhibit negligible cross-resistance to medicines of the benzothiazinones class, including BTZ043 and PBTZ169. Further studies are required to confirm that Rv0678 mutations will impair the efficacy of BTZ043 and PBTZ169 in treating TB in humans [109].
LZD belongs to the oxazolidinone class and demonstrates antibacterial action by interfering with bacterial protein synthesis in M. abscessus [38]. Earlier studies have confirmed the involvement of efflux pumps, including drrABC, rv0987, lmrS, acrAB, mmpL9, acrF, and optrA, in extrusion of LZD. The molecular analysis of LZD-resistant M. abscessus harbors 23S rRNA mutations. Also, there are enhanced lmrS and mmpL9 transcriptional levels in resistance strains compared to LZD-susceptible strains [110]. The anti-TB drugs/agents and the mutation-induced resistance is summarised in Table 4.
Table 4 Anti-Tubercular Drugs/Agents and Mutation-Associated Resistant Mechanism
| Drug/agent | Mutation | Ref. |
|---|---|---|
| Isoniazid | Suppression of KatG leads to decreased activation of INH. | [71, 72] |
| Rifampicin | The rpoB mutation causes a conformational change in the β-subunit of RNA polymerase, leading to decreased binding affinity for rifampicin. | [75, 76] |
| Pyrazinamide | A mutation in the pncA gene prevents pyrazinamide from being converted into its active acid form. | [79–81] |
| Ethambutol | Mutations in the embCAB operon encode arabinosyl transferases, which are responsible for EMB resistance. | [84, 86, 87] |
| Amikacin/Kanamycin | Modulation of the 16S rRNA target site (1400 and 1401 rrs gene) and overexpression of aminoglycoside acetyltransferase results in increased drug inactivation. | [36, 37] |
| Capreomycin | A mutation in the tlyA gene reduces the activity of rRNA methyltransferase and is associated with cross-resistance to aminoglycosides. | [276] |
| Streptomycin | Mutations in the rrs, rpsL, and gidB genes encode 16S rRNA, protein S12, and 7-methylguanosine methyltransferase, which are linked to STR resistance. | [88] |
| Fluoroquinolones | Mutations in the gyrA and gyrB genes alter the binding of DNA gyrase A/B to fluoroquinolones. | [94] |
| Ethionamide | Mutations in the ethA and inhA genes result in reduced prodrug activation and overexpression of InhA, which also contributes to cross-resistance with INH. | [277] |
| Cycloserine | The overexpression of alrA leads to a reduction in the effectiveness of cycloserine. A point mutation in a cycA gene causes resistance to D-cycloserine. | [91] |
| P-aminosalicyclic acid | thyA mutations reduce the activated concentrations of P-aminosalicylic acid, while folC mutations change the binding site of P-aminosalicylic acid. | [276] |
| Clofazimine | A mutation in Rv0678 leads to upregulation of MmpL5. | [235, 236] |
| Clarithromycin | Expression of emr37 leads to modulation of the 23S rRNA site. | [111] |
| Bedaquiline | atpE mutations cause modulation in the binding site for bedaquiline. | [98] |
| Q203 | Mutation in the b subunit of the cytochrome bc1 complex. | [278] |
| Delamanid and pretomanid | Gene mutations (ddn, fgd1, fbiA, fbiB, fbiC, and fbiD) are involved in the F420-dependent bioactivation pathway. | [103, 104] |
Enzymes mediate antibiotic resistance in Mycobacterium spp.
Mycobacterial enzymes transform drugs, rendering drugs inactive and unable to bind to targets. Mycobacterium spp. synthesize several enzymes that can cleave and degrade several classes of antibiotics, including β-lactams, aminoglycosides, and macrolides [111]. Mycobacterium produces ambler class-A β-lactamase, which is encoded by the blaC gene and cleaves β-lactam antibiotics into an inactive form. Mycobacterium spp., such as M. smegmatis and M. fortuitum, have been reported to possess ambler class-A β-lactamase [112]. Aminoglycosides, such as neomycin, kanamycin, gentamycin, tobramycin, and ribostamycin, have amino groups. The mycobacterial N-acetyltransferase enzyme carries out acetylation to these amino groups and renders the amino acid-containing antibiotics inactive. The M. abscessus MAB_4395 gene encodes 2′ N– acetyltransferase, which is responsible for acetylating gentamicin C, dibekacin, tobramycin, and kanamycin B, and rendering the antibiotics inactive [111]. Notably, several Mycobacterium spp., such as M.tb, M. kansasii, M. fortuitum, and M. smegmatis, have been shown to produce N-acetyltransferase. In contrast, the MAB_2385 gene encodes 3′-O-phosphotransferase, which is responsible for inactivating STR and resistance in M. abscessus. The binding sites for aminoglycoside antibiotics, such as capreomycin and viomycin, are located within the ribosomal subunit interface of 16S rRNA helix 44 and 23S rRNA helix 69. Nucleotides, including C1409 of 16S rRNA and C1920 of 23S rRNA, are methylated at the 2’-hydroxyl. Methyltransferase tlyA deactivation leads to unmethylated ribosomes, inactivating capreomycin and viomycin [113, 114].
Azithromycin and clarithromycin belong to the macrolide chemical class and interfere with antibacterial protein synthesis by inhibiting 50S ribosomal subunits [34, 35]. M. chelonae, M. abscessus, and M. fortuitum exhibit methylase gene resistance. The transcriptional regulator, WhiB7, regulates expression of the erm (41) gene, which is responsible for macrolide-induced resistance. Azithromycin exposure is more accountable for rapid macrolide resistance in M. abscessus than clarithromycin [115]. Also, the erm gene is responsible for explicit methylation of the 23S rRNA A2058 nucleotide, which leads to reduced ribosome affinity to macrolides and thus leads to macrolide resistance [111, 116]. Interestingly, deleting erm 41 genes in M. massiliense and M. abscessus leads to increased susceptibility to the macrolide and lowers the MIC [111]. In addition to the above-mentioned enzymes, Mycobacterium also biosynthesizes RIF ADP-ribosyltransferase, which attaches an ADP-ribose unit to a hydroxyl residue at position 23 of RIF, which leads to rifamycin resistance in Mycobacterium spp. [117].
M.tb Rv1026 (ppx2) is a novel exopolyphosphatase responsible for hydrolytic activity in the long-chain poly(P). M.tb knockdown with Rv1026 expression displays slow growth, poly(P) accumulation, and increased cell wall thickness. These factors lead to decreased INH activity and enhanced resilience towards heat and acidic pH, and Mycobacterium spp. can increase intracellular macrophage infection at a speedy rate [118].
Biofilm-mediated antibiotic resistance in Mycobacterium spp.
Biofilms are a three-dimensional extracellular matrix made up of polymeric substances (EPSs) used to cover the microorganisms. This matrix is also known as a “safe haven” for microorganisms because the matrix provides a biological barrier against antimicrobial agents and hosts immune component cells. Notably, under biofilm protection, the microorganism not only survives the effects of antibiotics but can resume growth after withdrawal of antibiotics [119]. Several studies have suggested that efflux pumps are critical in intrinsic drug resistance, acquired drug resistance, and biofilm formation [120]. Also, efflux pump overexpression enhances biofilm formation [121]. For example, Rv1877, a protein belonging to M.tb major facilitator superfamily (MFS). Rv1877 expression in Escherichia coli results in an increase in resistance to several classes of antibiotics. Additionally, Rv1877 expression increases biofilm formation in E. coli. The above finding suggests that EPIs, potentially inhibiting biofilm formation and impairing the activity of the efflux pump, can be a promising strategy to restore the activity of existing antibiotics [121].
Efflux pumps and gene involvement in antibiotic resistance among Mycobacterium spp.
Efflux is a physiologic procedure that expels foreign substances and endogenous metabolic waste through an efflux pump to maintain normal cell function [122]. The increased bacterial resistance to several antibiotic classes is primarily due to enhanced expression of multi-drug efflux mechanisms [123]. Efflux can transport numerous substrates out of cells or specific types of antibiotics [124].
The bacterial efflux pumps are classified into five different superfamilies of proteins, including ATP-binding cassette (ABC), small multi-drug resistance (SMR), multi-drug and toxic compound extrusion (MATE), MFS, and resistance-nodulation-division (RND) [66, 125, 126]. Efflux pumps belonging to ABC, MFS, RND, and SMR families are present in Mycobacterium [122]. Transporters require energy to extrude the substrates outside cells. The ABC superfamily uses ATP as an energy source and other transporters utilise the transmembrane PMF or an Na+/H+ ion electrochemical gradient to move substrates from the pumps [66, 125, 126].
A recent study conducted by Kanji et al. discovered SNPs in efflux pump genes. According to a comparative analysis, the detected alleles are either exclusive to DS-TB strains or found in XDR strains. Importantly, XDR-TB expresses more Rv2688 and drrB than DS-TB strains [127]. Another study reported epistatic relations between the drug-resistance gene in MDR and XDR-TB [128].
ABC transporters
Eukaryotes and prokaryotes have ABC protein transporters. These transporters extrude substances across biological membranes, including ions, antibiotics, polysaccharides, peptides, amino acids, and proteins [129]. The ABC superfamily transporter-encoding genes are Rv1456c, Rv1457c, Rv1458c, Rv1217c, Rv1218c, and Rv1473. M.tb clinical isolates overexpressing Rv1456c, Rv1457c, and Rv1458c are responsible for RIF, INH, STR, and EMB resistance. In contrast, Rv1473 is involved in M. smegmatis macrolide resistance via efflux and the transcription factor, WhiB7, positively regulates Rv1473 on exposure to macrolides [129]. MSMEG-3762 and Rv1687c are annotated as ATP-binding proteins of the ABC efflux pump in M. smegmatis and M.tb, respectively. Siena et al. demonstrated that M. smegmatis contains the MSMEG-3762/63 protein complex, which acts as an efflux pump. This efflux pump has been shown to be responsible for effluxing RIF and ciprofloxacin, and biofilm development [130]. M.tb and M. smegmatis with the Rv2686c-Rv2687c-Rv2688c locus are involved in ciprofloxacin resistance, and to some extent, other FQs, such as norfloxacin, moxifloxacin, and sparfloxacin. Notably, FQ resistance can be decreased by reserpine, CCCP, and verapamil. Rv1217c and Rv1218c overexpression in M.tb results in higher RIF MICs, whereas overexpression of Rv1218c alone leads to a higher INH MIC in M.tb [131]. Rv1217c and Rv1218c in M.tb are significantly upregulated during biofilm formation. Compounds, such as piperine and 1-(1-naphthylmethyl)-piperazine (NMP), inhibit the dose-dependent expression of Rv1217c and Rv1218c genes. This finding suggests that EPIs, such as piperine and NMP, can be used as adjuvants to primary anti-TB drugs for biofilm inhibition [132]. Liu et al. carried out point mutations in M.tb (V219A and S292L) in Rv1258c. Interestingly, mutated V219A and S292L are responsible for PZA, INH, and STR resistance in M.tb. Notably, the V219A point mutation is responsible for low PZA, INH, and STR resistance, whereas the S292L mutation is associated with increased PZA, INH, and STR resistance. Piperine reduces INH and PZA resistance in the S292L mutant but not in the V219A mutant [133].
The Rv1258c (TAP-2) efflux pump encodes tetracycline/aminoglycosides resistance in M.tb and is implicated in drug resistance to RIF, INH, FQs, and aminoglycosides [134]. EPIs, such as verapamil and piperine, inhibit Rv1258c. Cloete et al. performed molecular modeling and simulation studies of Rv1258c and reported the probable binding sites of the substrate and inhibitors. The substrate, which has similar binding, was effluxed by the Rv1258c pump and inhibited efflux by piperine. The modeling studies revealed that spectinamide binds differently than other substrates, suggesting spectinamide can escape Rv1258c efflux pump activity [134].
MFS transporters
MFS proteins identified in bacteria include Rv2209, Rv3331, Rv1902c, Rv1672c, Rv0191, Rv0849, Rv2456c, Rv0037c, Rv0842, Rv0876c, Rv1258c, Rv2508c, Rv0783c, Rv1250, Rv1410c, Rv1634, Rv1877, Rv2333c, Rv2459, Rv2846c, Rv3239c, Rv3728, Rv2265, Rv1200, Rv3476c, Rv0261c, Rv0267, Rv1737c, Rv2329c, and Rv2994 [121].
EfpA is part of the QacA transporter family, which belongs to the MFS. Several studies have confirmed upregulation of EfpA in several clinical M.tb strains, including the M.tb strains resistant to INH, RIF, OFX, or multidrug treatments. BRD-8000.3 and BRD-9327 are potent inhibitors of EfpA [135, 136]. Recently, Wang et al. reported that the cryo-EM structures of EfpA in an outward open conformation bind to endogenous lipids or inhibitor BRD-8000.3. Three lipids in EfpA extend from the inner to the outer leaflet of the membrane. BRD-8000.3 occupies a lipid site within the inner membrane leaflet, which competitively inhibits lipid binding. Based on this study, Wang et al. stated that the EfpA structure is similar to the lysophospholipid transporter, MFSD2A, and the lipid-binding site functions as lipid flippase. Notably, the inhibitor, BRD-8000.3, prevents lipid entry into the pathway, while BRD-9327 likely disrupts the lipid flipping process between site A and site B [137].
Umar et al. evaluated the impact of anti-tubercular drugs on the mRNA efflux pump gene, Rv1250, in M.tb patients. A polymerase chain reaction revealed that the susceptible (n = 11) and resistant (n = 40) groups had different levels of Rv1250 mRNA expression (5.961 ± 0.414 and 10.192 ± 1.978 fold changes; P < 0.05). The extent of Rv1250 expression depends upon exposure to antimicrobial drug treatments [138]. Chloramphenicol resistance is linked to Rv0191 of the MFS transporter. Notably, Rv1353c controls Rv0191 transcription. Hence, when Rv1353c is overexpressed, transcription of Rv0191 is downregulated, making the strain more susceptible to chloramphenicol [139]. The TetR-related transcriptional repressor, LfrR, regulates the primary facilitator family gene LfrA expression, which actively extrudes FQs, cationic dyes, and anthracyclines out of the cell and promotes innate resistance to antibiotics. The increased expression of LfrA has been shown to be responsible for FQ resistance [139].
EfpA belongs to the MFS family and is highly conserved in M.tb and slow- and fast-growing Mycobacterium spp. M.tb with EfpA overexpression is responsible for resistance to FQs, INH, RIF, tetracycline, and clofazimine [140]. A recent study demonstrated that controlled expression of the EfpA gene in M. smegmatis increases drug tolerance by 32-fold to RIF, INH, STR, and amikacin. At the same time, M. smegmatis and M. bovis BCG overexpressing EfpA have higher drug tolerance to moxifloxacin by >180- and 8-fold, respectively [141].
There are 112 M. smegmatis mc2155 MFS members, which typically include 6–15 conserved transmembrane domains. The MSMEG_3705 gene of M. smegmatis mc2155 encodes a transporter protein containing 12 alpha-helical transmembrane domains. Zhang et al. deleted MSMEG_3705, which was designated as M. smegmatis mc2155 △MSMEG_3705. Interestingly, the mutant strain, M. smegmatis mc2155 △MSMEG_3705, is more sensitive to capreomycin and accumulates more EtBr than the wild strain. Deleting MSMEG_3705 also influences and affects bacterial growth. Deleting MSMEG_3705 causes upregulation of MSMEG_3706, a gene that encodes an isocitrate lyase (ICL), and downstream MSMEG_3705, which might be responsible for faster growth in the mutant strain (M. smegmatis mc2155 △MSMEG_3705) [142].
Small multidrug resistance (SMR) family
M.tb TBsmr (previously mmr [Rv3065]) of the SMR superfamily has approximately 43% sequence homology to E. coli EmrE. Antibiotics and fluorescent dyes are among the substances that TBsmr, a secondary active multi-drug transporter, transports [143]. Rv3066 controls Mmr expression [144]. Additionally, it has been shown that Mmr is significantly overexpressed in M.tb strains exposed to INH. Overexpressed mmr reduces sensitivity to safranin O, acriflavine, and EtBr in M.tb, and simultaneously verapamil and CCCP restores responsiveness [145].
Resistance-nodulation-division (RND) Superfamily
The RND family efflux pumps, such as membrane proteins large (MmpL) and mycobacterial small membrane protein (MmpS), are present in M.tb. The genomic sequence of M.tb H37Rv reveals 14 types of mmpL genes. Different species have different numbers of mmpL genes. For example, M. leprae has 5 mmpL genes, whereas M. abscessus has 31 mmpL genes. Specifically, MmpS5/MmpL5 is responsible for an efflux of azoles, clofazimine, BDQ, and ionophores [125]. Notably, alteration in the Rv0678 gene is involved in upregulating the mmpS5-mmpL5 genes. Importantly, M.tb with MmpS5-MmL5 overexpression shows resistance to INH, RIF, and diarylquinolines [140]. M. smegmatis cell overexpression of the mmpL7 gene leads to INH efflux and is responsible for INH resistance. The level of INH resistance is reduced in the presence of reserpine and CCCP [140].
Previous work has shown that the Mmr transporter is overexpressed in strains exposed to INH and mmr knockout mutants have enhanced sensitivity to ethidium bromide, tetraphenylphosphonium, and cetyltrimethylammonium bromide (CTAB). Overexpressed mmr reduces sensitivity to EtBr, acriflavine, and safranin O, and this effect is decreased in the presence of verapamil and CCCP. Importantly, INH susceptibility in M.tb is independent of mmr expression [145].
The role of MmpL3, MmpL7, and critical genes, such as katG, inhA, and nat, was studied to understand INH resistance in M.tb. The investigation suggested that the katG gene has a substitution mutation AGC-ACC at codon 315, the inhA promoter expresses C replacement at nucleotide position 15, and a silent mutation GGA to GGG at codon 207 in the nat gene. Taken together, these mutations are thought to be the factors responsible for INH resistance. Remarkably, no isolates exhibited mutations in MmpL3 and MmpL7 [146]. The TetR family transcription factor, MAB_2299c gene, which codes for MmpS-MmpL efflux pumps, controls MAB_2300-MAB_2301 and MAB_1135c-MAB_1134c expression. Deleting MAB_1135c-1134c in M.tb increases susceptibility to clofazimine and BDQ. MAB_1135c-1134c encodes MmpS-MmpL in M. abscessus. Interestingly, deletion of MAB_2300-MAB_2301 and MAB_1135c- MAB_1134c genes in a mutant lacking MAB_2299c and MAB_2300-MAB_2301 exacerbate strain sensitivity to clofazimine and BDQ [147].
Several studies have demonstrated that MmpS4/MmpL4 and MmpS5/MmpL5 constitute part of the siderophore export system in M.tb. The MmpS4/MmpL4 and MmpS5/MmpL5 systems export and recycle siderophores, including mycobactins and carboxymycobactins, in M.tb [125]. A study was conducted using green fluorescent protein (GFP), which revealed that MmpL5 forms a homotrimeric functional complex in the inner membrane in the presence of MmpS5. MmpS5 expression facilitates assembly of monomeric MmpL5 into a homotrimer, which promotes extrusion of several antimycobacterial drugs [148]. The intracellular concentration of 3H-radiolabelled econazole was measured in M. bovis. The MmpS5/MmpL5 system was responsible for decreasing the concentration of econazole in the M. bovis, and after the addition of CCCP, the concentration was maintained inside M.bovis. This finding suggested that econazole is a substrate for the MmpS5/MmpL5 system in M. bovis [125]. Farina et al. studied the role of MmpS5/MmpL5 and its gene regulator (Rv0678) in MDR isolates. Genetic mutations in MmpL5 (Rv0676) due to insertion, deletion, and substitution could change the functionality of the efflux pump and cause a significant reduction in drug susceptibility, thus developing drug resistance in M.tb strains [149]. In another study, clofazimine-resistant M.tb strains were isolated, unexpectedly showing cross-resistance to BDQ. Furthermore, genomic analysis revealed that mutations in the transcriptional regulator, Rv0678, were shown to be responsible for the concomitant upregulation of the efflux pump, MmpL5, and thus were responsible for BDQ resistance [150]. Additionally, other studies have confirmed the role of the MmpS5-MmpL5 efflux system thiacetazones, azoles, and imidazo[1,2-b][1,2,4,5]tetrazines, and spiroketal indolyl Mannich bases [151, 152].
Overexpression of genes and transporters
The T28 erm(41) gene, positive regulatory gene, whiB7, and efflux genes, including MAB_2355c, MAB_1409c, and MAB_1846, are responsible for enhanced clarithromycin resistance [153]. Notably, MAB_2355c belongs to the ABC-F family protein and is involved in ribosome protection. MAB_2355c-deleted strains have increased sensitivity to macrolide [154]. In addition, M. abscessus strains exposed to clarithromycin had efflux pump gene (MAB_3142 and MAB_1409) overexpression. However, verapamil increases clarithromycin susceptibility to M. abscessus by 4- to ≥64-fold. The above finding suggests that T28 erm(41) polymorphisms and overexpressed efflux genes are responsible for macrolide resistance in M. abscessus subsp. abscessus and bolletii [155].
Rodrigues et al. exposed M. bovis BCG to INH to induce INH-resistant strains, which demonstrated an increased extrusion activity and enhanced expression of efpA, mmpL7, mmr, p55, and the Tap-like gene, Rv1258c. Notably, EPIs (chlorpromazine, thioridazine, and verapamil) decreased INH resistance in INH-induced resistance [156]. Overexpression of jefA (Rv2459) in M.tb showed resistance to INH and EMB [157].
Overexpression of genes in Mycobacterium, including Rv0191, Rv3756c, Rv3008, and Rv1667c, are responsible for resistance to PZA and pyrazinoic acid. Notably, reserpine, piperine, and verapamil increase M.tb strain susceptibility to PZA, having overexpressed Rv0191, Rv3756c, Rv3008, and Rv1667c [158]. Zimic et al. determined the vulnerability of M.tb and M. smegmatis based on the efflux of pyrazinoic acid. Zimic et al. reported the efflux of pyrazinoic acid was within the range and resistant M.tb strains exhibited below this range. Also, pyrazinoic acid efflux was higher in M. smegmatis than M.tb [159].
Caleffi-Ferracioli et al. exposed M.tb H37Rv to RIF, verapamil, and RIF + verapamil for 16 h and 72 h. Expression of efflux genes was determined after 72 h, which included Rv1456, Rv3065, Rv1458, Rv1457, Rv2846, Rv1258, Rv2942, Rv1217, Rv2459 and Rv1410 for RIF, Rv3065, Rv1457, and Rv1410 for verapamil, and Rv1457, Rv2846, Rv1258 and Rv2942 for RIF + verapamil. Treatment with RIF + verapamil lowered all the efflux pump genes, except for Rv2846, compared to RIF. Additionally, combining these drugs leads to cumulative morphologic changes (wrinkles and rounding) to M.tb [160].
In another study, Narang et al. evaluated the role of MmpL2 (Rv0507), MmpL5 (Rv0676c), Rv0194, and Rv1250 in RIF-resistant M.tb. The RIF-resistant strain had an Rv0194 induction of 50%, mmpL5 induction of 44%, Rv1250 induction of 25%, and mmpL2 induction of 12.5%. RIF-susceptible isolates exhibited Rv0194 induction of 18%, mmpL5 induction of 9%, Rv1250 induction of 18%, and mmpL2 induction of 9%. This study found There was no combination role for Rv0194, MmpL2, MmpL5, and Rv1250 genes in RIF resistance. However, the RIF-resistant isolates had more Rv0194 and mmpL5 overexpression than RIF-susceptible isolates [161].
THP-1 infected with M.tb H37Rv were treated with RIF and verapamil alone and in combination. RIF exposure led to enhanced expression of efflux pump genes in the SMR, ABC, and MFS families. In association with verapamil, RIF led to downregulation of eight efflux genes, which suggests that verapamil exhibits efflux pump inhibitory activity inside the macrophage and thus could be beneficial in treating TB in humans [162].
Li et al. reported that RIF-monoresistant M.tb isolates had mutations in rpoB overexpressed with one or two of efflux pump genes, such as Rv2333, drrB, drrC, Rv0842, bacA, and efpA. Notably, CCCP, VP, TZ, and CPZ modulated RIF activity in clinical strains [163].
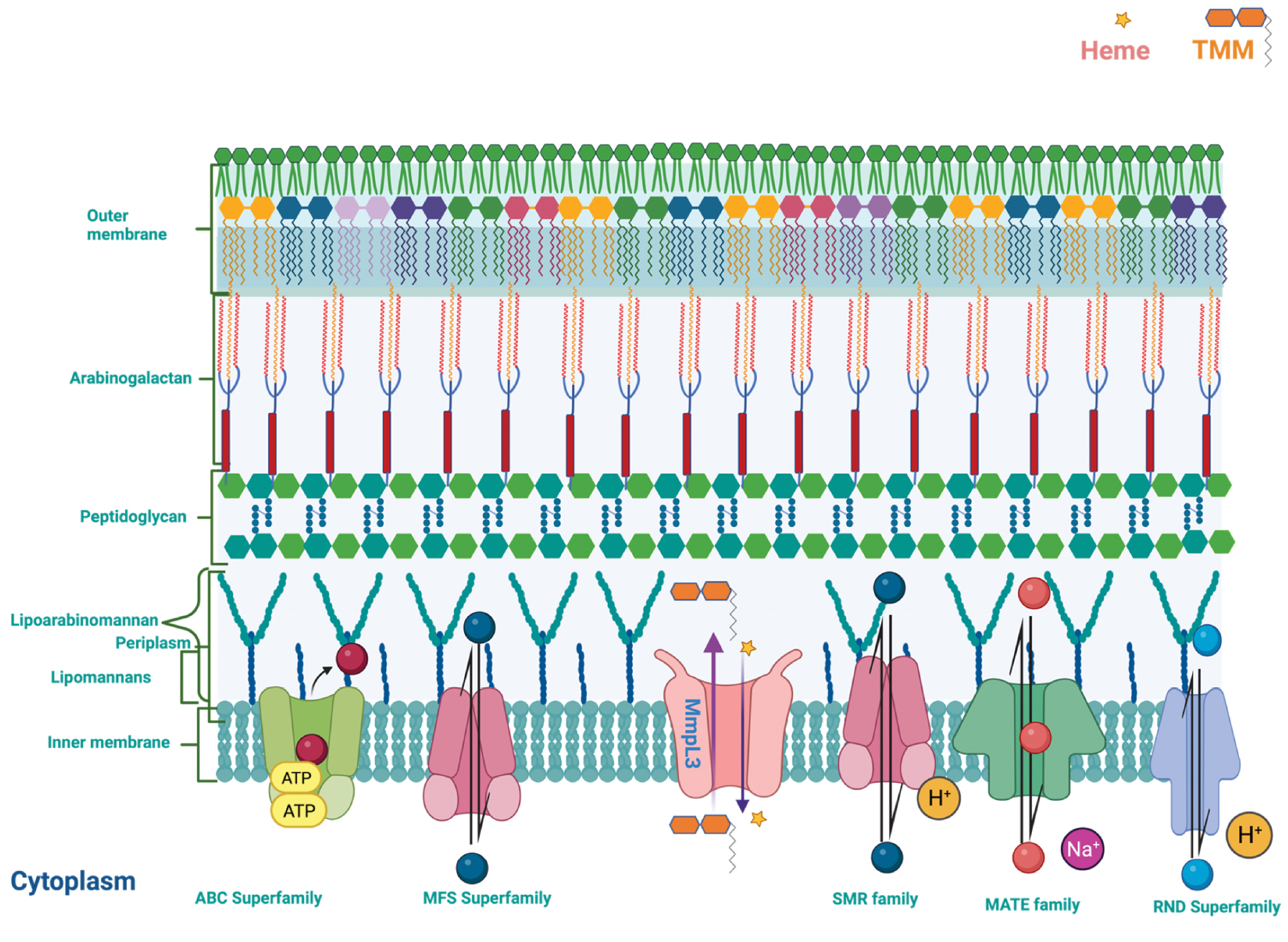
Machado et al. investigated the relationship between mutations and the role of efflux pump genes in drug resistance. After exposure of anti-TB drugs to M.tb isolates, the efflux pump genes (mmr, mmpL7, Rv1258c, p55, and efpA) were shown to be overexpressed. Additionally, these strains had mutations, including C-15T (inhA prom), S94A (inhA ORF), I194T (inhA ORF), S315T (katG), D735A (katG), S315T (katG), S531L (rpoB RRDR), D516Y (rpoB RRDR), A1401G (rrs 1400 region), wt/A1401G* (rrs 1400 region), G-10A (eis prom), ins GT at POS 755/756 (tly), D94N (gyrA QRDR), A90V (gyrA QRDR), and S91P (gyrA QRDR). Efflux of the substrates was reduced using EPIs. However, these EPIs were not able to reverse antibiotic resistance [164]. Therefore, the level of drug resistance seen in these strains resulted from efflux activity and a mutation in a target gene for drug resistance [164]. Figure 4 depicts the Mycobacterium membrane and efflux pumps. The efflux pump superfamily and the efflux pumps with substrate and inhibitors are listed in Table 5.
Figure 4 A schematic representation depicting the Mycobacterium membrane and the efflux pump families.
Table 5 Efflux Pump Transporter Family and Efflux Pumps with their Substrate and known Inhibitors
| Efflux Pump Transporter Family | Efflux Pump | Antibiotic (substrates) | Efflux Pump Inhibitors | Ref. |
|---|---|---|---|---|
| ABC superfamily | Rv2686c/87c/88c | Floroquinolones (ciprofloxacin, norfloxacin, moxifloxacin, and sparfloxacin) | Carbonyl cyanide m-chlorophenyl hydrazone, reserpine, and verapamil | [279] |
| MSMEG_3762/63/65 | Rifampicin and ciprofloxacin | [130] | ||
| drrA/drrB | Rifampicin and isoniazid | Reserpin and, verapamil | [280] | |
| Rv1473 | Roxithromycin, erythromycin, and azithromycin | Carbonyl cyanide m-chlorophenyl hydrazone, reserpine, an verapamil | [129] | |
| Rv1218c | Isoniazid, rifampicin, moxifloxacin, linezolid, bedaquiline, PA-824, and lofazimine | Piperine and 1-(1-naphthylmethyl)-piperazine | [132] | |
| Rv1217c | Rifampicin | Piperine ane 1-(1-naphthylmethyl)-piperazine | [131] | |
| Rv1218c | Isoniazid | Piperine and 1-(1-naphthylmethyl)-piperazine | [131] | |
| MAV_3306 | Azithromycin | [281] | ||
| Rv0191 | Pyrazinamide | Reserpine, verapamil, and piperine | [158] | |
| Rv1667c | Pyrazinamide | Reserpine, verapamil, and piperine | [158] | |
| Rv3756c | Pyrazinamide and pyrazinoic acid | Reserpine, verapamil, an piperine | [158, 159] | |
| Rv0194 | RIF | Reserpine and verapamil | [161] | |
| MFS superfamily | MAV_1406 | Azithromycin | [281] | |
| Rv0849 | ||||
| Rv0191 | Pyrazinamide, pyrazinoic acid, and chloramphenicol | Reserpine, verapamil, and piperine | [139] | |
| Rv0783 | [156, 167] | |||
| Rv0849 | [156, 167, 282] | |||
| Rv1258c | Rifampicin, isoniazid, and kanamycin | Piperine and verapamil | [133, 156, 167, 282] | |
| Rv1634 | Florroquinolones | [121] | ||
| Rv2333c | Tetracylinees | [121] | ||
| Rv2459 | Isoniazid, | Verapamil | [121, 157, 160] | |
| Rv2846c | [121] | |||
| Rv2994 | [121] | |||
| Rv3008 | Pyrazinamide and pyrazinoic acid | [158] | ||
| Rv3728 | [121] | |||
| EfpA | Verapamil | [156] | ||
| lfrA | [283] | |||
| SMR | Rv3065 | Safranin O, acriflavine, and ethidium bromide | Verapamil and carbonyl cyanide m-chlorophenyl hydrazone | [123] |
| Mmr | Ethidium bromide, acriflavine, and safranin O | Verapamil and carbonyl cyanide m-chlorophenyl hydrazone | [145] | |
| RND superfamily | MmpS5/MmpL5 (Rv0678) | Azoles, clofazimine, bedaquiline, ionophores isoniazid, and rifampicin, | BM212, SQ109, indolcarboxamides, THPPs, and adamantyl ureas | [125] |
| MmpL7 | Isoniazid | Carbonyl cyanide m-chlorophenyl hydrazone, and reserpine | [284] | |
| Rv2942 | Rifampicin | Verapamil | [160] |
EPIs
EPIs target bacterial efflux proteins to prevent the extrusion of substrates from the bacterial membrane. When used in combination with antibiotics, EPIs offer a promising strategy to combat efflux pump activity. The benefits of EPIs include the following: (i) inhibiting the efflux of anti-TB drugs, thereby reversing drug resistance in M.tb; (ii) producing synergistic effects with anti-TB medications; and (iii) reducing the required dosage of the drugs [165, 166].
Natural products and analogs acting as Mycobacterium efflux pump inhibitors
Piperine (1) is an alkaloid that exhibits moderate antimycobacterial activity against M.tb H37Rv with an MIC value of 128 μg/mL. Immunomodulatory activity, and in combination with rifampicin, improved efficacy and decreased bacterial loads in the lungs of infected mice were shown [167]. Moreover, piperine showed inhibitory activity against Rv1258c, a putative multidrug efflux pump of M.tb. Piperine reduced the MIC and the mutation prevention rifampicin concentration in M.tb H37Rv and MDR M.tb clinical isolates. Furthermore, piperine enhanced the bactericidal activity of rifampicin and significantly extended the post-antibiotic effect [168]. In another study, piperine showed synergistic activity with streptomycin against M.tb H37Rv [169].
Farnesol (2 [32 μg/mL]), in combination with ethidium bromide (EtBr), modulates farnesol (2) activity, reduces the MIC by 4-fold, and enhances the accumulation of EtBr in M. smegmatis. Moreover, farnesol (2) exhibits synergism with RIF against M. smegmatis [170].
Reserpine (3; Figure 5) has anti-mycobacterial activity against M.tb H37Rv with an MIC value of 128 μg/mL [171]. Reserpine inhibits the efflux pump of M.tb and increases the intracellular concentration of the ciprofloxacin (MIC value of 4 μg/mL against MDR M.tb) [171]. The reserpine mode of action reduces drug excretion and decreases the energy supply through ATP hydrolysis. Reserpine increases the intracellular concentration of ciprofloxacin in M.tb and susceptibility to bacilli [172].
Figure 5 Chemical structures of natural products as EPIs.
Gröblacher et al. isolated diarylheptanoid scaffold compounds, including trans,trans-1,7-diphenylhep-ta-4,6-dien-3-one (4; Figure 5), (5R)-trans-1,7-diphenyl-5-hydroxyhept-6-en-3-one (5; Figure 5), (3S,5S)-trans-1,7-diphe-nylhept-1-ene-3,5-diol (6; Figure 5), and in the flavonoid category, pinocembrin (7; Figure 5), from Alpinia katsumadai belonging to family Zingiberaceae [173]. The identified compoundsefflux pump inhibitory and anti-mycobacterial activities were assessed in M. smegmatis mc2 155. Compounds (4–7) inhibited M. smegmatis mc2 155 with MIC values of ≥128, 64, ≥128, and 128 mg/L, respectively. Compound 4 (64 mg/L), 5 (32 mg/L), 6 (64 mg/L), and 7 (64 mg/L) modulated RIF activity by 4- to 8-, 4-, 4-, and 2-fold, EMB activity by 1-, 1- to 2-, 2-, and 2- to 4-fold, INH by 1-, 1-, 2-, and 1-fold, ciprofloxacin activity by 1-, 2-, 2-, and 1- to 2-fold, and EtBr activity by 4-, 8-, 8-, and 4-fold, respectively [173]. Although diarylheptanoids and pinocembrin exhibited weak anti-mycobacterial activity, these compounds accumulated EtBr and decreased the RIF MIC. Thus, the diarylheptanoids class of compounds could be developed to reverse anti-mycobacterial drug resistance. Furthermore, compounds 4–7 should be evaluated in other Mycobacterium spp. and in vivo models of TB infection. Also, the safety should be considered in the cell model of TB infection [173].
de Vasconcelos et al. evaluated carvacrol (8; Figure 5) and its derivatives for efflux pump inhibitory activity. Compound 8 promoted the accumulation of EtBr in M.tb H37Rv, which exhibits a relative final fluorescence (RFF) of 0.38 (similar to verapamil with an RFF of 0.20). Carvacrol displayed anti-mycobacterial activity against M.tb H37Rv with an MIC value of 76 μg/mL. Carvacrol demonstrated synergistic activity with RIF. However, carvacrol derivatives showed no anti-mycobacterial activity [174]. Additionally, compound 8 demonstrated EPI activity in several strains, including Salmonella enteritidis 1129 strains, S. typhimurium LT2 DT104, E. coli ATCCR 35218, and Staphylococcus aureus ATCCR 25923 [175]. Sieniawska et al. evaluated natural terpenes (R-limonene [9; Figure 5]), S-limonene (10; Figure 5), myrcene (11; Figure 5), sabinene (12; Figure 5), and β-elemene (13; Figure 5) in combination with first-line anti-TB drugs [176]. compounds 9–13 showed anti-mycobacterial activity against M.tb with MIC values of 64, 64, 32, 32, and 2 μg/mL. Of all the compounds examined, compound 10 showed the most synergistic efficacy against M. tb with all first-line anti-TB drugs. Compound 10 modulated the EMB MIC from 16 to 0.475 μg/mL, the RIF MIC from 16 to 0.237 μg/mL, and the INH MIC from 32 to 0.475 μg/mL. Compounds 9 and 11–13, and 12, in association with first-line antibiotics, modulated the antibiotic MIC in M.tb (from 3.9 to 0.475 μg/mL for EMB, 15 to 0.475 μg/mL for INH, and 0.475 to 0.237 μg/mL for RIF). However, α-pinene in combination with EMB and INH showed the opposite effect in M.tb and increased MIC values from 16 to 125 μg/mL for EMB and from 32 to 125 μg/mL for INH [176].
Solnier et al. evaluated flavonoids (skullcapflavone II [14; Figure 5], nobiletin [15; Figure 5], tangeretin [16; Figure 5], baicalein [17; Figure 5], and wogonin [18; Figure 5]) against M. smegmatis mc2 155, M. aurum ATCC 23366, and M. bovis BCG ATCC 35734 for anti-mycobacterial and efflux pump inhibition activity [177]. The anti-mycobacterial activity results indicated that compounds 15–18 had an MIC99 value ≤31.25 mg/L against M. aurum and compound 14 had an MIC99 value of 7.8 mg/L. Compounds14, 15, and 18 had anti-mycobacterial activity against M. smegmatis mc2 155 with an MIC99 value ≤128 mg/L and compound 17 inhibits M. smegmatis mc2 155 with an MIC99 value of 31.25 mg/L. Compounds 14–18 had activities against M. bovis BCG ATCC 35734 with MIC99 values of 31.25, 62.5, 31.25, 250, and 500 mg/L, respectively. Among these flavonoids, compounds 14 (32 mg/L), 15 (32 mg/L), 16 (32 mg/L), 17 (8 mg/L), and 18 (32 mg/L) modulated the activity of EtBr and RIF in M. smegmatis mc2 155 with modulation factors of 128 and 8, 2 and 32, 2 and 32, 8 and 32, and 4 and 16, respectively. These results suggested that flavonoids could be used as adjuvants in combination with RIF to reverse drug resistance [177].
Seven carvotacetones were extracted from Sphaeranthus africanus by Tran et al. and screened against the mycobacterial model strains Mycobacterium smegmatis mc2 155, M. aurum ATCC 23366, and M. bovis BCG ATCC 35734 for determination of anti-mycobacterial and EPI activities [178]. All carvotacetones exhibited moderate anti-mycobacterial activity against M. smegmatis mc2 155, ranging between 32 and 128 μg/mL. Representative carvotacetones (compound 19; Figure 5) and (compound 20; Figure 5) strongly modulated the activity of RIF and EtBr in M. smegmatis mc2 155. Compound 19 (at 1 and 16 mg/L concentrations) modulated EtBr activity by 8-fold and RIF activity by 2-fold in M. smegmatis mc2 155. Compound 20 (1 mg/mL) was only able to modulate EtBr activity 8-fold in M. smegmatis mc2 155. Moreover, compounds 19 and 20 inhibited M. aurum ATCC 23366 with MIC values of 85.9 and 78.5 μM and M. bovis BCG ATCC 35734 with MIC values of 42.9 and 78.5 μM, respectively [178].
Stephania tetrandra S. Moore’s root contains tetrandrine (compound 21; Figure 5). Tetrandrine is used in traditional Chinese medicine to treat angina pectoris, cardiac arrhythmia, and hypertension. Tetramdrome has calcium channel antagonist properties and interacts with the slowly gated K (Ca+2) channel and the voltage-activated L- and T-type Ca+2 channels with variable degrees of specificity and affinity [179]. Zhang et al. evaluated compound 21 against 200 isolates resistant to isoniazid and ethambutol. Incubation of compound 21 with M.tb strains (resistant to INH and EMB) showed no activity against growth at 30 μg/mL. Compound 21, in association with EMB, modulated EMB activity and reduced the MIC by > 64-fold from 80 μg/mL to 1.25 μg/mL. Also, Compound 21reduced the MIC of INH by > 32-fold from 8 μg/mL to 0.25 μg/mL. Notably, compound 21 showed a reversal in MIC in 82% resistant M.tb strains, which was more potent than verapamil. This finding suggested that compound 21 could be developed as an adjuvant in combination with first-line anti-TB drugs [180].
Synthetic compounds as Mycobacterium spp. efflux pump modulators
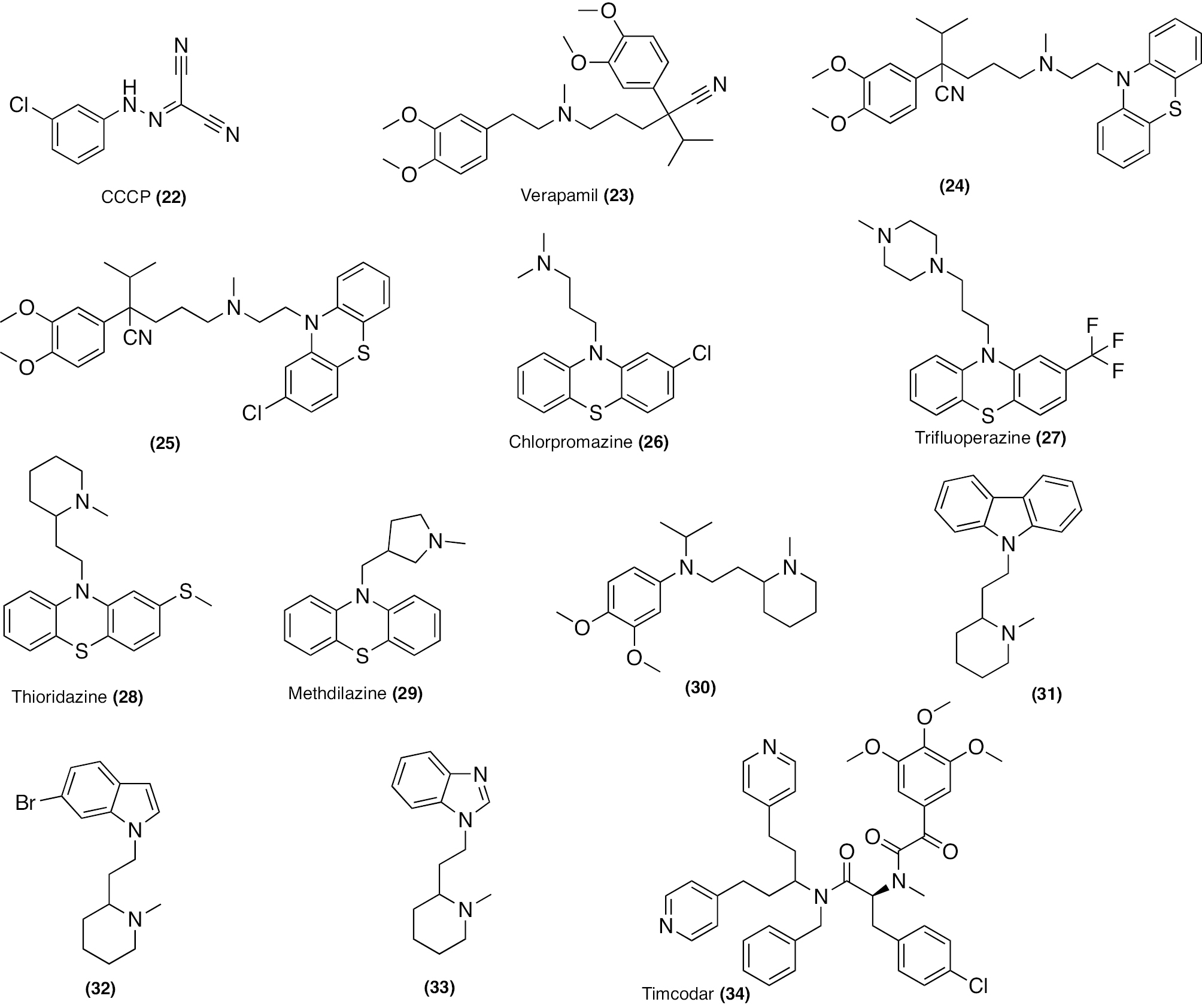
Proton influx into the cytosol is induced by the protonophore, CCCP (22; Figure 6). Compound 22 controls proton influx by generating electrochemical potentials from CCCP and CCCP– anions. This process is not dependent on the electrochemical H+ gradient. Furthermore, this action leads to disruption of energy sources in the membrane [181]. As a result of the diminished F1 F0 -ATP synthase activity, the principal active ABC transporters are also affected. All transport processes are hampered by a lack of energy, including importing and exporting nutrients through the cell membrane components. CCCP treatment decreases overexpression of the mmpL7 gene and can reverse the resistance associated with the mmpL7 gene [182].
Figure 6 Chemical structures of synthetic compounds acting as EPIs.
Clinically, verapamil (23; Figure 6) alleviates heart conditions. Verapamil moderately inhibits M.tb H37Rv with an MIC value of 128 μg/mL. Verapamil enhances the effectiveness of several anti-tubercular drugs in vivo and in vitro. The cationic amphiphile nature disrupts the bacterial membrane and creates a stress response in Mycobacterium spp. [183]. The homo-tetrameric bacterial Ca2+ channel inhibits ATP-binding cassette transporters and Ca2+ selectivity filter from the intracellular side [160, 184]. Moreover, verapamil modulates the potency of anti-TB drugs. Moreover, in association with RIF, verapamil helps lower the fractional inhibitory concentration index (FICI), reduces viable cell counts, and decrease efflux pump genes [160]. Verapamil also regulates downregulation of efflux pump genes in M.tb inside the macrophages and enhances the accumulation of ethidium bromide in the cells [162, 185]. The BDQ and clofazimine MICs were lowered by verapamil against drug-susceptible and -resistant M.tb strains [186]. Additionally, verapamil increases systemic BDQ exposure, which boosts the effectiveness of BDQ in treating murine TB infection. In contrast, verapamil does not influence the oral bioavailability or effectiveness of clofazimine in mice [187]. Moreover, verapamil increases Q203 activity against M.tb in vitro and ex vivo in an unidentified overexpressed efflux pump [188, 189]. Verapamil and its analogues (compounds 24 and 25; Figure 6) exhibit improved synergy with RIF and comparable anti-TB activity [190, 191].
Two lines of evidence indicate that the M.tb drug efflux pumps have a role in the drug tolerance induced by macrophages. In a virulent laboratory strain (CDC1551), the genetic knockdown of the MFS efflux pump Tap (Rv1258c) leads to loss of RIF tolerance, which is usually observed when M.tb infects macrophages. It has also been observed that drug tolerance induced by macrophages is inhibited by various structurally distinct drugs that are known to block bacterial efflux pumps [192]. Lake et al. reported that verapamil reduces M.tb RIF efflux and macrophage-induced RIF tolerance. Notably, verapamil and proton pump inhibitors (PPIs), including omeprazole, pantoprazole, lansoprazole, and lansoprazole sulfide, decrease RIF efflux, intramacrophage proliferation, and macrophage-induced tolerance. Most anti-TB drugs have shown macrophage-induced tolerance. It will be interesting to determine if anti-TB agents and PPIs reverse drug resistance and tolerance. Further, in vivo studies should be conducted to assess the translation value [193].
Initially, phenothiazines were identified as anti-psychotic, but scientists were subsequently curious about potential anti-mycobacterial effects [194]. According to the investigations, phenothiazines selectively block M.tb type II NADH dehydrogenases. As a result, phenothiazines hinders oxidative phosphorylation and the respiratory chain electron transfer from NADH to menaquinone. PMF and ATP depletion are caused by respiratory chain disturbance. Additionally, thioridazine enhances cell wall permeability by changing plasma membrane composition [184]. Among phenothiazines, chlorpromazine (26), trifluoperazine (27), thioridazine (28), and methdilazine (29; Figure 6) were shown to have anti-mycobacterial activity [195]. Kumar et al. synthesised hybrid compounds (having verapamil and tricyclic chemo-sensitizer core of phenothiazines), which exhibited intrinsic anti-mycobacterial and intracellular activities in macrophages [196]. In free form, thioridazine (28) is toxic to cells and zebrafish embryos. The encapsulation of thioridazine in poly(lactic-co-glycolic) acid nanoparticles results in no toxicity. Combined with RIF nanoparticles, the encapsulated thioridazine showed a slight boost in the ability of macrophages to kill M. bovis BCG and M.tb [197]. Thioridazine exhibits bactericidal against M. avium by inhibiting intracellular growth at 2X MIC by blocking EtBr efflux [198].
A panel of drug-resistant M.tb was used to assess the mycobacterial potential of EPIs with ion channel blocking capabilities, such as verapamil, thioridazine, chlorpromazine, flupenthixol, and haloperidol. These compounds demonstrate synergistic activity with INH and RIF and inhibit efflux pumps. In addition, these compounds in combination displayed activity against M.tb. The bactericidal action of compounds was due to blocking of ion channels, which decreased intracellular ATP levels. These substances also decreased the intracellular mycobacterial burden by activating lysosomal hydrolase and phagosome acidification. Therefore, ion channel inhibitors could be developed as adjuvants for treating TB caused by susceptible and resistant isolates [199].
Furthermore, Pieroni et al. synthesized hybrid scaffolds containing thioridazine, verapamil, and the representative synthesized compounds (30–33; Figure 6), demonstrated higher efflux inhibitory activity, less cytotoxicity, and a synergistic effect with anti-TB drugs in vitro and in intracellular macrophages [200]. Timcodar (34; Figure 6) showed weak inhibition against M.tb with an MIC value of 18.7 μg/mL. Synergistic activity with RIF, BDQ, and clofazimine was shown but not with other anti-TB agents. In addition, timcodar exhibited a 10-fold increase in growth inhibition against M.tb cultured in host macrophages and demonstrated synergy with RIF, moxifloxacin, and BDQ [201]. Moreover, the efficacy of RIF and INH was demonstrated in a mouse model of M.tb infection than each drug alone [163].
Li et al. reported lowering the RIF MIC in combination with EPIs (carbonyl cyanide m-chlorophenyl hydrazine, verapamil, thioridazine, and chlorpromazine) on RIF-monoresistant isolates (rpoB) overexpressed with one or two putative efflux pump genes (Rv2333, drrB, drrC, Rv0842, bacA, and efpA) [202].
M. abscessus subsp. abscessus was exposed to antibiotics, such as amikacin, cefoxitin, clarithromycin, clofazimine, and tigecycline, for 24 h in another study. RNA sequencing was evaluated after antibiotic exposure. All five antibiotics (amikacin, cefoxitin, clarithromycin, clofazimine, and tigecycline) induced expression of one or more genes responsible for efflux transporter systems. In addition, antibiotics acting through ribosomes exhibited the most pronounced effects in expressing genes responsible for efflux pumps. A time-kill kinetics study showed that amikacin, cefoxitin, clarithromycin, clofazimine, and tigecycline in combination with EPIs (thioridazine and verapamil) had synergistic activity. One interesting observation from this study was that ribosome-targeting antibiotics resulted in pronounced efflux transporter expression. This finding suggested that antibiotics targeting ribosomes could be explored for expression of efflux pumps and possible efflux pump inhibition activity [203].
Thirty-one phenotypically and genotypically identified M. abscessus subsp. abscessus, M. abscessus subsp. massiliense, and M. abscessus subsp. bolletii clinical isolates were screened for BDQ activity in the presence and absence of verapamil and reserpine. Interestingly, verapamil reduced the BDQ MIC in all 100% isolates of M. abscessus subsp. by modulating BDQ activity from 4- to 32-fold and achieved maximum activity in M. abscessus subsp. bolletii. While reserpine reduced the BDQ MIC in all 54.8% isolates of M. abscessus subsp. and modulated the activity by 2- and 4-fold. This finding suggested that BDQ could be developed as combination therapy with verapamil as an adjuvant for treating M. abscessus-associated infections in humans [204].
Menichini et al. evaluated clarithromycin in combination with berberine or CCCP, piperine, or tetrandrine in 12 MAC clinical isolates. Mutations were present in domain V of the 23S RNA gene that affect the activity of EPIs on clarithromycin-resistant strains of MAC. Specifically, it was observed that the nucleotide change (A→C at position 2059) in strains, such as MAH 2587, MAH 2854, and MAH 2915, maintains a resistant phenotype despite a significant reduction in the MIC of clarithromycin when EPIs (berberine, CCCP, and piperine) are present. In contrast, strain MAH 1565 exhibited a point mutation at position 2058, with a nucleotide change from A→G, which restored sensitivity to clarithromycin in the presence of tetrandrine. Additionally, strain MI 2895 exhibited a nucleotide change (A→T), resulting in an intermediate phenotype when treated with berberine and tetrandrine. The overall finding suggested that on treatment with EPIs (berberine, CCCP, piperine, or tetrandrinAC-resistant strain was made susceptible to clarithromycin [205].
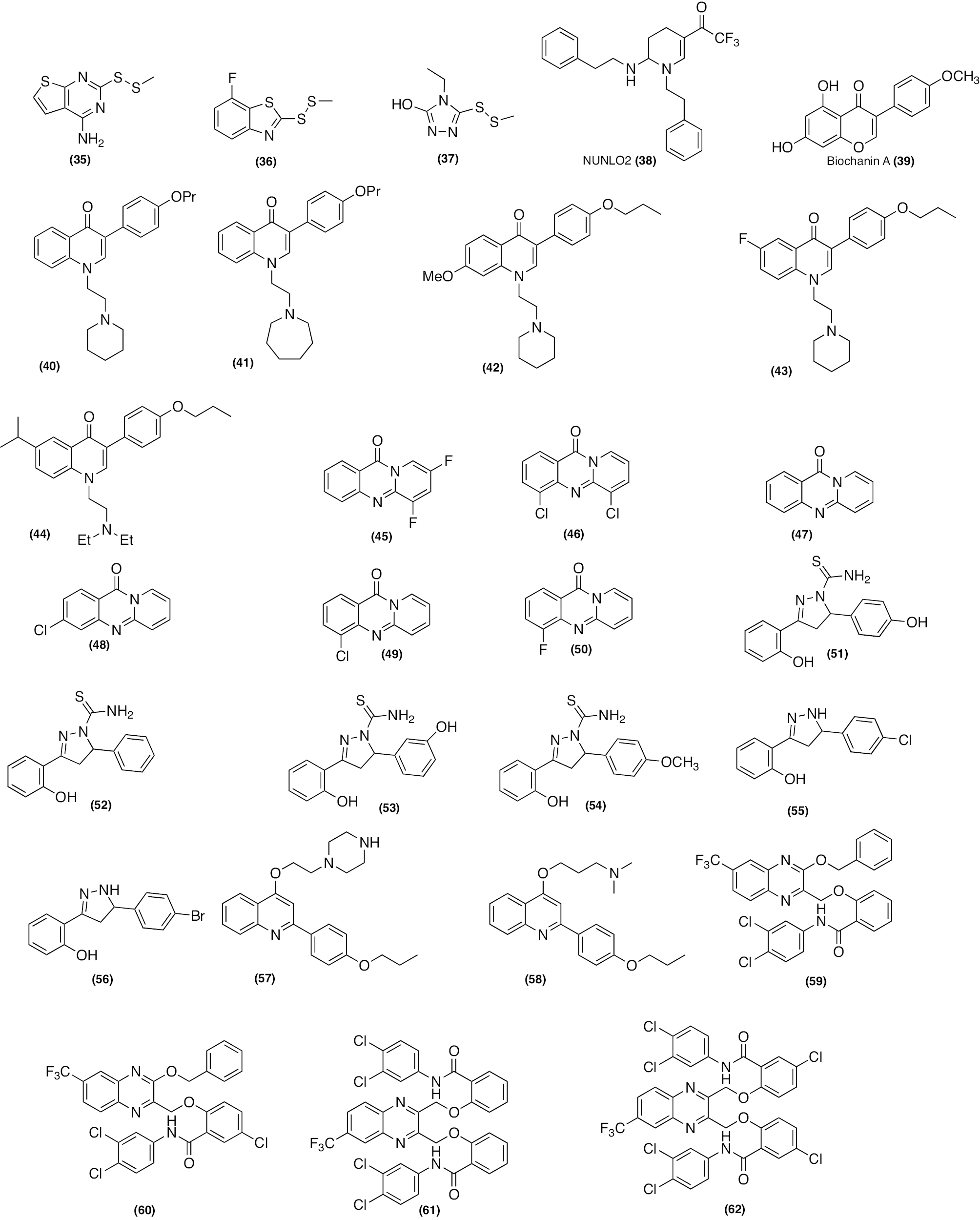
Disulfides are obtained from Allium Stipitatum and possess antibacterial properties. Disulfide analogues (35–37; Figure 7) showed anti-mycobacterial effects by suppressing the drug efflux pump and biofilm formation [206]. The category of synthetic compounds included tetrahydropyridine derivatives, which possess calcium channel regulators. With this in view, Halicki et al. evaluated 2,2,2-trifluoro-1-(1,4,5,6-tetrahydropyridin-3-yl) ethanone (NUNL02) [compound 38; Figure 7] in combination with anti-TB drugs against M.tb-resistant strains. Compound 38 inhibited susceptible M.tb H37Rv and FURG-2 (MDR) with MIC values of 25 and 12.5 μg/mL, respectively. Also, compound 38 (1.56 μg/mL) in association with RIF reduced the RIF MIC by 16-fold (from 256 to 16 μg/mL). Consequently, compound 38 makes the MDR-TB strains more susceptible to RIF. Notably, compound 38 increases EtBr accumulation in the FURG-2 strain more than susceptible M.tb H37Rv and the RFF value was 4-fold higher than verapamil. Interestingly, compound 38 exhibited sterilizing activity against M.tb H37Rv and was non-toxic to VERO cells. Furthermore, compound 38 should be evaluated in combination with RIF and another first-line anti-TB antibiotic in an animal model of TB to assess compound 38 efficacy [207].
Figure 7 Chemical structures of synthetic compounds acting as EPIs.
Biochanin A (39; Figure 7) belongs to isoflavone, which shows promising efflux pump inhibition in M. smegmatis. Cannalire et al. performed bioisosteric replacement by structural modification in biochanin A replacing oxygen with nitrogen [208]. In addition, the phenolic group was removed from the parent compound 39 and the 4′-methoxy group was substituted by a propyloxy, which resulted in the formation of a 2-phenylquinoline scaffold. Additionally, several 3-phenylquinoline derivatives were synthesized and evaluated for the ability to block the efflux pump in M. smegmatis mc2 155 and M. avium. The representative 3-phenyl quinoline derivatives (40; Figure 7) and (41; Figure 7) displayed the highest efflux pump activity. Compounds 40 and 41 modulated the activity of macrolides, including clarithromycin, erythromycin, azithromycin, and fluoroquinolones, such as ciprofloxacin and ofloxacin, and EtBr in M. avium strains. Importantly, modulation activity was better than standard verapamil, chlorpromazine, and thioridazine. Furthermore, preclinical and clinical studies are required to evaluate the safety and potency of 3-phenylquinoline to use as an adjuvant with antibiotics to reverse drug resistance in M. avium. This finding also suggested the combination of 3-phenylquinoline and FQs for treating M.tb infections in humans [208].
The 3-phenylquinolone compound class showed antibacterial activity against M. smegmatis and M. avium. 3-phenylquinolone also demonstrated activity against S. aureus, which overexpresses the NorA pump. Felicetti et al. modified the C6 and C7 positions of 3-phenylquinolone to improve the toxicity profile of the EPIs [209]. Representative compounds (42–44; Figure 7) showed synergistic activity with clarithromycin, ciprofloxacin, and EtBr at low concentrations against M. avium. A substantial synergistic action was demonstrated by compound 44 (4 μg/mL [1/32 MIC]) with clarithromycin, which also caused a 6-fold modulation in the MIC. In an ex-vivo animal harbouring M. avium 104 infections compound 44 demonstrated an 80% reduction in bacterial viability. Additionally, these compounds did not display toxicity to human macrophages. This finding suggested that these compounds can be developed as potential EPIs against M. avium [209].
Sen et al. synthesized quinazolinone analogs and evaluated quinazolinone analogs for efflux pump modulation activity in M. smegmatis. All quinazolinone analogs (45–50; Figure 7) had a higher MIC value (200 μg/mL) against M. smegmatis [210]. Representative (compounds 45 [100 μg/mL] and 46 [100 μg/mL]) modulated EtBr activity in M. smegmatis by 16-fold. In contrast, compounds 47 (25 μg/mL), 48 (100 μg/mL), and 49 (25 μg/mL) modulated norfloxacin activity in M. smegmatis by a modulation factor of 8. In comparison, compound 50 (12.5 μg/mL) showed the best modulation activity against M. smegmatis and modulated norfloxacin activity by a modulation factor of 16. Furthermore, these compounds exhibited the least moderate toxicity in a hepatocyte cell line (CC-1) based on a cell-based cytotoxicity assay [210]. Mycobactin analogs with diaryl-substituted pyrazoline were synthesized. Under iron-deprived conditions, compounds 51–54 (Figure 7) exhibited activity against M.tb but were inactive against M. smegmatis. Notably, pyrazoline with diaryl acetyl failed to show any activity against M.tb and M. smegmatis. Pyrazoline (55 and 56; Figure 7) demonstrated activity in M. smegmatis and M.tb without any substitution at the N1 position. The mode of action for pyrazoline is MbtA inhibition and efflux pump inhibition in M. smegmatis [211].
The 2-phenyl quinoline derivatives (57 and 58; Figure 7) have NorA efflux pump inhibition activity [212, 213]. Compounds 57 and 58 displayed anti-mycobacterial activity against M. smegmatis mc2 155 and M. avium 104 with the same MIC value (32 μg/mL). All compounds (at a concentration of 1/2 MIC) increased the accumulation of EtBr in M. smegmatis and M. avium. The representative phenyl quinoline (57 and 58; Figure 7) showed potent efflux pump activity with RFF values of 5.91 and 5.84, respectively (P < 0.01). Compound 57 (1/2 MIC) modulates clarithromycin activity and decreases the MIC from 8 μg/mL to < 0.125 μg/mL (> 64-fold reduction) and to 4 μg/mL (2-fold drop) at one-fourth the MIC against the M. smegmatis mc2 155 strain. In contrast, compound 58 decreases the half-MIC of CLA from 8 to 1 μg/mL (8-fold reduction) and alters clarithromycin activty. Furthermore, compound 57 shows potent EtBr efflux inhibitory activity against M. avium 104 (RFF of 1.23; P < 0.01) comparable to verapamil (RFF of 1.77; P < 0.01) and better than thioridazine and chlorpromazine. Compound 57 exhibits superior EtBr efflux inhibitory action compared to verapamil against the M. avium 104CLA3 strain with an RFF index of 2.11 (P < 0.05). Moreover, compound 57 (1/4th MIC) lowers the clarithromycin MIC against M. avium 104 by 8-fold. Simultaneously, MIC reductions of 32- and 16-fold were noted when erythromycin and azithromycin were combined with compound 57. Additionally, compound 57 has synergistic activity with clarithromycin in an ex vivo macrophage model infected with M. avium 104 [213].
Quinoxaline derivatives exhibit a high MIC value (> 256 μg/mL) against M. abscessus and M. simiae [213]. Furthermore, these derivatives were assessed for EPI activity against M. abscessus and M. simiae. Compounds 59–62 (Figure 7) modulate azithromycin activty in M. abscessus by 8-, 8-, 8-, and 2-fold, respectively. In addition, compounds 59 and 62 modulate amikacin activity at a similar 2-fold value. Among these synthesized compounds, compound 59 was the most potent and able to modulate azithromycin and amikacin activity against M. simiae. The SAR study suggested that introducing the side chain and chlorine atoms to quinoxaline did not improve EPI activity of the compound. Instead, quinoxaline had deleterious effects on the efflux pump inhibition activity. The above finding is based on the in vitro studies. Furthermore, in vivo studies are needed to determine efficacy and safety [213]. The compounds with potential efflux pump inhibition activity are summarised in Table 6.
Table 6 Mycobacterium Species Efflux Pump Inhibitors
| Compounds | Modulation of antibiotic activity by efflux pump inhibition | Strain | Additional information | Ref. |
|---|---|---|---|---|
| Piperine | Piperine modulated the activity of RIF. | M.tb H37Rv | Piperine demonstrated synergistic activity with RIF and STR. | [167] |
| Farnesol | Farnesol modulated the activity of EtBr. | M. smegmatis | Farnesol exhibited synergism with RIF. | [170] |
| Reserpine | Reserpine modulated the activity of ciprofloxacin. | M. tuberculosis | Reserpine increases the intracellular concentration of ciprofloxacin in M.tb. and increases susceptibility to antibiotics. | [171, 172] |
| Compounds 4, 5,6, and 7 | Compounds 4, 5,6, and 7 modulated the activity of RIF, EMB, and EtBr. | M. smegmatis mc2 | [173] | |
| Carvacrol | Carvacrol modulated the activity of EtBr. | M.tb H37Rv | Carvacrol causes the accumulation activity of EtBr. | [174] |
| S-limonene | S-limonene modulated the activity of EMB, RIF, and INH. | M. tuberculosis | [176] | |
| R-limonene, myrcene, sabinene, and β-elemene | R-limonene, myrcene, sabinene, and β-elemene modulated the activity of EMB, RIF, and INH. | M. tuberculosis | [176] | |
| Skullcapflavone II, nobiletin, tangeretin, baicalein, and wogonin | Skullcapflavone II, nobiletin, tangeretin, baicalein, and wogonin modulated the activity of EMB RIF. | M. smegmatis mc2 155 | [177] | |
| Carvotacetone | Carvotacetone modulated the activity of EMB and RIF. | M. smegmatis mc2 155 | Carvotacetone significantly inhibits M. aurum ATCC 23366 with an MIC value of 85.9 μM. | [177] |
| Tetrandrine | Tetrandrine modulated the activity of EMB and INH. | M. tuberculosis | Tetrandrine has calcium channel antagonist activity. | [178] |
| Verapamil | Verapamil modulated the activity of EtBr. | M. tuberculosis | In association with RIF, verapamil causes a decrease in the overexpression of the efflux pump gene in M.tb. | [182–184] |
| Verapamil modulated the activity of BDQ. | M. abscessus subsp. bolletii | [186] | ||
| PPIs (omeprazole, pantoprazole, lansoprazole, and lansoprazole sulphide) | PPIs modulated the activity of RIF. | M. tuberculosis | PPIs reduced the RIF efflux. | [192] |
| Thioridazine | Thioridazine modulated the activity of EtBr. | M. avium | Thioridazine blocks the efflux of EtBr. | [183] |
| Timcodar | Timcodar modulated the activity of RIF and INH. | M. tuberculosis | Timcodar showed synergistic activity with RIF, moxifloxacin, and BDQ in a mouse model of M.tb infection. | [200] |
| 2,2,2-trifluoro-1-(1,4,5,6-tetrahydropyridin-3-yl) | It modulated the activity of RIF. | M.tb H37Rv | 2,2,2-trifluoro-1-(1,4,5,6-tetrahydropyridin-30yl) also increases the EtBr accumulation in the FURG-2 (MDR M.tb) strain. | [206] |
| Biochanin A and its derivatives 40 and 41. | These compounds modulated the activity of macrolides (clarithromycin, erythromycin, and azithromycin), fluoroquinolones (ciprofloxacin and ofloxacin), and EtBr. | M.avium | [207] | |
| 3-phenylquinolone derivative (44) | 44 modulated the activity of clarithromycin. | M. avium 104 | [208] | |
| Quinazolinone analogs (45-50) | Quinazolinone analogs modulated the activity of EtBr and norfloxacin | M. smegmatis | [209] | |
| Phenyl quinoline derivatives (54 and 55) | Phenyl quinoline derivatives modulated the activity of clarithromycin. | M. smegmatis mc2 155 | 54 showed synergistic activity with clarithromycin in an ex vivo macrophage model infected with M. avium 104. | [211, 212] |
| Quinoxaline derivatives (59–62) | Quinoxaline derivatives modulated the activity of azithromycin. | M. abscessus | [213] |
Conclusions and future perspective
The inappropriate use of antibiotics for non-microbial conditions in humans, animals, and plants has led to the concerning emergence of MDR strains. In light of this challenge, one promising avenue to address antibiotic resistance is through the inhibition of efflux pumps. The use of EPIs as adjuvants alongside antibiotics has the potential to enhance effectiveness and broaden the antimicrobial spectrum, thereby restoring sensitivity and extending the duration of use. Moreover, incorporating EPIs into treatment protocols may also contribute to decelerating the development of antibiotic resistance. EPIs, like verapamil, reserpine, CCCP, piperine, and phenothiazines, have immense potential to be used as adjuvants because EPIs have shown great promise as adjuvants and exhibit potent activities in macrophages and in vivo mouse models of MDR infection. The robust efficacy in macrophages and in vivo mouse models of MDR Mycobacterium spp. infection underscores the potential to significantly enhance treatment strategies. Moreover, the use of adjuvants has demonstrated the potential to reduce the dose of existing antibiotics that are known to exhibit additional side effects, cytotoxicity, and low efficacy. Furthermore, to use EPIs as an adjuvant to existing antibiotics, in-depth biological studies are needed to develop treating Mycobacterium spp. diseases.
There are several technical challenges associated with the discovery of EPIs. One significant issue is the lack of available crystal structures for Mycobacterium spp. efflux pumps, as well as the absence of co-crystallized proteins with ligands. This limits our ability to conduct a thorough investigation of these proteins and the binding residues. To accelerate the identification of EPIs, it is essential to screen larger chemical libraries through high-throughput screening methods. These screenings should utilize in silico technologies and complementary in vitro and in vivo biological studies to develop effective EPIs against MDR strains.
Acknowledgements
GK thanks Director BITS Pilani for the research facilities. The authors also acknowledge the support of Dr. Nisha Rathor, Professor, Department of Microbiology, Nims Institute of Medical, and Dr. Suresh Kumar, Professor, Department of Humanity of Nims University Rajasthan, for their kind technical review of the manuscript.
Conflicts of Interest
The authors declare no competing financial interests.
Abbreviations
AhpC, alkyl hydroperoxide reductase; ABC, ATP-binding cassette, AMI, amikacin; Araf, D-arabinofuranose; CAP, capreomycin; CCCP, carbonyl cyanide m-chlorophenylhydrazone; COPD, chronic obstructive pulmonary disease; CTAB, cetyltrimethylammonium bromide; DPA, decaprenylphosphoryl-β-D-arabinofuranose; DprE1, decaprenylphosphoryl-β-D-ribose 2′-epimerase 1; DNA, deoxyribonucleic acid; EMB, ethambutol; EPIs, efflux pump inhibitors; ETH, ethionamide; ERDR, ETB resistance-determining region; FIC; fractional inhibitory concentration; FQs, fluoroquinolones; HGT, horizontal gene transfer; ICL, isocitrate lyase; INH, isoniazid; KAN, kanamycin; LAM, lipoarabinomannan; LZD, linezolid; MABA, microplate alamar blue assay; MAC, Mycobacterium avium complex; MATE, multidrug and toxic compound extrusion; MDR-TB, multidrug-resistant TB; MFS; major facilitator superfamily; MmpL; membrane proteins large; MmpS, mycobacterial small membrane protein; mRNA, messenger RNA; M.sm, Mycobacterium smegmatis; MTBC, Mycobacterium tuberculosis complex; M.tb, Mycobacterium tuberculosis; MXF, moxifloxacin; NMP, 1-(1-naphthylmethyl)-piperazine; NTM, non-tuberculous mycobacteria; OM, outer membrane; PMF; proton-motive force; PPIs, proton pump inhibitors; PZA, pyrazinamide; QRDR, quinolone resistance determining region; RFF, relative final fluorescence; RIF, rifampicin; RDDR, RIF-resistance determining region; RNA, ribonucleic acid; RT-qPCR, quantitative real-time PCR; SMR; small multidrug resistance; RND; resistance-nodulation-division; SAR, structure-activity relationship; SNPs, single-nucleotide polymorphisms; STR, streptomycin; TB, tuberculosis; XDR TB, extensively drug-resistant
Graphical abstract
Highlights
- Emerging Threat of Tuberculosis Drug Resistance
A comprehensive overview of the global challenge posed by multidrug-resistant Mycobacterium tuberculosis (MDR-TB) and the rising incidence of drug resistance in non-tuberculous mycobacteria (NTM). - Efflux Pumps: A Key Mechanism Driving Resistance
In-depth exploration of efflux pump systems as critical contributors to intrinsic and acquired drug resistance in mycobacteria, with emphasis on the roles in therapeutic failure and persistence. - Natural Compounds as Promising Efflux Pump Inhibitors
Evaluation of naturally derived molecules with potential to inhibit mycobacterial efflux pumps, highlighting the role in restoring drug susceptibility and enhancing existing tuberculosis treatments.
In brief
This review highlights the critical role of efflux pumps in mediating drug resistance in Mycobacterium tuberculosis and non-tuberculous mycobacteria, and explores the promising potential of natural products as efflux pump inhibitors to restore antibiotic efficacy and combat multidrug resistance.
References
- Pushkaran AC, Biswas R, Mohan CG. Impact of target-based drug design in anti-bacterial drug discovery for the treatment of tuberculosis. In: Mohan C, editor. Structural bioinformatics: applications in preclinical drug discovery process. Challenges and advances in computational chemistry and physics. Springer, Cham.; 2019. pp. 307-46.
- Feehan KT, Gilroy DW. Is resolution the end of inflammation? Trends Mol Med 2019;25(3):198-214. [PMID: 30795972 DOI: 10.1016/j.molmed.2019.01.006]
- Bagcchi S. WHO’s global tuberculosis report 2022. Lancet Microbe 2023;4(1):e20. [PMID: 36521512 DOI: 10.1016/S2666-5247(22)00359-7]
- Momin MAM, Tucker IG, Das SC. High dose dry powder inhalers to overcome the challenges of tuberculosis treatment. Int J Pharm 2018;550(1-2):398-417. [PMID: 30179703 DOI: 10.1016/j.ijpharm.2018.08.061]
- Reuter A, Hughes J, Furin J. Challenges and controversies in childhood tuberculosis. Lancet 2019;394(10202):967-78. [PMID: 31526740 DOI: 10.1016/S0140-6736(19)32045-8]
- Torrey HL, Keren I, Via LE, Lee JS, Lewis K. High persister mutants in Mycobacterium tuberculosis. PLoS One 2016;11(5):e0155127. [PMID: 27176494 DOI: 10.1371/journal.pone.0155127]
- Fedrizzi T, Meehan CJ, Grottola A, Giacobazzi E, Fregni Serpini G, et al. Genomic characterization of nontuberculous mycobacteria. Sci Rep 2017;7(1):45258.
- Falkinham JO. Current epidemiologic trends of the nontuberculous mycobacteria (NTM). Curr Environ Health Rep 2016;3(2):161-7. [PMID: 27020801 DOI: 10.1007/s40572-016-0086-z]
- Falkinham JO. Ecology of nontuberculous mycobacteria. Microorganisms 2021;9(11):1-10. [PMID: 34835388 DOI: 10.3390/microorganisms9112262]
- Porvaznik I, Solovič I, Mokrý J. Non-tuberculous mycobacteria: classification, diagnostics, and therapy. Adv Exp Med Biol 2016;944:19-25. [PMID: 27826888 DOI: 10.1007/5584_2016_45]
- Poonam, Yennamalli RM, Bisht GS, Shrivastava R. Ribosomal maturation factor (RimP) is essential for survival of nontuberculous mycobacteria Mycobacterium fortuitum under in vitro acidic stress conditions. 3 Biotech 2019;9(4):127. [PMID: 30863706 DOI: 10.1007/s13205-019-1659-y]
- Swain A, Gnanasekar P, Prava J, Rajeev AC, Kesarwani P, et al. A comparative genomics approach for shortlisting broad-spectrum drug targets in nontuberculous mycobacteria. Microb Drug Resist 2021;27(2):212-26. [PMID: 32936741 DOI: 10.1089/mdr.2020.0161]
- Huang HL, Lu PL, Lee CH, Chong IW. Treatment of pulmonary disease caused by Mycobacterium kansasii. J Formos Med Assoc 2020;119:S51-7. [PMID: 32505588 DOI: 10.1016/j.jfma.2020.05.018]
- Mogami R, Goldenberg T, de Marca PG, Mello FC, Lopes AJ. Pulmonary infection caused by Mycobacterium kansasii: findings on computed tomography of the chest. Radiol Bras 2016;49(4):209-13. [PMID: 27777472 DOI: 10.1590/0100-3984.2015.0078]
- Coolen-Allou N, Touron T, Belmonte O, Gazaille V, Andre M, et al. Clinical, radiological, and microbiological characteristics of Mycobacterium simiae infection in 97 patients. Antimicrob Agents Chemother 2018;62(7):e00395-18. [PMID: 9760130 DOI: 10.1128/AAC.00395-18]
- Sette CS, Wachholz PA, Masuda PY, da Costa Figueira RBF, de Oliveira Mattar FR, et al. Mycobacterium marinum infection: a case report. J Venom Anim Toxins Incl Trop Dis 2015;21(1):7. [PMID: 25806076 DOI: 10.1186/s40409-015-0008-9]
- Johnson MG, Stout JE. Twenty-eight cases of Mycobacterium marinum infection: retrospective case series and literature review. Infection 2015;43(6):655-62. [PMID: 25869820 DOI: 10.1007/s15010-015-0776-8]
- Chen Y, Jiang J, Jiang H, Chen J, Wang X, et al. Mycobacterium gordonae in patient with facial ulcers, nosebleeds, and positive T-SPOT.TB test, China. Emerg Infect Dis 2017;23(7):1204-6. [PMID: 28628445 DOI: 10.3201/eid2307.162033]
- Utsugi H, Usui Y, Nishihara F, Kanazawa M, Nagata M. Mycobacterium gordonae-induced humidifier lung. BMC Pulm Med 2015;15(1):108. [PMID: 26420433 DOI: 10.1186/s12890-015-0107-y]
- Lotfi H, Sankian M, Meshkat Z, Khalifeh Soltani A, Aryan E. Mycobacterium szulgai pulmonary infection in a vitamin D–deficient patient: a case report. Clin Case Rep 2021;9(3):1146-9. [PMID: 33768798 DOI: 10.1002/ccr3.3692]
- Wilson JW, Jagtiani AC, Wengenack NL. Mycobacterium scrofulaceum disease: experience from a tertiary medical centre and review of the literature. Infect Dis 2019;51(8):602-9. [PMID: 31264491 DOI: 10.1080/23744235.2019.1634281]
- Busatto C, Vianna JS, da Silva LV, Ramis IB, da Silva PEA. Mycobacterium avium: an overview. Tuberculosis 2019;114:127-34. [PMID: 30711152 DOI: 10.1016/j.tube.2018.12.004]
- Gutierrez C, Somoskovi A. Human pathogenic mycobacteria. In: Reference Module in Biomedical Sciences. Third Edition. Elsevier; 2014. pp. 1-15.
- Lee MR, Sheng WH, Hung CC, Yu CJ, Lee LN, et al. Mycobacterium abscessus complex infections in humans. Emerg Infect Dis 2015;21(9):1638-46. [PMID: 26295364 DOI: 10.3201/2109.141634]
- Okamori S, Asakura T, Nishimura T, Tamizu E, Ishii M, et al. Natural history of Mycobacterium fortuitum pulmonary infection presenting with migratory infiltrates: a case report with microbiological analysis. BMC Infect Dis 2018;18(1):1. [PMID: 29291713 DOI: 10.1186/s12879-017-2892-9]
- Johansen MD, Herrmann JL, Kremer L. Non-tuberculous mycobacteria and the rise of Mycobacterium abscessus. Nat Rev Microbiol 2020;18(7):392-407. [PMID: 32086501 DOI: 10.1038/s41579-020-0331-1]
- Gopalaswamy R, Shanmugam S, Mondal R, Subbian S. Of tuberculosis and non-tuberculous mycobacterial infections – a comparative analysis of epidemiology, diagnosis and treatment. J Biomed Sci 2020;27(1):74. [PMID: 32552732 DOI: 10.1186/s12929-020-00667-6]
- Nguta JM, Appiah-Opong R, Nyarko AK, Yeboah-Manu D, Addo PGA. Current perspectives in drug discovery against tuberculosis from natural products. Int J Mycobacteriol 2015;4(3):165-83. [PMID: 27649863 DOI: 10.1016/j.ijmyco.2015.05.004]
- Zumla A, Nahid P, Cole ST. Advances in the development of new tuberculosis drugs and treatment regimens. Nat Rev Drug Discov 2013;12(5):388-404. [PMID: 23629506 DOI: 10.1038/nrd4001]
- Kumar G, Kapoor S. Targeting mycobacterial membranes and membrane proteins: progress and limitations. Bioorg Med Chem 2023;81:117212. [PMID: 36804747 DOI: 10.1016/j.bmc.2023.117212]
- Kumar G, Adhikrao PA. Targeting Mycobacterium tuberculosis iron-scavenging tools: a recent update on siderophores inhibitors. RSC Med Chem 2023;14(10):1885-913. [PMID: 37859726 DOI: 10.1039/d3md00201b]
- Kumar G, Amrutha C. Natural products and their analogues acting against Mycobacterium tuberculosis: aA recent update. Drug Dev Res 2023;84(5):779-804. [PMID: 37086027 DOI: 10.1002/ddr.22063]
- Soni I, De Groote MA, Dasgupta A, Chopra S. Challenges facing the drug discovery pipeline for non-tuberculous mycobacteria. J Med Microbiol 2016;65(1):1-8. [PMID: 26515915 DOI: 10.1099/jmm.0.000198]
- Coolen N, Morand P, Martin C, Hubert D, Kanaan R, et al. Reduced risk of nontuberculous mycobacteria in cystic fibrosis adults receiving long-term azithromycin. J Cyst Fibros 2015;14(5):594-9. [PMID: 25735458 DOI: 10.1016/j.jcf.2015.02.006]
- van Ingen J. Treatment of pulmonary disease caused by non-tuberculous mycobacteria. Lancet Respir Med 2015;3(3):179-80. [PMID: 25773208 DOI: 10.1016/S2213-2600(15)00033-8]
- Palencia A, Li X, Bu W, Choi W, Ding CZ, et al. Discovery of novel oral protein synthesis inhibitors of mycobacterium tuberculosis that target leucyl-tRNA synthetase. Antimicrob Agents Chemother 2016;60(10):6271-80. [PMID: 27503647 DOI: 10.1128/AAC.01339-16]
- Raaijmakers J, Schildkraut JA, Hoefsloot W, van Ingen J. The role of amikacin in the treatment of nontuberculous mycobacterial disease. Expert Opin Pharmacother 2021;22(15):1961-74. [PMID: 34292097 DOI: 10.1080/14656566.2021.1953472]
- Kim TS, Choe JH, Kim YJ, Yang CS, Kwon HJ, et al. Activity of LCB01-0371, a novel oxazolidinone, against Mycobacterium abscessus. Antimicrob Agents Chemother 2017;61(9):e02752. [PMID: 28674049 DOI: 10.1128/AAC.02752-16]
- Brown-Elliott BA, Wallace RJ. In vitro susceptibility testing of tedizolid against nontuberculous mycobacteria. J Clin Microbiol 2017;55(6):1747-54. [PMID: 28330892 DOI: 10.1128/JCM.00274-17]
- Brown-Elliott BA, Wallace RJ. In vitro susceptibility testing of eravacycline against nontuberculous mycobacteria. Antimicrob Agents Chemother 2022;65(3):e0068922. [PMID: 35943269 DOI: 10.1128/aac.00689-22]
- Alffenaar JW, Märtson AG, Heysell SK, Cho JG, Patanwala A, et al. Therapeutic drug monitoring in non-tuberculosis mycobacteria infections. Clin Pharmacokinet 2021;60(6):711-25. [PMID: 33751415 DOI: 10.1007/s40262-021-01000-6]
- Muñoz-Egea MC, Carrasco-Antón N, Esteban J. State-of-the-art treatment strategies for nontuberculous mycobacteria infections. Expert Opin Pharmacother 2020;21(8):969-81. [PMID: 32200657 DOI: 10.1080/14656566.2020.1740205]
- Locher CP, Jones SM, Hanzelka BL, Perola E, Shoen CM, et al. A novel inhibitor of gyrase B is a potent drug candidate for treatment of tuberculosis and nontuberculosis mycobacterial infections. Antimicrob Agents Chemother 2015;59(3):1455-65. [PMID: 25534737 DOI: 10.1128/AAC.04347-14]
- Brown-Elliott BA, Rubio A, Wallace RJ. In vitro susceptibility testing of a novel benzimidazole, SPR719, against nontuberculous mycobacteria. Antimicrob Agents Chemother 2018;62(11):e01503-18. [PMID: 30126964 DOI: 10.1128/AAC.01503-18]
- Stokes SS, Vemula R, Pucci MJ. Advancement of GyrB inhibitors for treatment of infections caused by Mycobacterium tuberculosis and non-tuberculous mycobacteria. ACS Infect Dis 2020;6(6):1323-31. [PMID: 32183511 DOI: 10.1021/acsinfecdis.0c00025]
- Aziz DB, Low JL, Wu ML, Gengenbacher M, Teo JWP, et al. Rifabutin is active against Mycobacterium abscessus complex. Antimicrob Agents Chemother 2017;61(6):1-17. [PMID: 28396540 DOI: 10.1128/AAC.00155-17]
- Kim DH, Kim SY, Huh HJ, Lee NY, Koh WJ, et al. In vitro activity of rifamycin derivatives against nontuberculous mycobacteria, including macrolide- and amikacin-resistant clinical isolates. Antimicrob Agents Chemother 2021;65(5):e02611-20. [PMID: 33685889 DOI: 10.1128/AAC.02611-20]
- Kim DH, Jhun BW, Moon SM, Kim SY, Jeon K, et al. In vitro activity of bedaquiline and delamanid against nontuberculous mycobacteria, including macrolide-resistant clinical isolates. Antimicrob Agents Chemother 2019;63(8):e00665-19. [PMID: 31182533 DOI: 10.1128/AAC.00665-19]
- Pfaeffle HOI, Alameer RM, Marshall MH, Houpt ER, Albon DP, et al. Clofazimine for treatment of multidrug-resistant non-tuberculous mycobacteria. Pulm Pharmacol Ther 2021;70:102058. [PMID: 34293446 DOI: 10.1016/j.pupt.2021.102058]
- Kumar G, Engle K. Natural products acting against S. aureus through membrane and cell wall disruption. Nat Prod Rep 2023;40(10):1608-46. [DOI: 10.1039/D2NP00084A]
- Muñoz-Muñoz L, Aínsa JA, Ramón-García S. Repurposing β-lactams for the treatment of Mycobacterium kansasii infections: an in vitro study. Antibiotics 2023;12(2):335. [PMID: 36830246 DOI: 10.3390/antibiotics12020335]
- Lefebvre AL, Le Moigne V, Bernut A, Veckerlé C, Compain F, et al. Inhibition of the β-lactamase BlaMab by avibactam improves the in vitro and in vivo efficacy of imipenem against Mycobacterium abscessus. Antimicrob Agents Chemother 2017;61(4):e02440. [PMID: 28096155 DOI: 10.1128/AAC.02440-16]
- Negatu DA, González del Río R, Cacho-Izquierdo M, Barros-Aguirre D, Lelievre J, et al. Activity of oral tebipenem-avibactam in a mouse model of Mycobacterium abscessus lung infection. Antimicrob Agents Chemother 2023;67(2):e0145922. [PMID: 36688684 DOI: 10.1128/aac.01459-22]
- Le Run E, Atze H, Arthur M, Mainardi JL. Impact of relebactam-mediated inhibition of Mycobacterium abscessus BlaMab β-lactamase on the in vitro and intracellular efficacy of imipenem. J Antimicrob Chemother 2019;75(2):379-83. [PMID: 31637424 DOI: 10.1093/jac/dkz433]
- Kaushik A, Ammerman NC, Parrish NM, Nuermberger EL. New β-lactamase inhibitors nacubactam and zidebactam improve the in vitro activity of β-lactam antibiotics against Mycobacterium abscessus complex clinical isolates. Antimicrob Agents Chemother 2019;63(9):e00733-19. [PMID: 31209013 DOI: 10.1128/AAC.00733-19]
- Seung KJ, Keshavjee S, Rich ML. Multidrug-resistant tuberculosis and extensively drug-resistant tuberculosis. Cold Spring Harb Perspect Med 2015;5(9):a017863. [PMID: 25918181 DOI: 10.1101/cshperspect.a017863]
- Tran T, Bonham AJ, Chan ED, Honda JR. A paucity of knowledge regarding nontuberculous mycobacterial lipids compared to the tubercle bacillus. Tuberculosis 2019;115:96-107. [PMID: 30948183 DOI: 10.1016/j.tube.2019.02.008]
- World Health Organization. WHO Bacterial Priority Pathogens List, 2024:Bacterial pathogens of public health importance to guide research, development and strategies to prevent and control antimicrobial resistance. 2024.
- Mabhula A, Singh V. Drug-resistance in Mycobacterium tuberculosis: where we stand. Medchemcomm 2019;10(8):1342-60. [PMID: 31534654 DOI: 10.1039/c9md00057g]
- Singh R, Dwivedi SP, Gaharwar US, Meena R, Rajamani P, et al. Recent updates on drug resistance in Mycobacterium tuberculosis. J Appl Microbiol 2020;128(6):1547-67. [PMID: 31595643 DOI: 10.1111/jam.14478]
- Smith T, Wolff KA, Nguyen L. Molecular biology of drug resistance in Mycobacterium tuberculosis. Curr Top Microbiol Immunol 2013;374:53-80. [PMID: 23179675 DOI: 10.1007/82_2012_279]
- Poulton NC, Rock JM. Unraveling the mechanisms of intrinsic drug resistance in Mycobacterium tuberculosis. Front Cell Infect Microbiol 2022;12:997283. [PMID: 36325467 DOI: 10.3389/fcimb.2022.997283]
- Dheda K, Gumbo T, Maartens G, Dooley KE, McNerney R, Murray M, et al. The epidemiology, pathogenesis, transmission, diagnosis, and management of multidrug-resistant, extensively drug-resistant, and incurable tuberculosis. Lancet Respir Med 2017;5(4):291-360. [DOI: 10.1016/S2213-2600(17)30079-6]
- Saxena S, Spaink HP, Forn-Cuní G. Drug resistance in nontuberculous mycobacteria: mechanisms and models. Biology (Basel) 2021;10(2):96. [PMID: 33573039 DOI: 10.3390/biology10020096]
- Sachan RSK, Mistry V, Dholaria M, Rana A, Devgon I, et al. Overcoming Mycobacterium tuberculosis drug resistance: novel medications and repositioning strategies. ACS Omega 2023;8(36):32244-57. [PMID: 37720746 DOI: 10.1021/acsomega.3c02563]
- Song L, Wu X. Development of efflux pump inhibitors in antituberculosis therapy. Int J Antimicrob Agents 2016;47(6):421-9. [PMID: 27211826 DOI: 10.1016/j.ijantimicag.2016.04.007]
- Domínguez J, Boettger EC, Cirillo D, Cobelens F, Eisenach KD, et al. Clinical implications of molecular drug resistance testing for Mycobacterium tuberculosis: a TBNET/RESIST-TB consensus statement. Int J Tuberc Lung Dis 2016;20(1):24-42. [PMID: 26688526 DOI: 10.5588/ijtld.15.0221]
- Dookie N, Rambaran S, Padayatchi N, Mahomed S, Naidoo K. Evolution of drug resistance in Mycobacterium tuberculosis: a review on the molecular determinants of resistance and implications for personalized care. J Antimicrob Chemother 2018;73(5):1138-51. [PMID: 29360989 DOI: 10.1093/jac/dkx506]
- Almeida Da Silva PE, Palomino JC. Molecular basis and mechanisms of drug resistance in Mycobacterium tuberculosis: classical and new drugs. J Antimicrob Chemother 2011;66(7):1417-30. [PMID: 21558086 DOI: 10.1093/jac/dkr173]
- Danilchanka O, Pires D, Anes E, Niederweis M. The Mycobacterium tuberculosis outer membrane channel protein cpnt confers susceptibility to toxic molecules. Antimicrob Agents Chemother 2015;59(4):2328-36. [PMID: 25645841 DOI: 10.1128/AAC.04222-14]
- Tseng ST, Tai CH, Li CR, Lin CF, Shi ZY. The mutations of katG and inhA genes of isoniazid-resistant Mycobacterium tuberculosis isolates in Taiwan. J Microbiol Immunol Infect 2015;48(3):249-55. [PMID: 24184004 DOI: 10.1016/j.jmii.2013.08.018]
- Bakhtiyariniya P, Khosravi AD, Hashemzadeh M, Savari M. Detection and characterization of mutations in genes related to isoniazid resistance in Mycobacterium tuberculosis clinical isolates from Iran. Mol Biol Rep 2022;49(7):6135-43. [PMID: 35366177 DOI: 10.1007/s11033-022-07404-2]
- Viswanathan G, Yadav S, Raghunand TR. Identification of novel loci associated with mycobacterial isoniazid resistance. Tuberculosis 2016;96:21-6. [PMID: 26786650 DOI: 10.1016/j.tube.2015.09.008]
- Yimcharoen M, Saikaew S, Wattananandkul U, Phunpae P, Intorasoot S, et al. The regulation of ManLAM-related gene expression in Mycobacterium tuberculosis with different drug resistance profiles following isoniazid treatment. Infect Drug Resist 2022;15:399-412. [PMID: 35153492 DOI: 10.2147/IDR.S346869]
- Alifano P, Palumbo C, Pasanisi D, Talà A. Rifampicin-resistance, rpoB polymorphism and RNA polymerase genetic engineering. J Biotechnol 2015;202:60-77. [PMID: 25481100 DOI: 10.1016/j.jbiotec.2014.11.024]
- de Knegt GJ, Bruning O, ten Kate MT, de Jong M, van Belkum A, et al. Rifampicin-induced transcriptome response in rifampicin-resistant Mycobacterium tuberculosis. Tuberculosis 2013;93(1):96-101. [PMID: 23182912 DOI: 10.1016/j.tube.2012.10.013]
- Zaw MT, Emran NA, Lin Z. Mutations inside rifampicin-resistance determining region of rpoB gene associated with rifampicin-resistance in Mycobacterium tuberculosis. J Infect Public Health 2018;11(5):605-10. [PMID: 29706316 DOI: 10.1016/j.jiph.2018.04.005]
- Casali N, Nikolayevskyy V, Balabanova Y, Harris SR, Ignatyeva O, et al. Evolution and transmission of drug-resistant tuberculosis in a Russian population. Nat Genet 2014;46(3):279-86. [PMID: 24464101 DOI: 10.1038/ng.2878]
- Merker M, Kohl TA, Barilar I, Andres S, Fowler PW, et al. Phylogenetically informative mutations in genes implicated in antibiotic resistance in Mycobacterium tuberculosis complex. Genome Med 2020;12(1):27. [PMID: 32143680 DOI: 10.1186/s13073-020-00726-5]
- Junaid M, Khan MT, Malik SI, Wei DQ. Insights into the mechanisms of the pyrazinamide resistance of three pyrazinamidase mutants N11K, P69T, and D126N. J Chem Inf Model 2019;59(1):498-508. [PMID: 30481017 DOI: 10.1021/acs.jcim.8b00525]
- Miotto P, Cabibbe AM, Feuerriegel S, Casali N, Drobniewski F, et al. Mycobacterium tuberculosis pyrazinamide resistance determinants: a multicenter study. mBio 2014;5(5):e01819-14. [PMID: 25336456 DOI: 10.1128/mBio.01819-14]
- Sheen P, Requena D, Gushiken E, Gilman RH, Antiparra R, et al. A multiple genome analysis of Mycobacterium tuberculosis reveals specific novel genes and mutations associated with pyrazinamide resistance. BMC Genomics 2017;18(1):769. [PMID: 29020922 DOI: 10.1186/s12864-017-4146-z]
- Zhang L, Zhao Y, Gao Y, Wu L, Gao R, et al. Structures of cell wall arabinosyltransferases with the anti-tuberculosis drug ethambutol. Science 2020;368(6496):1211-9. [PMID: 32327601 DOI: 10.1126/science.aba9102]
- Xu Y, Jia H, Huang H, Sun Z, Zhang Z. Mutations found in embCAB, embR, and ubiA genes of ethambutol-sensitive and -resistant Mycobacterium tuberculosis clinical isolates from China. Biomed Res Int 2015;2015:951706. [PMID: 26417605 DOI: 10.1155/2015/951706]
- He L, Wang X, Cui P, Jin J, Chen J, et al. ubiA (Rv3806c) encoding DPPR synthase involved in cell wall synthesis is associated with ethambutol resistance in Mycobacterium tuberculosis. Tuberculosis 2015;95(2):149-54. [PMID: 25547657 DOI: 10.1016/j.tube.2014.12.002]
- Brossier F, Sougakoff W, Bernard C, Petrou M, Adeyema K, et al. Molecular analysis of the embCAB locus and embR gene involved in ethambutol resistance in clinical isolates of Mycobacterium tuberculosis in France. Antimicrob Agents Chemother 2015;59(8):4800-8. [PMID: 26033726 DOI: 10.1128/AAC.00150-15]
- Zhao LL, Sun Q, Liu HC, Wu XC, Xiao TY, et al. Analysis of embCAB mutations associated with ethambutol resistance in multidrug-resistant Mycobacterium tuberculosis isolates from China. Antimicrob Agents Chemother 2015;59(4):2045-50. [PMID: 25605360 DOI: 10.1128/AAC.04933-14]
- Shippy DC, Fadl AA. RNA modification enzymes encoded by the gid operon: Implications in biology and virulence of bacteria. Microb Pathog 2015;89:100-7. [DOI: 10.1016/j.micpath.2015.09.008]
- Hlaing YM, Tongtawe P, Tapchaisri P, Thanongsaksrikul J, Thawornwan U, et al. Mutations in streptomycin resistance genes and their relationship to streptomycin resistance and lineage of Mycobacterium tuberculosis thai isolates. Tuberc Respir Dis (Seoul) 2017;80(2):159. [PMID: 28416956 DOI: 10.4046/trd.2017.80.2.159]
- Reeves AZ, Campbell PJ, Sultana R, Malik S, Murray M, et al. Aminoglycoside cross-resistance in Mycobacterium tuberculosis due to mutations in the 5′ untranslated region of whiB7. Antimicrob Agents Chemother 2013;57(4):1857-65. [PMID: 23380727 DOI: 10.1128/AAC.02191-12]
- Batson S, De Chiara C, Majce V, Lloyd AJ, Gobec S, et al. Inhibition of D-Ala:D-Ala ligase through a phosphorylated form of the antibiotic D-cycloserine. Nat Commun 2017;8(1):1939. [DOI: 10.1038/s41467-017-02118-7]
- Chen J, Zhang S, Cui P, Shi W, Zhang W, et al. Identification of novel mutations associated with cycloserine resistance in Mycobacterium tuberculosis. J Antimicrob Chemother 2017;72(12):3272-6. [PMID: 28961957 DOI: 10.1093/jac/dkx316]
- Malik S, Willby M, Sikes D, Tsodikov OV, Posey JE. New insights into fluoroquinolone resistance in Mycobacterium tuberculosis: functional genetic analysis of gyrA and gyrB mutations. PLoS One 2012;7(6):e39754. [PMID: 22761889 DOI: 10.1371/journal.pone.0039754]
- Eilertson B, Maruri F, Blackman A, Guo Y, Herrera M, et al. A novel resistance mutation in eccC5 of the ESX-5 secretion system confers ofloxacin resistance in Mycobacterium tuberculosis. J Antimicrob Chemother 2016;71(9):2419-27. [PMID: 27261264 DOI: 10.1093/jac/dkw168]
- Aragaw WW, Cotroneo N, Stokes S, Pucci M, Critchley I, et al. In vitro resistance against DNA gyrase inhibitor SPR719 in Mycobacterium avium and Mycobacterium abscessus. Microbiol Spectr 2022;10(1):e0132121. [PMID: 35019671 DOI: 10.1128/spectrum.01321-21]
- Durcik M, Tomašič T, Zidar N, Zega A, Kikelj D, et al. ATP-competitive DNA gyrase and topoisomerase IV inhibitors as antibacterial agents. Expert Opin Ther Pat 2019;29(3):171-80. [PMID: 30686070 DOI: 10.1080/13543776.2019.1575362]
- Preiss L, Langer JD, Yildiz Ö, Eckhardt-Strelau L, Guillemont JEG, et al. Structure of the mycobacterial ATP synthase Fo rotor ring in complex with the anti-TB drug bedaquiline. Sci Adv 2015;1(4):e1500106. [PMID: 26601184 DOI: 10.1126/sciadv.1500106]
- Andries K, Villellas C, Coeck N, Thys K, Gevers T, et al. Acquired resistance of Mycobacterium tuberculosis to bedaquiline. PLoS One 2014;9(7):e102135. [PMID: 25010492 DOI: 10.1371/journal.pone.0102135]
- Almeida D, Ioerger T, Tyagi S, Li SY, Mdluli K, et al. Mutations in pepQ confer low-level resistance to bedaquiline and clofazimine in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2016;60(8):4590-9. [PMID: 27185800 DOI: 10.1128/AAC.00753-16]
- Gler MT, Skripconoka V, Sanchez-Garavito E, Xiao H, Cabrera-Rivero JL, et al. Delamanid for multidrug-resistant pulmonary tuberculosis. N Engl J Med 2012;2151-60. [PMID: 22670901 DOI: 10.1056/NEJMoa1112433]
- Ryan NJ, Lo JH. Delamanid: first global approval. Drugs 2014;74(9):1041-5. [PMID: 24923253 DOI: 10.1007/s40265-014-0241-5]
- Blair HA, Scott LJ. Delamanid: a review of its use in patients with multidrug-resistant tuberculosis. Drugs 2015;75(1):91-100. [PMID: 25404020 DOI: 10.1007/s40265-014-0331-4]
- Nguyen TVA, Anthony RM, Cao TTH, Bañuls AL, Nguyen VAT, et al. Delamanid resistance: update and clinical management. Clin Infect Dis 2020;71(12):3252-9. [PMID: 32521000 DOI: 10.1093/cid/ciaa755]
- Fujiwara M, Kawasaki M, Hariguchi N, Liu Y, Matsumoto M. Mechanisms of resistance to delamanid, a drug for Mycobacterium tuberculosis. Tuberculosis 2018;108:186-94. [PMID: 29523322 DOI: 10.1016/j.tube.2017.12.006]
- Liu Y, Shi J, Li L, Wu T, Chu P, et al. Spontaneous mutational patterns and novel mutations for delamanid resistance in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2022;66(12):e0053122. [PMID: 36448833 DOI: 10.1128/aac.00531-22]
- Rifat D, Li SY, Ioerger T, Shah K, Lanoix JP, et al. Mutations in fbiD (Rv2983) as a novel determinant of resistance to pretomanid and delamanid in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2021;65(1):e01948. [PMID: 33077652 DOI: 10.1128/AAC.01948-20]
- Batt SM, Jabeen T, Bhowruth V, Quill L, Lund PA, et al. Structural basis of inhibition of Mycobacterium tuberculosis DprE1 by benzothiazinone inhibitors. Proc Natl Acad Sci U S A 2012;109(28):11354-9. [PMID: 22733761 DOI: 10.1073/pnas.1205735109]
- Brecik M, Centárová I, Mukherjee R, Kolly GS, Huszár S, et al. DprE1 is a vulnerable tuberculosis drug target due to its cell wall localization. ACS Chem Biol 2015;10(7):1631-6. [PMID: 25906160 DOI: 10.1021/acschembio.5b00237]
- Poulton NC, Azadian ZA, DeJesus MA, Rock JM. Mutations in rv0678 confer low-level resistance to benzothiazinone DprE1 inhibitors in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2022;66(9):e0090422. [PMID: 35920665 DOI: 10.1128/aac.00904-22]
- Ye M, Xu L, Zou Y, Li B, Guo Q, et al. Molecular analysis of linezolid-resistant clinical isolates of Mycobacterium abscessus. Antimicrob Agents Chemother 2019;63(2):e01842-18. [PMID: 30478161 DOI: 10.1128/AAC.01842-18]
- Luthra S, Rominski A, Sander P. The role of antibiotic-target-modifying and antibiotic-modifying enzymes in Mycobacterium abscessus drug resistance. Front Microbiol 2018;9:2179. [PMID: 30258428 DOI: 10.3389/fmicb.2018.02179]
- Fisher JF, Mobashery S. β-Lactam resistance mechanisms: gram-positive bacteria and Mycobacterium tuberculosis. Cold Spring Harb Perspect Med 2016;6(5):a025221. [PMID: 27091943 DOI: 10.1101/cshperspect.a025221]
- Witek MA, Kuiper EG, Minten E, Crispell EK, Conn GL. A novel motif for S-adenosyl-l-methionine binding by the ribosomal RNA methyltransferase TlyA from Mycobacterium tuberculosis. J Biol Chem 2017;292(5):1977-87. [PMID: 28031456 DOI: 10.1074/jbc.M116.752659]
- Laughlin ZT, Nandi S, Dey D, Zelinskaya N, Witek MA, et al. 50S subunit recognition and modification by the Mycobacterium tuberculosis ribosomal RNA methyltransferase TlyA. Proc Natl Acad Sci 2022;119(14):e212035119. [PMID: 35357969 DOI: 10.1073/pnas.2120352119]
- Richard M, Gutiérrez AV, Kremer L. Dissecting erm(41)-mediated macrolide-inducible resistance in Mycobacterium abscessus. Antimicrob Agents Chemother 2020;64(2):e01879-19. [PMID: 31791943 DOI: 10.1128/AAC.01879-19]
- Griffith DE, Daley CL. Treatment of Mycobacterium abscessus pulmonary disease. Chest 2022;161(1):64-75. [PMID: 34314673 DOI: 10.1016/j.chest.2021.07.035]
- Rominski A, Roditscheff A, Selchow P, Böttger EC, Sander P. Intrinsic rifamycin resistance of Mycobacterium abscessus is mediated by ADP-ribosyltransferase MAB_0591. J Antimicrob Chemother 2017;72(2):376-84. [PMID: 27999011 DOI: 10.1093/jac/dkw466]
- Chuang YM, Bandyopadhyay N, Rifat D, Rubin H, Bader JS, et al. Deficiency of the novel exopolyphosphatase Rv1026/PPX2 leads to metabolic downshift and altered cell wall permeability in Mycobacterium tuberculosis. mBio 2015;6(2):e02428. [PMID: 25784702 DOI: 10.1128/mBio.02428-14]
- Hajiagha MN, Kafil HS. Efflux pumps and microbial biofilm formation. Infect Genet Evol 2023;115:105459. [PMID: 37271271 DOI: 10.1016/j.meegid.2023.105459]
- Fouladi MD, Besharati S, Farnia P, Khosravi A. A concise review of the effect of efflux pump on biofilm intensity in bacteria with a special view to Mycobacterium. J Prev Diagn Treat Strate Med 2024;3(1):1-5. [DOI: 10.4103/jpdtsm.jpdtsm_119_23]
- Li P, Gu Y, Li J, Xie L, Li X, et al. Mycobacterium tuberculosis major facilitator superfamily transporters. J Membr Biol 2017;250(6):573-85. [PMID: 28852815 DOI: 10.1007/s00232-017-9982-x]
- Kanji A, Hasan R, Hasan Z. Efflux pump as alternate mechanism for drug resistance in Mycobacterium tuberculosis. Indian J Tuberc 2019;66(1):20-5. [PMID: 30797276 DOI: 10.1016/j.ijtb.2018.07.008]
- Balganesh M, Dinesh N, Sharma S, Kuruppath S, Nair AV, et al. Efflux pumps of Mycobacterium tuberculosis play a significant role in antituberculosis activity of potential drug candidates. Antimicrob Agents Chemother 2012;56(5):2643-51. [PMID: 22314527 DOI: 10.1128/AAC.06003-11]
- Papkou A, Hedge J, Kapel N, Young B, MacLean RC. Efflux pump activity potentiates the evolution of antibiotic resistance across S. aureus isolates. Nat Commun 2020;11(1):3970. [PMID: 32769975 DOI: 10.1038/s41467-020-17735-y]
- Briffotaux J, Huang W, Wang X, Gicquel B. MmpS5/MmpL5 as an efflux pump in Mycobacterium species. Tuberculosis 2017;107:13-9. [PMID: 29050760 DOI: 10.1016/j.tube.2017.08.001]
- Kumar G, Kiran Tudu A. Tackling multidrug-resistant Staphylococcus aureus by natural products and their analogues acting as NorA efflux pump inhibitors. Bioorg Med Chem 2023;80:117187. [PMID: 36731248 DOI: 10.1016/j.bmc.2023.117187]
- Kanji A, Hasan R, Ali A, Zaver A, Zhang Y, et al. Single nucleotide polymorphisms in efflux pumps genes in extensively drug resistant Mycobacterium tuberculosis isolates from Pakistan. Tuberculosis 2017;107:20-30. [PMID: 29050768 DOI: 10.1016/j.tube.2017.07.012]
- Coll F, Phelan J, Hill-Cawthorne GA, Nair MB, Mallard K, et al. Genome-wide analysis of multi- and extensively drug-resistant Mycobacterium tuberculosis. Nat Genet 2018;50(2):307-16. [PMID: 29358649 DOI: 10.1038/s41588-017-0029-0]
- Duan W, Li X, Ge Y, Yu Z, Li P, et al. Mycobacterium tuberculosis Rv1473 is a novel macrolides ABC efflux pump regulated by WhiB7. Future Microbiol 2019;14(1):47-59. [PMID: 30539658 DOI: 10.2217/fmb-2018-0207]
- De Siena B, Campolattano N, D’Abrosca G, Russo L, Cantillon D, et al. Characterization of the mycobacterial MSMEG-3762/63 efflux pump in Mycobacterium smegmatis drug efflux. Front Microbiol 2020;11:575828. [PMID: 33343518 DOI: 10.3389/fmicb.2020.575828]
- Wang K, Pei H, Huang B, Zhu X, Zhang J, et al. The expression of ABC efflux pump, Rv1217c-Rv1218c, and its association with multidrug resistance of Mycobacterium tuberculosis in China. Curr Microbiol 2013;66(3):222-6. [PMID: 23143285 DOI: 10.1007/s00284-012-0215-3]
- Ma F, Zhou H, Yang Z, Wang C, An Y, et al. Gene expression profile analysis and target gene discovery of Mycobacterium tuberculosis biofilm. Appl Microbiol Biotechnol 2021;105(12):5123-34. [PMID: 34125278 DOI: 10.1007/s00253-021-11361-4]
- Liu J, Shi W, Zhang S, Hao X, Maslov DA, et al. Mutations in efflux pump Rv1258c (Tap) cause resistance to pyrazinamide, isoniazid, and streptomycin in M. tuberculosis. Front Microbiol 2019;10:216. [PMID: 30837962 DOI: 10.3389/fmicb.2019.00216]
- Cloete R, Kapp E, Joubert J, Christoffels A, Malan SF. Correction: Molecular modelling and simulation studies of the Mycobacterium tuberculosis multidrug efflux pump protein Rv1258c. PLoS One 2018;13(11):e0207605. [PMID: 30475855 DOI: 10.1371/journal.pone.0209717]
- Johnson EO, Office E, Kawate T, Orzechowski M, Hung DT. Large-scale chemical-genetic strategy enables the design of antimicrobial combination chemotherapy in mycobacteria. ACS Infect Dis 2020;6(1):56-63. [PMID: 31721551 DOI: 10.1021/acsinfecdis.9b00373]
- Johnson EO, LaVerriere E, Office E, Stanley M, Meyer E, et al. Large-scale chemical–genetics yields new M. tuberculosis inhibitor classes. Nature 2019;571(7763):72-8. [PMID: 31217586 DOI: 10.1038/s41586-019-1315-z]
- Wang S, Wang K, Song K, Lai ZW, Li P, Li D, et al. Structures of the Mycobacterium tuberculosis efflux pump EfpA reveal the mechanisms of transport and inhibition. Nat Commun 2024;15(1):7710. [PMID: 39231991 DOI: 10.1038/s41467-024-51948-9]
- Umar F, Hatta M, Husain DR, Natzir R, Dwiyanti R, et al. The effect of anti-tuberculosis drugs therapy on mRNA efflux pump gene expression of Rv1250 in Mycobacterium tuberculosis collected from tuberculosis patients. New Microbes New Infect 2019;32:100609. [PMID: 33014381 DOI: 10.1016/j.nmni.2019.100609]
- Li X, Li P, Ruan C, Xie LX, Gu Y, et al. Mycobacterium tuberculosis Rv0191 is an efflux pump of major facilitator superfamily transporter regulated by Rv1353c. Arch Biochem Biophys 2019;667(May):59-66. [PMID: 31054279 DOI: 10.1016/j.abb.2019.04.010]
- Thakur V, Uniyal A, Tiwari V. A comprehensive review on pharmacology of efflux pumps and their inhibitors in antibiotic resistance. Eur J Pharmacol 2021;903:174151. [PMID: 33964293 DOI: 10.1016/j.ejphar.2021.174151]
- Rai D, Mehra S. The mycobacterial efflux pump EfpA can induce high drug tolerance to many antituberculosis drugs, including moxifloxacin, in Mycobacterium smegmatis. Antimicrob Agents Chemother 2021;65(11):e0026221. [PMID: 34424047 DOI: 10.1128/AAC.00262-21]
- Zhang Z, Wang R, Xie J. Mycobacterium smegmatis MSMEG_3705 encodes a selective major facilitator superfamily efflux pump with multiple roles. Curr Microbiol 2015;70(6):801-9. [PMID: 25697715 DOI: 10.1007/s00284-015-0783-0]
- Mörs K, Hellmich UA, Basting D, Marchand P, Wurm JP, Haase W, et al. A lipid-dependent link between activity and oligomerization state of the M. tuberculosis SMR protein TBsmr. Biochim Biophys Acta 2013;1828(2):561-7. [PMID: 23103507 DOI: 10.1016/j.bbamem.2012.10.020]
- Machado D, Lecorche E, Mougari F, Cambau E, Viveiros M. Insights on Mycobacterium leprae efflux pumps and their implications in drug resistance and virulence. Front Microbiol 2018;9:3072. [PMID: 30619157 DOI: 10.3389/fmicb.2018.03072]
- Rodrigues L, Villellas C, Bailo R, Viveiros M, Aínsa JA. Role of the mmr efflux pump in drug resistance in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2013;57(2):751-7. [PMID: 23165464 DOI: 10.1128/AAC.01482-12]
- Unissa AN, Dusthackeer VNA, Kumar MP, Nagarajan P, Sukumar S, et al. Variants of katG, inhA and nat genes are not associated with mutations in efflux pump genes (mmpL3 and mmpL7) in isoniazid-resistant clinical isolates of Mycobacterium tuberculosis from India. Tuberculosis 2017;107:144-8. [PMID: 29050763 DOI: 10.1016/j.tube.2017.07.014]
- Gutiérrez AV, Richard M, Roquet-Banères F, Viljoen A, Kremer L. The TetR family transcription factor MAB_2299c regulates the expression of two distinct MmpS-MmpL efflux pumps involved in cross-resistance to clofazimine and bedaquiline in Mycobacterium abscessus. Antimicrob Agents Chemother 2019;63(10):e01000-19. [PMID: 31332077 DOI: 10.1128/AAC.01000-19]
- Yamamoto K, Nakata N, Mukai T, Kawagishi I, Ato M. Coexpression of MmpS5 and MmpL5 contributes to both efflux transporter MmpL5 trimerization and drug resistance in Mycobacterium tuberculosis. mSphere 2021;6(1):e00518. [PMID: 33408221 DOI: 10.1128/mSphere.00518-20]
- Farnia P, Besharati S, Farina P, Ayoubi S, Marjani M, et al. The role of efflux pumps transporter in multi-drug resistant tuberculosis: mycobacterial memberane protein(MmpL5). Int J Mycobacteriol 2024;13(1):7-14. [PMID: 38771273 DOI: 10.4103/ijmy.ijmy_37_24]
- Hartkoorn RC, Uplekar S, Cole ST. Cross-resistance between clofazimine and bedaquiline through upregulation of mmpl5 in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2014;58(5):2979-81. [PMID: 24590481 DOI: 10.1128/AAC.00037-14]
- Li M, Nyantakyi SA, Go ML, Dick T. Resistance against membrane-inserting MmpL3 inhibitor through upregulation of MmpL5 in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2020;64(12):e01100-20. [PMID: 32958714 DOI: 10.1128/AAC.01100-20]
- Shur KV, Frolova SG, Akimova NI, Danilenko VN, Maslov DA. A test system for in vitro screening antimycobacterial drug candidates for MmpS5-MmpL5 mediated resistance. Russ J Genet 2021;57(1):114-6. [DOI: 10.1134/S1022795421010154]
- Guo Q, Chen J, Zhang S, Zou Y, Zhang Y, et al. Efflux pumps contribute to intrinsic clarithromycin resistance in clinical, Mycobacterium abscessus isolates. Infect Drug Resist 2020;13:447-54. [PMID: 32104016 DOI: 10.2147/IDR.S239850]
- Guo Q, Zhang Y, Fan J, Zhang H, Zhang Z, et al. MAB_2355c confers macrolide resistance in Mycobacterium abscessus by ribosome protection. Antimicrob Agents Chemother 2021;65(8):e0033021. [PMID: 34097497 DOI: 10.1128/AAC.00330-21]
- Vianna JS, Machado D, Ramis IB, Silva FP, Bierhals DV, et al. The contribution of efflux pumps in Mycobacterium abscessus complex resistance to clarithromycin. Antibiotics 2019;8(3):153. [PMID: 31540480 DOI: 10.3390/antibiotics8030153]
- Rodrigues L, Machado D, Couto I, Amaral L, Viveiros M. Contribution of efflux activity to isoniazid resistance in the Mycobacterium tuberculosis complex. Infect Genet Evol 2012;12(4):695-700. [PMID: 21871582 DOI: 10.1016/j.meegid.2011.08.009]
- Gupta AK, Reddy VP, Lavania M, Chauhan DS, Venkatesan K, et al. jefA (Rv2459), a drug efflux gene in Mycobacterium tuberculosis confers resistance to isoniazid & ethambutol. Indian J Med Res 2010;132(8):176-88. [PMID: 20716818]
- Zhang Y, Zhang J, Cui P, Zhang Y, Zhang W. Identification of novel efflux proteins Rv0191, Rv3756c, Rv3008, and Rv1667c involved in pyrazinamide resistance in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2017;61(8):e00940-17. [PMID: 28584158 DOI: 10.1128/AAC.00940-17]
- Zimic M, Fuentes P, Gilman RH, Gutiérrez AH, Kirwan D, et al. Pyrazinoic acid efflux rate in Mycobacterium tuberculosis is a better proxy of pyrazinamide resistance. Tuberculosis 2012;92(1):84-91. [PMID: 22004792 DOI: 10.1016/j.tube.2011.09.002]
- Caleffi-Ferracioli KR, Amaral RCR, Demitto FO, Maltempe FG, Canezin PH, et al. Morphological changes and differentially expressed efflux pump genes in Mycobacterium tuberculosis exposed to a rifampicin and verapamil combination. Tuberculosis 2016;97:65-72. [PMID: 26980498 DOI: 10.1016/j.tube.2015.12.010]
- Narang A, Garima K, Porwal S, Bhandekar A, Shrivastava K, et al. Potential impact of efflux pump genes in mediating rifampicin resistance in clinical isolates of Mycobacterium tuberculosis from India. PLoS One 2019;14(9):e0223163. [PMID: 31557231 DOI: 10.1371/journal.pone.0223163]
- Canezin PH, Caleffi-Ferracioli KR, Scodro RBL, Siqueira VLD, Pavan FR, et al. Intramacrophage Mycobacterium tuberculosis efflux pump gene regulation after rifampicin and verapamil exposure. J Antimicrob Chemother 2018;73(7):1770-6. [PMID: 29579201 DOI: 10.1093/jac/dky091]
- Li G, Zhang J, Guo Q, Wei J, Jiang Y, et al. Study of efflux pump gene expression in rifampicin-monoresistant Mycobacterium tuberculosis clinical isolates. J Antibiot (Tokyo) 2015;68(7):431-5. [PMID: 25690361 DOI: 10.1038/ja.2015.9]
- Machado D, Coelho TS, Perdigão J, Pereira C, Couto I, et al. Interplay between mutations and efflux in drug resistant clinical isolates of Mycobacterium tuberculosis. Front Microbiol 2017;8:711. [PMID: 28496433 DOI: 10.3389/fmicb.2017.00711]
- Pule CM, Sampson SL, Warren RM, Black PA, van Helden PD, et al. Efflux pump inhibitors: targeting mycobacterial efflux systems to enhance TB therapy. J Antimicrob Chemother 2016;71(1):17-26. [PMID: 26472768 DOI: 10.1093/jac/dkv316]
- Sharma S, Kalia NP, Suden P, Chauhan PS, Kumar M, et al. Protective efficacy of piperine against Mycobacterium tuberculosis. Tuberculosis 2014;94(4):389-96. [PMID: 24880706 DOI: 10.1016/j.tube.2014.04.007]
- Sharma S, Kumar M, Sharma S, Nargotra A, Koul S, et al. Piperine as an inhibitor of Rv1258c, a putative multidrug efflux pump of Mycobacterium tuberculosis. J Antimicrob Chemother 2010;65(8):1694-701. [PMID: 20525733 DOI: 10.1093/jac/dkq186]
- Hegeto LA, Caleffi-Ferracioli KR, Nakamura-Vasconcelos SS, Almeida AL, Baldin VP, et al. In vitro combinatory activity of piperine and anti-tuberculosis drugs in Mycobacterium tuberculosis. Tuberculosis 2018;111:35-40. [PMID: 30029912 DOI: 10.1016/j.tube.2018.05.006]
- Jin J, Zhang JY, Guo N, Sheng H, Li L, et al. Farnesol, a potential efflux pump inhibitor in Mycobacterium smegmatis. Molecules 2010;15(11):7750-62. [PMID: 21042264 DOI: 10.3390/molecules15117750]
- Viveiros M, Martins M, Rodrigues L, Machado D, Couto I, et al. Inhibitors of mycobacterial efflux pumps as potential boosters for anti-tubercular drugs. Expert Rev Anti Infect Ther 2012;10(9):983-98. [PMID: 23106274 DOI: 10.1586/eri.12.89]
- Huang TS, Kunin CM, Wang HM, Yan BS, Huang SP, et al. Inhibition of the Mycobacterium tuberculosis reserpine-sensitive efflux pump augments intracellular concentrations of ciprofloxacin and enhances susceptibility of some clinical isolates. J Formos Med Assoc 2013;112(12):789-94. [PMID: 24331109 DOI: 10.1016/j.jfma.2012.03.009]
- Gröblacher B, Kunert O, Bucar F. Compounds of Alpinia katsumadai as potential efflux inhibitors in Mycobacterium smegmatis. Bioorg Med Chem 2012;20(8):2701-6. [PMID: 22459211 DOI: 10.1016/j.bmc.2012.02.039]
- Nakamura de Vasconcelos SS, Caleffi-Ferracioli KR, Hegeto LA, Baldin VP, Nakamura CV, et al. Carvacrol activity & morphological changes in Mycobacterium tuberculosis. Future Microbiol 2018;13(8):877-88. [PMID: 29877104 DOI: 10.2217/fmb-2017-0232]
- Miladi H, Zmantar T, Chaabouni Y, Fedhila K, Bakhrouf A, et al. Antibacterial and efflux pump inhibitors of thymol and carvacrol against food-borne pathogens. Microb Pathog 2016;99:95-100. [PMID: 27521228 DOI: 10.1016/j.micpath.2016.08.008]
- Sieniawska E, Swatko-Ossor M, Sawicki R, Skalicka-Woźniak K, Ginalska G. Natural terpenes influence the activity of antibiotics against isolated Mycobacterium tuberculosis. Med Princ Pract 2017;26(2):108-12. [PMID: 27883995 DOI: 10.1159/000454680]
- Solnier J, Martin L, Bhakta S, Bucar F. Flavonoids as novel efflux pump inhibitors and antimicrobials against both environmental and pathogenic intracellular mycobacterial species. Molecules 2020;25(3):734. [PMID: 32046221 DOI: 10.3390/molecules25030734]
- Tran HT, Solnier J, Pferschy-Wenzig EM, Kunert O, Martin L, et al. Antimicrobial and efflux pump inhibitory activity of carvotacetones from Sphaeranthus africanus against mycobacteria. Antibiotics 2020;9(7):390. [PMID: 32650510 DOI: 10.3390/antibiotics9070390]
- Bhagya N, Chandrashekar KR. Tetrandrine – a molecule of wide bioactivity. Phytochemistry 2016;125:5-13. [PMID: 26899361 DOI: 10.1016/j.phytochem.2016.02.005]
- Zhang Z, Yan J, Xu K, Ji Z, Li L. Tetrandrine reverses drug resistance in isoniazid and ethambutol dual drug-resistant Mycobacterium tuberculosis clinical isolates. BMC Infect Dis 2015;15(1):153. [PMID: 25887373 DOI: 10.1186/s12879-015-0905-0]
- Plášek J, Babuka D, Hoefer M. H+ translocation by weak acid uncouplers is independent of H+ electrochemical gradient. J Bioenerg Biomembr 2017;49(5):391-7. [PMID: 28900787 DOI: 10.1007/s10863-017-9724-x]
- Black PA, Warren RM, Louw GE, Van Helden PD, Victor TC, et al. Energy metabolism and drug efflux in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2014;58(5):2491-503. [PMID: 24614376 DOI: 10.1128/AAC.02293-13]
- Umar F, Hatta M, Husain DR, Bahar B, Bukhari A, et al. Verapamil as an efflux inhibitor against drug resistant Mycobacterium tuberculosis: a review. Sys Rev Pharm 2019;10(1):S43-8. [DOI: 10.5530/srp.2019.1s.22]
- Remm S, Earp JC, Dick T, Dartois V, Seeger MA. Critical discussion on drug efflux in Mycobacterium tuberculosis. FEMS Microbiol Rev 2022;46(1):fuab050. [PMID: 34637511 DOI: 10.1093/femsre/fuab050]
- Li G, Zhang J, Li C, Guo Q, Jiang Y, et al. Antimycobacterial activity of five efflux pump inhibitors against Mycobacterium tuberculosis clinical isolates. J Antibiot (Tokyo) 2016;69(3):173-5. [PMID: 26464012 DOI: 10.1038/ja.2015.101]
- Gupta S, Cohen KA, Winglee K, Maiga M, Diarra B, et al. Efflux inhibition with verapamil potentiates bedaquiline in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2014;58(1):574-6. [PMID: 24126586 DOI: 10.1128/AAC.01462-13]
- Xu J, Tasneen R, Peloquin CA, Almeida DV, Li SY, et al. Verapamil increases the bioavailability and efficacy of bedaquiline but not clofazimine in a murine model of tuberculosis. Antimicrob Agents Chemother 2018;62(1):e01692-17. [PMID: 29038265 DOI: 10.1128/AAC.01692-17]
- Jang J, Kim R, Woo M, Jeong J, Park DE, et al. Efflux attenuates the antibacterial activity of Q203 in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2017;61(7):e02637-16. [PMID: 28416541 DOI: 10.1128/AAC.02637-16]
- Gupta S, Tyagi S, Bishaia WR. Verapamil increases the bactericidal activity of bedaquiline against Mycobacterium tuberculosis in a mouse model. Antimicrob Agents Chemother 2015;59(1):673-6. [PMID: 25331694 DOI: 10.1128/AAC.04019-14]
- Caleffi-Ferracioli KR, Cardoso RF, De Souza JVP, Murase LS, Canezin PH, et al. Modulatory effects of verapamil in rifampicin activity against Mycobacterium tuberculosis. Future Microbiol 2019;14(3):185-94. [PMID: 30648892 DOI: 10.2217/fmb-2018-0277]
- Singh K, Kumar M, Pavadai E, Naran K, Warner DF, et al. Synthesis of new verapamil analogues and their evaluation in combination with rifampicin against Mycobacterium tuberculosis and molecular docking studies in the binding site of efflux protein Rv1258c. Bioorg Med Chem Lett 2014;24(14):2985-90. [PMID: 24894561 DOI: 10.1016/j.bmcl.2014.05.022]
- Adams KN, Szumowski JD, Ramakrishnan L. Verapamil, and its metabolite norverapamil, inhibit macrophage-induced, bacterial efflux pump-mediated tolerance to multiple anti-tubercular drugs. J Infect Dis 2014;210(3):456-66. [PMID: 24532601 DOI: 10.1093/infdis/jiu095]
- Lake MA, Adams KN, Nie F, Fowler E, Verma AK, et al. The human proton pump inhibitors inhibit Mycobacterium tuberculosis rifampicin efflux and macrophage-induced rifampicin tolerance. Proc Natl Acad Sci USA 2023;120(7):e2215512120. [PMID: 36763530 DOI: 10.1073/pnas.2215512120]
- Garrison AT, Abouelhassan Y, Kallifidas D, Bai F, Ukhanova M, et al. Halogenated phenazines that potently eradicate biofilms, MRSA persister cells in non-biofilm cultures, and Mycobacterium tuberculosis. Angew Chem Int Ed Engl 2015;127(49):15032-6. [PMID: 26480852 DOI: 10.1002/anie.201508155]
- Kristiansen JE, Dastidar SG, Palchoudhuri S, Roy DS, Das S, et al. Phenothiazines as a solution for multidrug resistant tuberculosis: From the origin to present. Int Microbiol 2015;18(1):1-12. [PMID: 26415662 DOI: 10.2436/20.1501.01.229]
- Kumar M, Singh K, Naran K, Hamzabegovic F, Hoft DF, et al. Design, synthesis, and evaluation of novel hybrid efflux pump inhibitors for use against Mycobacterium tuberculosis. ACS Infect Dis 2016;2:714-25. [PMID: 27737555 DOI: 10.1021/acsinfecdis.6b00111]
- Vibe CB, Fenaroli F, Pires D, Wilson SR, Bogoeva V, et al. Thioridazine in PLGA nanoparticles reduces toxicity and improves rifampicin therapy against mycobacterial infection in zebrafish. Nanotoxicology 2016;10(6):680-8. [PMID: 26573343 DOI: 10.3109/17435390.2015.1107146]
- Ruth MM, Pennings LJ, Koeken VACM, Schildkraut JA, Hashemi A, et al. Thioridazine is an efflux pump inhibitor in Mycobacterium avium complex but of limited clinical relevance. Antimicrob Agents Chemother 2020;64(7):e00181. [PMID: 32312774 DOI: 10.1128/AAC.00181-20]
- Machado D, Pires D, Perdigão J, Couto I, Portugal I, et al. Ion channel blockers as antimicrobial agents, efflux inhibitors, and enhancers of macrophage killing activity against drug resistant Mycobacterium tuberculosis. PLoS One 2016;11(2):e0149326. [PMID: 26919135 DOI: 10.1371/journal.pone.0149326]
- Pieroni M, Machado D, Azzali E, Santos Costa S, Couto I, et al. Rational design and synthesis of thioridazine analogues as enhancers of the antituberculosis therapy. J Med Chem 2015;58(15):5842-53. [PMID: 26197353 DOI: 10.1021/acs.jmedchem.5b00428]
- Grossman TH, Shoen CM, Jones SM, Jones PL, Cynamon MH, et al. The efflux pump inhibitor timcodar improves the potency of antimycobacterial agents. Antimicrob Agents Chemother 2015;59(3):1534-41. [PMID: 25534740 DOI: 10.1128/AAC.04271-14]
- Dinesh N, Sharma S, Balganesh M. Involvement of efflux pumps in the resistance to peptidoglycan synthesis inhibitors in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2013;57(4):1941-3. [PMID: 23335736 DOI: 10.1128/AAC.01957-12]
- Mudde SE, Schildkraut JA, Ammerman NC, de Vogel CP, de Steenwinkel JEM, et al. Unraveling antibiotic resistance mechanisms in Mycobacterium abscessus: the potential role of efflux pumps. J Glob Antimicrob Resist 2022;31:345-52. [PMID: 36347496 DOI: 10.1016/j.jgar.2022.10.015]
- Martin A, Bouyakoub Y, Soumillion K, Mantu EON, Colmant A, et al. Targeting bedaquiline mycobacterial efflux pump to potentially enhance therapy in Mycobacterium abscessus. Int J Mycobacteriol 2020;9(1):71-5. [PMID: 32474492 DOI: 10.4103/ijmy.ijmy_181_19]
- Menichini M, Lari N, Rindi L. Effect of efflux pump inhibitors on the susceptibility of Mycobacterium avium complex to clarithromycin. J Antibiot (Tokyo) 2020;73(2):128-32. [PMID: 31624335 DOI: 10.1038/s41429-019-0245-1]
- Danquah CA, Kakagianni E, Khondkar P, Maitra A, Rahman M, et al. Analogues of disulfides from allium stipitatum demonstrate potent anti-tubercular activities through drug efflux pump and biofilm inhibition. Sci Rep 2018;8(1):1150. [PMID: 29348586 DOI: 10.1038/s41598-017-18948-w]
- Halicki PCB, Vianna JS, Zanatta N, de Andrade VP, de Oliveira M, et al. 2,2,2-trifluoro-1-(1,4,5,6-tetrahydropyridin-3-yl)ethanone derivative as efflux pump inhibitor in Mycobacterium tuberculosis. Bioorg Med Chem Lett 2021;42:128088. [PMID: 33964440 DOI: 10.1016/j.bmcl.2021.128088]
- Cannalire R, Machado D, Felicetti T, Santos Costa S, Massari S, et al. Natural isoflavone biochanin A as a template for the design of new and potent 3-phenylquinolone efflux inhibitors against Mycobacterium avium. Eur J Med Chem 2017;140:321-30. [PMID: 28964936 DOI: 10.1016/j.ejmech.2017.09.014]
- Felicetti T, Machado D, Cannalire R, Astolfi A, Massari S, et al. Modifications on C6 and C7 positions of 3-phenylquinolone efflux pump inhibitors led to potent and safe antimycobacterial treatment adjuvants. ACS Infect Dis 2019;5(6):982-1000. [PMID: 30907573 DOI: 10.1021/acsinfecdis.9b00041]
- Sen T, Neog K, Sarma S, Manna P, Deka Boruah HP, et al. Efflux pump inhibition by 11H-pyrido[2,1-b]quinazolin-11-one analogues in mycobacteria. Bioorg Med Chem 2018;26(17):4942-51. [PMID: 30190182 DOI: 10.1016/j.bmc.2018.08.034]
- Shyam M, Verma H, Bhattacharje G, Mukherjee P, Singh S, et al. Mycobactin analogues with excellent pharmacokinetic profile demonstrate potent antitubercular specific activity and exceptional efflux pump inhibition. J Med Chem 2022;65(1):234-56. [DOI: 10.1021/acs.jmedchem.1c01349]
- Felicetti T, Cannalire R, Pietrella D, Latacz G, Lubelska A, et al. 2-Phenylquinoline S. aureus NorA efflux pump inhibitors: evaluation of the importance of methoxy group introduction. J Med Chem 2018;61(17):7827-48. [PMID: 30067360 DOI: 10.1021/acs.jmedchem.8b00791]
- Machado D, Cannalire R, Santos Costa S, Manfroni G, Tabarrini O, et al. Boosting effect of 2-Phenylquinoline efflux inhibitors in combination with macrolides against Mycobacterium smegmatis and Mycobacterium avium. ACS Infect Dis 2015;1(12):593-603. [PMID: 27623057 DOI: 10.1021/acsinfecdis.5b00052]
- Corona P, Ibba R, Piras S, Molicotti P, Bua A, et al. Quinoxaline-based efflux pump inhibitors restore drug susceptibility in drug-resistant nontuberculous mycobacteria. Arch Pharm (Weinheim) 2022;355(8):e2100492. [PMID: 35532283 DOI: 10.1002/ardp.202100492]
- Duran M, Araiza A, Surani SR, Vakil A, Varon J. Pulmonary infection caused by Mycobacterium terrae: a case report and literature review. Cureus 2019;11(11):1-9. [PMID: 31890427 DOI: 10.7759/cureus.6228]
- Little JS, Dedrick RM, Freeman KG, Cristinziano M, Smith BE, et al. Bacteriophage treatment of disseminated cutaneous Mycobacterium chelonae infection. Nat Commun 2022;13(1):2313. [PMID: 35504908 DOI: 10.1038/s41467-022-29689-4]
- Pinto-Gouveia M, Gameiro A, Ramos L, Cardoso JC, Brites MM, et al. Mycobacterium chelonae is an ubiquitous atypical mycobacterium. Case Rep Dermatol 2015;7(2):207-11. [PMID: 26351432 DOI: 10.1159/000438898]
- Saffo Z, Ognjan A. Mycobacterium smegmatis infection of a prosthetic total knee arthroplasty. IDCases 2016;5:80-2. [PMID: 27516972 DOI: 10.1016/j.idcr.2016.07.007]
- Koul A, Arnoult E, Lounis N, Guillemont J, Andries K. The challenge of new drug discovery for tuberculosis. Nature 2011;469(7331):483-90. [PMID: 21270886 DOI: 10.1038/nature09657]
- Protopopova M, Hanrahan C, Nikonenko B, Samala R, Chen P, et al. Identification of a new antitubercular drug candidate, SQ109, from a combinatorial library of 1,2-ethylenediamines. J Antimicrob Chemother 2005;56(5):968-74. [PMID: 16172107 DOI: 10.1093/jac/dki319]
- Foss MH, Pou S, Davidson PM, Dunaj JL, Winter RW, et al. Diphenylether-modified 1,2-diamines with improved drug properties for development against Mycobacterium tuberculosis. ACS Infect Dis 2016;2(7):500-8. [PMID: 27626102 DOI: 10.1021/acsinfecdis.6b00052]
- Tahlan K, Wilson R, Kastrinsky DB, Arora K, Nair V, et al. SQ109 targets MmpL3, a membrane transporter of trehalose monomycolate involved in mycolic acid donation to the cell wall core of Mycobacterium tuberculosis. Antimicrob Agents Chemother 2012;56(4):1797-809. [PMID: 22252828 DOI: 10.1128/AAC.05708-11]
- Hoagland DT, Liu J, Lee RB, Lee RE. New agents for the treatment of drug-resistant Mycobacterium tuberculosis. Adv Drug Deliv Rev 2016;102:55-72. [PMID: 27151308 DOI: 10.1016/j.addr.2016.04.026]
- Riccardi G, Pasca MR, Chiarelli LR, Manina G, Mattevi A, et al. The DprE1 enzyme, one of the most vulnerable targets of Mycobacterium tuberculosis. Appl Microbiol Biotechnol 2013;97(20):8841-8. [PMID: 24037308 DOI: 10.1007/s00253-013-5218-x]
- Chikhale RV, Barmade MA, Murumkar PR, Yadav MR. Overview of the development of DprE1 inhibitors for combating the menace of tuberculosis. J Med Chem 2018;61(19):8563-93. [PMID: 29851474 DOI: 10.1021/acs.jmedchem.8b00281]
- Hariguchi H, Chen X, Hayashi Y, Kawano Y, Fujiwara M, et al. OPC-167832, a novel carbostyril derivative with potent. Antimicrob Agents Chemother 2020;64(6):1-13. [DOI: 10.1128/AAC.02020-19]
- Pomar V, Martínez JA. MESA: Nuevas pautas de tratamiento en TB. Analysis of pooled delamanid clinical trials data and new Otsuka TB product developments. Rev Enf Emerg 2018;17(3):116-21.
- Butler MS, Paterson DL. Antibiotics in the clinical pipeline in October 2019. J Antibiot 2020;73(6):329-64. [PMID: 32152527 DOI: 10.1038/s41429-020-0291-8]
- Shirude PS, Shandil R, Sadler C, Naik M, Hosagrahara V, et al. Azaindoles: noncovalent DprE1 inhibitors from scaffold morphing efforts, kill Mycobacterium tuberculosis and are Efficacious in vivo. J Med Chem 2013;56:9701-8. [PMID: 24215368 DOI: 10.1021/jm401382v]
- Shetye GS, Franzblau SG, Cho S. New tuberculosis drug targets, their inhibitors, and potential therapeutic impact. Transl Res 2020;220:68-97. [PMID: 32275897 DOI: 10.1016/j.trsl.2020.03.007]
- Kempf DJ, Marsh KC. Assembling pharma resources to tackle diseases of underserved populations. ACS Med Chem Lett 2020;11(6):1094-100. [PMID: 32550987 DOI: 10.1021/acsmedchemlett.0c00051]
- Andries K, Verhasselt P, Guillemont J, Göhlmann HW, Neefs JM, et al. A diarylquinoline drug active on the ATP synthase of Mycobacterium tuberculosis. Science 2005;307(5707):223-7. [PMID: 15591164 DOI: 10.1126/science.1106753]
- Cohen J. Infectious disease. Approval of novel TB drug celebrated — with restraint. Science 2013;339(6116):130. [PMID: 23307714 DOI: 10.1126/science.339.6116.130]
- Sutherland HS, Tong AST, Choi PJ, Blaser A, Conole D, et al. 3,5-Dialkoxypyridine analogues of bedaquiline are potent antituberculosis agents with minimal inhibition of the hERG channel. Bioorg Med Chem 2019;27(7):1292-307. [PMID: 30803745 DOI: 10.1016/j.bmc.2019.02.026]
- Sarathy JP, Ragunathan P, Shin J, Cooper CB, Upton AM, et al. TBAJ-876 retains bedaquiline’s activity against subunits c and ε of Mycobacterium tuberculosis F-ATP synthase. Antimicrob Agents Chemother 2019;63(10):e01191-19. [PMID: 31358589 DOI: 10.1128/AAC.01191-19]
- O’Donnell MR, Padayatchi N, Metcalfe JZ. Elucidating the role of clofazimine for the treatment of tuberculosis. Int J Tuberc Lung Dis 2016;20(12):S52-7. [PMID: 28240574 DOI: 10.5588/ijtld.16.0073]
- Dalcolmo M, Gayoso R, Sotgiu G, D’Ambrosio L, Rocha JL, et al. Effectiveness and safety of clofazimine in multidrug-resistant tuberculosis: a nationwide report from Brazil. Eur Respir J 2017;49(3):1602445. [PMID: 28331044 DOI: 10.1183/13993003.02445-2016]
- Diacon AH, Dawson R, Von Groote-Bidlingmaier F, Symons G, Venter A, et al. Bactericidal activity of pyrazinamide and clofazimine alone and in combinations with pretomanid and bedaquiline. Am J Respir Crit Care Med 2015;191(8):943-53. [PMID: 25622149 DOI: 10.1164/rccm.201410-1801OC]
- Dey T, Brigden G, Cox H, Shubber Z, Cooke G, et al. Outcomes of clofazimine for the treatment of drug-resistant tuberculosis: a systematic review and meta-analysis. J Antimicrob Chemother 2013;68(2):284-93. [PMID: 23054996 DOI: 10.1093/jac/dks389]
- Lechartier B, Cole ST. Mode of action of clofazimine and combination therapy with benzothiazinones against Mycobacterium tuberculosis. Antimicrob Agents Chemother 2015;59(8):4457-63. [PMID: 25987624 DOI: 10.1128/AAC.00395-15]
- Cholo MC, Mothiba MT, Fourie B, Anderson R. Mechanisms of action and therapeutic efficacies of the lipophilic antimycobacterial agents clofazimine and bedaquiline. J Antimicrob Chemother 2017;72(2):338-53. [PMID: 27798208 DOI: 10.1093/jac/dkw426]
- Lu Y, Zheng M, Wang B, Fu L, Zhao W, et al. Clofazimine analogs with efficacy against experimental tuberculosis and reduced potential for accumulation. Antimicrob Agents Chemother 2011;55(11):5185-93. [PMID: 21844321 DOI: 10.1128/AAC.00699-11]
- Xu J, Wang B, Fu L, Zhu H, Guo S, et al. In vitro and in vivo activities of the riminophenazine TBI-166 against Mycobacterium tuberculosis. Antimicrob Agents Chemother 2019;63(5):e02155-18. [PMID: 30782992 DOI: 10.1128/AAC.02155-18]
- Pethe K, Bifani P, Jang J, Kang S, Park S, et al. Discovery of Q203, a potent clinical candidate for the treatment of tuberculosis. Nat Med 2013;19(9):1157-60. [PMID: 23913123 DOI: 10.1038/nm.3262]
- Scherr N, Bieri R, Thomas SS, Chauffour A, Kalia NP, et al. Targeting the Mycobacterium ulcerans cytochrome bc1:aa3 for the treatment of Buruli ulcer. Nat Commun 2018;9(1):5370. [PMID: 30560872 DOI: 10.1038/s41467-018-07804-8]
- Chauffour A, Robert J, Veziris N, Aubry A, Pethe K, et al. Q203 containing fully intermittent oral regimens exhibited high sterilizing activity against Mycobacterium ulcerans in mice. bioRxiv 2019;813253. [DOI: 10.1101/813253]
- Almeida DV, Converse PJ, Omansen TF, Tyagi S, Tasneen R, et al. Telacebec for ultrashort treatment of Buruli ulcer in a mouse model. Antimicrob Agents Chemother 2020;64(6):e00259-20. [PMID: 32205344 DOI: 10.1128/AAC.00259-20]
- Matsumoto M, Hashizume H, Tomishige T, Kawasaki M, Tsubouchi H, et al. OPC-67683, a nitro-dihydro-imidazooxazole derivative with promising action against tuberculosis in vitro and in mice. PLoS Med 2006;3(11):2131-44. [PMID: 17132069 DOI: 10.1371/journal.pmed.0030466]
- van den Boogaard J, Kibiki GS, Kisanga ER, Boeree MJ, Aarnoutse RE. New drugs against tuberculosis: problems, progress, and evaluation of agents in clinical development. Antimicrob Agents Chemother 2009;53(3):849-62. [PMID: 19075046 DOI: 10.1128/AAC.00749-08]
- Singh R, Manjunatha U, Boshoff HI, Ha YH, Niyomrattanakit P, et al. PA-824 kills nonreplicating mycobacterium tuberculosis by intracellular NO release. Science 2008;322(5906):1392-5. [PMID: 19039139 DOI: 10.1126/science.1164571]
- Stover CK, Warrener P, VanDevanter DR, Sherman DR, Arain TM, et al. A small-molecule nitroimidazopyran drug candidate for the treatment of tuberculosis. Nature 2000;405(6789):962-6. [PMID: 10879539 DOI: 10.1038/35016103]
- Aslam S, Musher DM. Nitazoxanide: clinical studies of a broad-spectrum anti-infective agent. Future Microbiol 2007;2(6):583-90. [PMID: 18041899 DOI: 10.2217/17460913.2.6.583]
- Stachulski AV, Pidathala C, Row EC, Sharma R, Berry NG, et al. Thiazolides as novel antiviral agents. 2. Inhibition of hepatitis c virus replication. J Med Chem 2011;54(24):8670-80. [PMID: 22059983 DOI: 10.1021/jm201264t]
- de Carvalho LPS, Lin G, Jiang X, Nathan C. Nitazoxanide kills replicating and nonreplicating Mycobacterium tuberculosis and evades resistance. J Med Chem 2009;52(19):5789-92. [PMID: 19736929 DOI: 10.1021/jm9010719]
- de Carvalho LPS, Darby CM, Rhee KY, Nathan C. Nitazoxanide disrupts membrane potential and intrabacterial pH homeostasis of Mycobacterium tuberculosis. ACS Med Chem Lett 2011;2(11):849-54. [PMID: 22096616 DOI: 10.1021/ml200157f]
- Singh KV, Coque TM, Murray BE. In vitro activity of the trinem sanfetrinem (GV104326) against gram-positive organisms. Antimicrob Agents Chemother 1996;40(9):2142-6. [PMID: 8878596 DOI: 10.1128/AAC.40.9.2142]
- El-Gamal MI, Brahim I, Hisham N, Aladdin R, Mohammed H, et al. Recent updates of carbapenem antibiotics. Eur J Med Chem 2017;131:185-95. [PMID: 28324783 DOI: 10.1016/j.ejmech.2017.03.022]
- Vilchèze C. Mycobacterial cell wall: a source of successful targets for old and new drugs. Appl Sci 2020;10(7):2278. [DOI: 10.3390/app10072278]
- Chopra I. Research and development of antibacterial agents. Curr Opin Microbiol 1998;1(5):495-501. [PMID: 10066523 DOI: 10.1016/s1369-5274(98)80080-5]
- Tripathi RP, Tewari N, Dwivedi N, Tiwari VK. Fighting tuberculosis: an old disease with new challenges. Med Res Rev 2005;25(1):93-131. [PMID: 15389729 DOI: 10.1002/med.20017]
- Rivers EC, Mancera RL. New anti-tuberculosis drugs with novel mechanisms of action. Curr Med Chem 2008;15(19):1956-67. [PMID: 18691051 DOI: 10.2174/092986708785132906]
- Leach KL, Brickner SJ, Noe MC, Miller PF. Linezolid, the first oxazolidinone antibacterial agent. Ann N Y Acad Sci 2011;1222(1):49-54. [PMID: 21434942 DOI: 10.1111/j.1749-6632.2011.05962.x]
- Zhu T, Friedrich SO, Diacon A, Wallis RS. Population pharmacokinetic/pharmacodynamic analysis of the bactericidal activities of sutezolid (PNU-100480) and its major metabolite against intracellular Mycobacterium tuberculosis in ex vivo whole-blood cultures of patients with pulmonary tuberculosis. Antimicrob Agents Chemother 2014;58(6):3306-11. [PMID: 24687496 DOI: 10.1128/AAC.01920-13]
- Jeong JW, Jung SJ, Lee HH, Kim YZ, Park TK, et al. In vitro and in vivo activities of LCB01-0371, a new oxazolidinone. Antimicrob Agents Chemother 2010;54(12):5359-62. [PMID: 20855730 DOI: 10.1128/AAC.00723-10]
- Cho YL, Jang J. Development of delpazolid for the treatment of tuberculosis. Appl Sci 2020;10(7):2211. [DOI: 10.3390/app10072211]
- Zong Z, Jing W, Shi J, Wen S, Zhang T, et al. Comparison of in vitro activity and MIC distributions between the novel oxazolidinone delpazolid and linezolid against multidrug-resistant and extensively drug-resistant mycobacterium tuberculosis in China. Antimicrob Agents Chemother 2018;62(8):e00165-18. [PMID: 29844043 DOI: 10.1128/AAC.00165-18]
- Mdluli K, Cooper C, Yang T, Lotlikar M, Betoudji F, et al. TBI-223: a safer oxazolidinone in pre-clinical development for tuberculosis. In Proceedings of the ASM Microbe 2017, 1-5 June 2017; New Orleans, LA, USA; 2017. Session 335-AAID-Sunday-50.
- Lociuro S. BioVersys receives QIDP designation from the U.S. FDA for the development of a fixed combination of BVL-GSK098 and ETH. BVL-GSK098 is being developed for the treatment of multi-drug resistant. Press Release BioVersys. 2020;1-3.
- Rosenthal IM, Zhang M, Williams KN, Peloquin CA, Tyagi S, et al. Daily dosing of rifapentine cures tuberculosis in three months or less in the murine model. PLoS Med 2007;4(12):e344. [PMID: 18092886 DOI: 10.1371/journal.pmed.0040344]
- Faustino C, Andrade JM, Ferreira IM, Almeida JF, Rijo P. Lead molecules from natural products: insight into tubercular targets. Studies in Natural Products Chemistry; 2020;65. pp.41-84.
- Igarashi M, Ishizaki Y, Takahashi Y. New antituberculous drugs derived from natural products: current perspectives and issues in antituberculous drug development. J Antibiot 2018;71(1):15-25. [DOI: 10.1038/ja.2017.126]
- Grosset JH, Singer TG, Bishai WR. New drugs for the treatment of tuberculosis: hope and reality. Int J Tuberc Lung Dis 2012;16(8):1005-14. [PMID: 22762423 DOI: 10.5588/ijtld.12.0277]
- Madhura DB, Liu J, Meibohm B, Lee RE. Phase II metabolic pathways of spectinamide antitubercular agents: a comparative study of the reactivity of 4-substituted pyridines to glutathione conjugation. Medchemcomm 2016;7(1):114-7. [PMID: 27042286 DOI: 10.1039/C5MD00349K]
- Lee RE, Hurdle JG, Liu J, Bruhn DF, Matt T, et al. Spectinamides: a new class of semisynthetic antituberculosis agents that overcome native drug efflux. Nat Med 2014;20(2):152-8. [PMID: 24464186 DOI: 10.1038/nm.3458]
- Bruhn DF, Scherman MS, Liu J, Scherbakov D, Meibohm B, et al. In vitro and in vivo evaluation of synergism between anti-tubercular spectinamides and non-classical tuberculosis antibiotics. Sci Rep 2015;5:13985. [DOI: 10.1038/srep13985]
- Lee SFK, Laughon BE, McHugh TD, Lipman M. New drugs to treat difficult tuberculous and nontuberculous mycobacterial pulmonary disease. Curr Opin Pulm Med 2019;25(3):271-80. [PMID: 30865034 DOI: 10.1097/MCP.0000000000000570]
- Gygli SM, Borrell S, Trauner A, Gagneux S. Antimicrobial resistance in Mycobacterium tuberculosis: mechanistic and evolutionary perspectives. FEMS Microbiology Reviews. Oxford University Press; 2017; 41. p. 354-73.
- Blondiaux N, Moune M, Desroses M, Frita R, Flipo M, et al. Reversion of antibiotic resistance in Mycobacterium tuberculosis by spiroisoxazoline SMARt-420. Science 2017;1211:1206-11. [PMID: 28302858 DOI: 10.1126/science.aag1006]
- Hoehl S, Rabenau H, Berger A, Kortenbusch M, Cinatl J, et al. Evidence of SARS-CoV-2 infection in returning travelers from Wuhan, China. N Engl J Med 2020;382(13):1278-80. [PMID: 32069388 DOI: 10.1056/NEJMc2001899]
- Pasca MR, Guglierame P, Arcesi F, Bellinzoni M, De Rossi E, et al. Rv2686c-Rv2687c-Rv2688c, an ABC fluoroquinolone efflux pump in Mycobacterium tuberculosis. Antimicrob Agents Chemother 2004;48(8):3175-8. [PMID: 15273144 DOI: 10.1128/AAC.48.8.3175-3178.2004]
- Khosravi AD, Sirous M, Absalan Z, Tabandeh MR, Savari M. Comparison of drrA and drrB Efflux pump genes expression in drug-susceptible and -resistant Mycobacterium tuberculosis strains isolated from tuberculosis patients in Iran. Infect Drug Resist 2019;12:3437-44. [PMID: 31807034 DOI: 10.2147/IDR.S221823]
- Schmalstieg AM, Srivastava S, Belkaya S, Deshpande D, Meek C, et al. The antibiotic resistance arrow of time: efflux pump induction is a general first step in the evolution of mycobacterial drug resistance. Antimicrob Agents Chemother 2012;56(9):4806-15. [PMID: 22751536 DOI: 10.1128/AAC.05546-11]
- Oh TS, Kim YJ, Kang HY, Kim CK, Cho SY, et al. RNA expression analysis of efflux pump genes in clinical isolates of multidrug-resistant and extensively drug-resistant Mycobacterium tuberculosis in South Korea. Infect Genet Evol 2017;49:111-5. [PMID: 28062386 DOI: 10.1016/j.meegid.2017.01.002]
- Takiff HE, Cimino M, Musso MC, Weisbrod T, Martinez R, et al. Efflux pump of the proton antiporter family confers low-level fluoroquinolone resistance in Mycobacterium smegmatis. Proc Natl Acad Sci U S A 1996;93(1):362-6. [PMID: 8552639 DOI: 10.1073/pnas.93.1.362]
- Pasca MR, Guglierame P, De Rossi E, Zara F, Riccardi G. mmpL7 gene of Mycobacterium tuberculosis is responsible for isoniazid efflux in Mycobacterium smegmatis. Antimicrob Agents Chemother 2005;49(11):4775-7. [PMID: 16251328 DOI: 10.1128/AAC.49.11.4775-4777.2005]